Study population
This prospective observational study utilized data from SNAC-K (www.snac-k.se), involving adults aged 60 years or older, residing in a central area of Stockholm, Sweden. In the initial phase (2001–2004), participants were randomly selected and stratified into 11 age cohorts (60, 66, 72, 78, 81, 84, 87, 90, 93, 96 and over 99 years of age). A total of 3,363 individuals were included and followed every 6 years (for ages 60–72 years) or 3 years (for ages ≥78 years)56. The present study included data from the baseline to the fifth follow-up, covering the period from 2001 to 2019, with a maximum follow-up duration of 15 years. From the baseline cohort (n = 3,363), individuals with at least one missing blood-based biomarker measurement were excluded (n = 1,116), resulting in a final study sample of 2,247 participants. Overall, excluded participants were older, more likely to be female and more dependent on activities of daily living. They had a higher number of chronic diseases and exhibited reduced cognitive function. SNAC-K was approved by the Regional Ethical Review Authority in Stockholm (Dnrs: KI 01-114, 04-929/3, Ö26-2007, 2009/595-32, 2010/447-31/2, 2013/828-31/3, and 2016/730-31/1), and written informed consent was obtained from participants or next of kin.
To validate our findings in an independent cohort, we used data from the BLSA, a continuously-enrolled cohort study of community-dwelling adults started in 1958. Enrolled participants are followed up at age-dependent intervals: every 4 years, for individuals younger than 60 years, every 2 years for those aged 60–79 years, and every year for those aged 80 years or older. Further details on the study design are published elsewhere57. This study incorporated data from participants’ first assessment with complete information through a maximum follow-up duration of 15 years. Individuals aged 60 years or older with available data on baseline blood-based biomarkers, key demographic characteristics and chronic conditions were included, obtaining a final study sample of 522 participants followed up on average 6.9 years (s.d. = 4.34). This subset of BLSA participants is, on average, slightly older than other participants aged 60 years or older observed during the same period (7 August 2006 to 4 March 2025). The BLSA study was approved by the National Institutes of Health Intramural Research Program Institutional Review Board, and informed consent was obtained from each participant.
The results of this study are reported in keeping with the Strengthening the Reporting of Observational Studies in Epidemiology recommendations58.
Data collection and definitions
In the SNAC-K cohort, a comprehensive data collection process, conducted by a multidisciplinary team of physicians, nurses and psychologists, was carried out during each visit at a dedicated research center. Consistent protocols were followed throughout the clinical examinations, in-person interviews and laboratory measurements to ensure data collection uniformity. Home visits were arranged for individuals who could not attend the visit site.
For the BLSA cohort, assessments were performed by the study staff either at the clinical research unit of the Intramural Research Program of the National Institute on Aging or at the individual’s home. These assessments encompassed interviews, clinical examinations and laboratory tests.
Disease assessment
In SNAC-K, diseases were identified at each visit through anamnestic medical history, physical examination and face-to-face and/or proxy interviews. Additional diseases were identified based on laboratory parameters, medication usage and the Swedish National Patient Register. These diagnoses were subsequently coded according to the 10th revision of the International Classification of Diseases (ICD-10). An international team of physicians and epidemiologists classified diseases as chronic when they persisted over time and were associated with either ongoing disability or the need for prolonged care, treatment or rehabilitation. The diseases identified as chronic were then grouped into 60 broad categories as previously described3. Incident cases of dementia, heart failure and ischemic heart disease were defined as new diagnoses made during the follow-up. A depressive episode was defined as the occurrence of either major or minor depression, as previously described59, during the follow-up.
In the BLSA cohort, diseases were identified based on a combination of clinical observations during comprehensive physical examination, self-reported medical history, clinical laboratory parameters and medication use. Where possible classifications were made based upon the 9th revision of the ICD (ICD-9) codes, and medications were coded according to the Anatomical Therapeutic Chemical (ATC) system.
Blood-based biomarkers
In the SNAC-K cohort, nonfasting venous blood samples were collected after informed consent. Blood was allowed to clot, and the serum separated from the clotted venipuncture sample was stored at −80 °C at the Karolinska Institutet BioBank. The 54 blood-based biomarkers examined at baseline were selected based on a comprehensive literature review and expert consensus. They encompass key biological processes—metabolism, inflammation, vascular function and neurodegeneration—relevant to aging and common geriatric syndromes (for example, multimorbidity, dementia and frailty), as well as routinely collected clinical parameters (for example, hemoglobin and creatinine). Several laboratory techniques were used for biomarker quantification; further details on biomarkers and assay platforms are provided in Supplementary Tables 4 and 5.
For BLSA, venous samples are collected after an overnight fast at each study visit. Clinical laboratory measurements are completed by the Clinical Laboratory Improvement Amendments certified clinical laboratory of MedStar Harbor Hospital. Samples for research measurements are prepared and stored by the Intramural Research Program of the National Institute on Aging Clinical Core Laboratory and Biorepository. Sample processing begins within 2 hours of collection with samples stored at ≤−70 °C.
Standard blood tests
As part of the SNAC-K protocol, standard blood tests were carried out at baseline in the hematological and biochemical laboratory at the Karolinska Hospital, in accordance with standardized institutional protocols, during the 12 hours following the assessment. The routinely performed measurements included: complete blood count, hemoglobin, total cholesterol, albumin, alkaline phosphatase, calcium, creatinine, folic acid, gamma-glutamyl transferase, thyroxine, HbA1c, thyroid stimulating hormone and vitamin B12.
In the BLSA, albumin, gamma-glutamyl transferase and HbA1c are measured as part of standard clinical laboratory panels. Fasting insulin and leptin are measured in the Intramural Research Program of the National Institute on Aging Laboratory of Clinical Investigation by enzyme-linked immunosorbent assay (ALPCO and Linco Research, Inc.). Fasting insulin is measured as part of an oral glucose tolerance test in a sample collected after an overnight fast but before glucose ingestion.
Protein biomarkers quantification by multiplexed immunoassay
In the SNAC-K cohort, quantification of protein biomarkers in serum was carried out at the Affinity Proteomics unit at SciLifeLab (Stockholm) using bead-based multiplexed assays performed with a Luminex system or Quanterix’s Single Molecule Arrays (Simoa). Biomarkers associated to inflammation and vascular disease were analyzed using a custom-designed magnetic Luminex assay—human premixed multi-analyte assay (Luminex Corporation). Samples were diluted following manufacturer instructions, processed in a 384 format using a validated standard operating procedure and analyzed on a FlexMap 3D (Luminex). The software xPONENT (Luminex) provides median fluorescence intensity values for samples and standards. Median fluorescence intensity is delivered for all proteins and represents a value for relative quantification. Curve fitting, extrapolation of concentrations and graphical representation were performed with Belysa Immunoassay Curve Fitting Software (Millipore), a software environment for the secondary analysis of data acquired from Luminex instruments. Standard curves were generated using a five-parameter logistic curve fit.
NfL and GFAP were assayed using the Simoa Neurology 2-Plex B kit; Aβ40, Aβ42 and T-tau were measured using the Simoa Neurology 3-Plex A kit and ptau181 was quantified using the Simoa ptau181 Advantage V2 kit. For each kit, 25 μl of sample were diluted 1:4 and the assays were performed according to manufacturer instructions. The Quanterix instrument provides average enzyme per bead values for calibrators, controls and samples. Curve fitting, extrapolation of concentrations and graphical representations are automatically performed within the Quanterix SR-X software using calibrators, a series of known concentrations of an analyte, and a four-parameter logistic curve fit. Data below the limit of detection were replaced using a not missing at random strategy, through single-value imputation, with a value of zero.
Baseline interleukin (IL-1β, IL-6, IL-8, IL-10 and IL-12p70), INFγ and TNF were measured at Accelerator Laboratory Services (Quanterix) using Simoa CorPlex Human Cytokine Panel 1 on the Quanterix SP-XTM imaging and analysis platform.
The full list of biomarkers analyzed, technology, catalog, lot number of the reagents used and analytical parameters for each assay are summarized in Supplementary Table 5. The average of intra and inter coefficient variations were calculated on a pool of samples representative of the whole SNAC-K sample set. The pool was analyzed in 8 replicates for each 384 plate (samples were distributed in 7 plates) in the Luminex assays and in three replicates in the Quanterix assays (samples were distributed in 39 plates). Average concentration in the pool and intra- and interplate coefficient variations are indicated for each protein. The average of intraplate coefficient variations is reported.
In the BLSA, GDF15 and Cystatin C were measured using the 7k SomaScan assay v4.1 (SomaLogic). Data were normalized and cross-batch harmonization was performed as previously described60.
Baseline biomarkers were employed as continuous variables and further converted into z-score to enable standardized comparisons.
Covariates
Sociodemographic data were collected through self-reported and/or nurse-administered questionnaires during SNAC-K visits. For people with cognitive impairment, information was gathered by interviewing a proxy or caregiver. Sociodemographic variables included age, sex, education level (elementary versus high school or above), marital status (partnered versus unpartnered) and residential status (institutionalized versus noninstitutionalized). In addition, clinical and functional status were collected through physical examination, including BMI, the number of activities of daily living lost, physical activity level (active versus inactive, based on self-reported intensity and frequency of physical activity engagement61), global cognitive function (evaluated through the MMSE score) and the number of prescribed daily medications (self-reported or obtained from medical records for participants who were institutionalized). All-cause mortality was retrieved through the Swedish Cause of Death Register and survival status was censored at the end of the study period.
Demographic characteristics of BLSA participants are collected in structured interviews conducted by trained study staff. Anthropometric measurements, including BMI, are objectively assessed using standardized protocols.
Statistical analyses
Baseline characteristics of the sample are reported as mean (s.d.) or median (interquartile range) for continuous variables, and count (%) for categorical variables.
Multimorbidity measures and latent class analysis
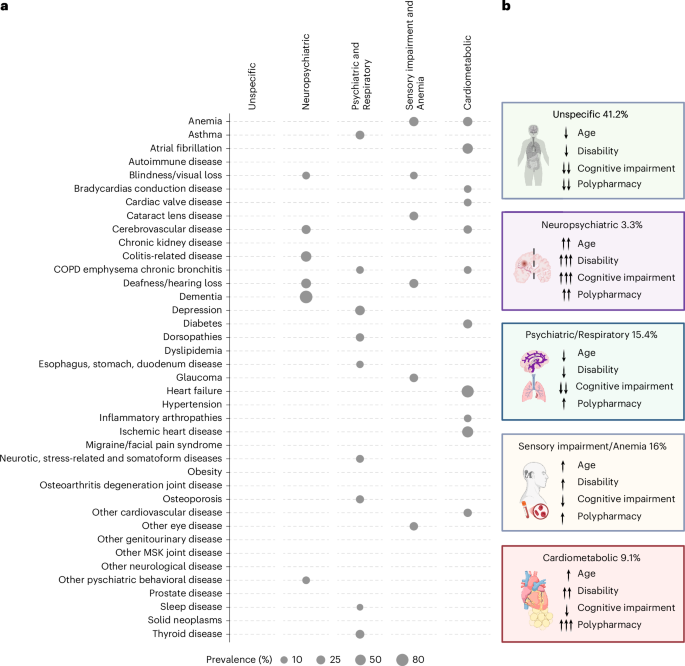
Multimorbidity measures were operationalized as follows. First, multimorbidity was defined as the number of chronic conditions, both at the cross-sectional and longitudinal levels. In addition, baseline chronic conditions with a prevalence of at least 2% in the overall multimorbid population were used to identify homogeneous groups of participants with multimorbidity (that is, two or more diseases), sharing similar combinations of chronic diseases (Supplementary Table 8)6,62. These baseline multimorbidity patterns were identified through LCA and further validated against several distal outcomes (Supplementary Tables 2 and 3). In the context of multimorbidity patterns identified through LCA, the term ‘homogeneous’ refers to the fact that participants within the same pattern tend to exhibit similar combinations of chronic conditions. The optimal number of patterns was determined using the adjusted Bayesian Information Criterion and theoretical interpretability (Extended Data Fig. 4). In addition, LCA models with different numbers of patterns were also compared in terms of entropy and assignment accuracy through five-fold cross validation to assess the stability and separation of the latent classes (Extended Data Fig. 4). Each pattern was labeled based on the distinctive set of chronic diseases that were disproportionately represented, that is overexpressed within that group. Chronic conditions were considered to be overexpressed in each pattern if they had an O/E ratio of ≥2 and an exclusivity of ≥25% (Supplementary Table 8)6,62,63,64. The O/E ratio was calculated by dividing the prevalence of a given disease within a specific pattern by its prevalence in the overall multimorbid population, while exclusivity was calculated by dividing the absolute frequency of a given disease within a specific pattern by its frequency in the overall multimorbid population. Furthermore, the LCA model provides, for each participant, a probability of belonging to one of the latent multimorbidity patterns based on their combination of baseline chronic diseases. While the specific disease combination makes participants more likely to belong to that pattern, they may still exhibit additional chronic diseases not overexpressed in their assigned pattern. To give a descriptive overview of the five patterns in terms of key baseline characteristics, participants were assigned to the pattern with the highest probability of membership (Supplementary Table 1). However, when incorporating classes derived from a LCA model into subsequent statistical models, it is essential to account for the inherently probabilistic nature of class membership. Assigning participants solely to their most likely class can lead to overly confident and potentially biased associations in downstream models. To account for the probabilistic nature of pattern membership, each subsequent analysis concerning multimorbidity patterns was performed across 1,000 resampled multimorbidity pattern membership datasets derived from the LCA models. Specifically, for each participant, class membership was sampled 1,000 times from a multinomial distribution parameterized by their class membership probabilities. Subsequently, each dataset was used to fit a regression model, enabling participants to contribute to each multimorbidity pattern in proportion to their membership probabilities across iterations.
For the clinical validation of the multimorbidity patterns against distal outcomes, the following models were employed across the 1,000 LCA-resampled multimorbidity pattern membership datasets: logistic regression models adjusted for age, sex and education for incident cases of dementia, heart failure and ischemic heart disease; Andersen–Gill Cox model for recurrent depressive events with age as time scale, adjusted by sex, education and past depressive episodes (time-dependent covariate) and Cox model with age as time scale, adjusted for sex and education for all-cause mortality. All models were fitted using, as the independent variable, the six baseline multimorbidity groups using no multimorbidity as the reference group. For each analysis, the final coefficients with their relative standard errors were calculated using Rubin’s rule.
Cross-sectional analysis
To assess the relationship between baseline biomarkers, a correlation matrix was constructed using Spearman’s rank correlation coefficients. To account for potential confounding effects of age, age-adjusted correlations were also calculated (Extended Data Fig. 1).
To address multicollinearity and select the most important predictors, LASSO with L1 regularization was performed to model the associations between baseline biomarkers and multimorbidity measures. A random seed was set for reproducibility. For the association between biomarkers and the baseline number of diseases, Gaussian LASSO adjusted for age, sex and education was applied with cross validation using 200 folds (10% of the sample) to optimize the regularization parameter. Model selection was based on mean squared error, and lambda 1se criterion was chosen to select the most regularized model such that the cross-validated error was within one standard error of the minimum.
Similarly, multinomial LASSO adjusted for age, sex and education was used to assess associations between biomarkers and baseline multimorbidity patterns, with a 200-fold cross validation. The multinomial LASSO was performed across 1,000 resampled multimorbidity pattern membership datasets derived from the LCA models. For each biomarker, the final coefficient was calculated as the mean of the nonzero coefficients across models. Model selection was based on deviance, and lambda 1se was applied as regularization criterion. Coefficients were reported using the No-multimorbidity group as the reference category. Biomarkers were considered repeatedly associated with a given pattern if they were selected in at least 70% of the LASSO models. Final coefficients for these biomarkers were calculated as the mean of the nonzero coefficients across the models.
The nonzero coefficients of the LASSO models indicate that the corresponding biomarkers contribute independently to the prediction of the dependent variables (that is, the multimorbidity measures). The absolute value of the coefficient reflects the strength of the association, while the sign of the coefficient indicates the effect (direct or reverse). Coefficients that are shrunk to zero imply that the corresponding biomarkers do not contribute meaningfully to the model when considering all other included variables.
Longitudinal analysis
Linear mixed-effect models with random intercepts and random slopes were used to assess the accumulation of diseases over time. The model was adjusted for age, sex and education. The estimated random slopes from the linear mixed-effect model were then used as outcomes in the Gaussian LASSO model to identify the biomarkers associated with a steeper trajectory of accumulation of disease.
Principal component analysis
To identify clusters and reduce dimensionality, PCA was performed. For each multimorbidity measure, PCA was performed on biomarkers selected by the LASSO analysis (that is, those with nonzero coefficients in the LASSO models), both in cross-section and longitudinally. The number of the principal components considered was selected considering the percentage of the total explained variance (minimum threshold set at 40%).
External validation of longitudinal findings
We first harmonized the chronic conditions available in BLSA using ICD and ATC codes, as well as study visit data to align with those in the SNAC-K dataset. The baseline characteristics of the BLSA study sample are reported in Supplementary Table 7. Due to limited biomarker availability in the BLSA, it was not feasible to replicate the full LASSO models developed in SNAC-K cohort. Instead, we validated the results of the LASSO analysis—which identified biomarkers associated with the rate of disease accumulation in SNAC-K, our most important results—by assessing the predictive accuracy of the LASSO-derived model in an independent cohort. We used MSE as the accuracy metric, consistent with the metric used during model training. This approach represents a standard and widely-accepted method for validating LASSO results, as it tests the model’s ability to generalize by applying it to an external dataset and comparing predictive performance65,66. This validation approach was particularly appropriate given that the BLSA cohort represents approximately 25% of the size of SNAC-K cohort and that not all biomarkers were measured, making full model retraining unfeasible but still allowing for a meaningful assessment of generalizability through external prediction. To evaluate the external performance of the SNAC-K LASSO-selected biomarkers in the BLSA dataset, we applied a multistep approach: (1) individual rates of disease accumulation in BLSA were estimated using a linear mixed-effects model with random intercept and random slope; (2) an age-adjusted linear regression model was fitted in BLSA using the estimated slopes as outcomes and the selected biomarkers as predictors; (3) the SNAC-K LASSO-derived coefficients were then applied to the BLSA data to generate predicted individual rates of disease accumulation; and (4) predictive accuracy was assessed by calculating the MSE between the predicted and observed slopes, and compared to the MSE obtained in SNAC-K. Statistical analysis was conducted using R software (version 4.2.3). Specifically, poLCA, glmnet, lme4, factoextra, corrplot, survival and ggplot2 packages were used for the analysis. The level of statistical significance was set at P < 0.05.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.