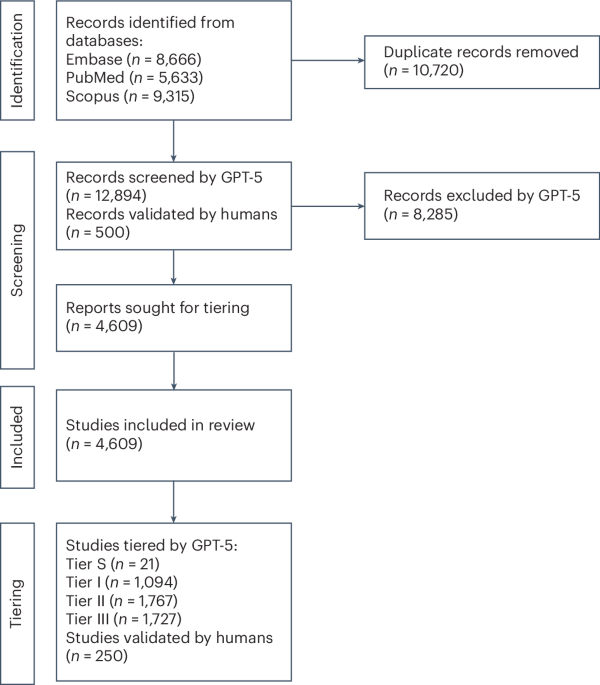
Search strategy and study selection
This review was conducted and reported in accordance with the Preferred Reporting Items for Systematic Reviews and Meta‑Analyses (PRISMA) 2020 guidelines. This review was not prospectively registered. A protocol was not prepared for this work.
We created a system for levels of evidence for LLM-based medical studies. We then introduced a scalable, LLM-assisted framework for evidence-tiered systematic review, and used this framework to assist in a systematic review of published studies evaluating LLMs in clinical medicine.
We searched PubMed, Embase and Scopus for studies published between 1 January 2022 and 6 September 2025. These databases were chosen to ensure comprehensive coverage of biomedical and clinical research. Search terms combined general descriptors of LLMs with specific model names (for example, GPT, ChatGPT, LLaMA, Claude, Gemini and Bard). To maximize specificity, we limited results to original research (articles, conference papers, preprints and letters) in health-related subject areas, and excluded reviews, meta-analyses, surveys and commentaries.
Database query string for PubMed
The data query string that was used for PubMed was as follows:
(“large language model”[Title/Abstract] OR “LLM”[Title/Abstract] OR “GPT”[Title/Abstract] OR “ChatGPT”[Title/Abstract] OR “LLaMA”[Title/Abstract] OR “Claude”[Title/Abstract] OR “Gemini”[Title/Abstract] OR “Bard”[Title/Abstract]) AND (humans[MeSH Terms]) NOT (review[Publication Type] OR meta-analysis[Publication Type] OR survey[Title])
Database query string for Scopus
The data query string that was used for Scopus was as follows:
TITLE-ABS (“large language model” OR llm OR gpt OR chatgpt OR llama OR claude OR gemini OR bard) AND PUBYEAR > 2021 AND PUBYEAR < 2026 AND (LIMIT-TO (SUBJAREA, “MEDI”) OR LIMIT-TO (SUBJAREA, “HEAL”) OR LIMIT-TO (SUBJAREA, “NURS”)) AND (LIMIT-TO (DOCTYPE, “ar”) OR LIMIT-TO (DOCTYPE, “le”) OR LIMIT-TO (DOCTYPE, “cp”))
Database query string for Embase
The data query string that was used for Embase was as follows:
(’large language model’:ab,ti OR ’llm’:ab,ti OR ’gpt’:ab,ti OR ’chatgpt’:ab,ti OR ’llama’:ab,ti OR ’claude’:ab,ti OR ’gemini’:ab,ti OR ’bard’:ab,ti) AND [humans]/lim NOT (’review’/it OR ’meta analysis’/it OR survey:ti) AND (2022:py OR 2023:py OR 2024:py OR 2025:py) AND (’Article’/it OR ’Article in Press’/it OR ’Conference Paper’/it OR ’Letter’/it OR ’Preprint’/it)
All retrieved records were deduplicated by digital object identifier and exact title. Titles and abstracts were then screened against predefined eligibility criteria by GPT-5 (methodology described below): studies were included if they reported original evaluations of LLMs on clinical tasks, and excluded if they used non-LLM models or applied LLMs solely to nonclinical contexts (for example, literature summarization or abstract screening). We additionally conducted a blinded manual human review of 500 randomly chosen studies from the initial pull (before LLM screening) to validate and characterize LLM screening performance rigorously. The precise inclusion and exclusion criteria (given to both humans and the LLM) can be found in the ‘screening_intructions.txt’ file under the Prompts folder on the GitHub repository.
Automated screening
Due to the immense number of unique studies found in the initial study pull, it was infeasible to screen for inclusion and/or exclusion entirely by hand. To undertake this challenge, we implemented an LLM-based screening pipeline using GPT-5 (reasoning mode) via OpenAI’s application programming interface and validated this approach by comparing it to human screening performance on a smaller subset of the data. We set GPT-5’s reasoning mode to ‘high’ for tasks that required complex decision-making, specifically inclusion and/or exclusion screening, evidence tier assignment and structured data extraction. We set reasoning mode to ‘minimal’ for tasks that were standard and well-validated for language models, specifically natural language classification tasks. Many frontier models are available for use in this screening process. Ideally, we would have performed benchmarks and compared the results of full analysis for several frontier models, but this proved to be prohibitively expensive at scale. As a result, we chose to use the most recently released frontier model, which was GPT-5 at the time of analysis.
Each title and abstract pair was submitted with a standardized prompt instructing the model to classify the record as ‘include’ if it evaluated one or more LLMs on a clinical task and to exclude if the study used a non-LLM model (for example, convolutional networks, recurrent neural networks or vision transformers) or if the study used an LLM in a healthcare-related context but for a nonclinical task (for example, abstract screening, paper-writing, electronic health record data extraction). The full screening prompt is available in the GitHub repository. In total, 4,609 studies were included out of the pulled studies.
Automated tiering
We additionally implemented a secondary ‘tiering’ phase, where we instructed GPT-5 with reasoning set to ‘high’, using a custom prompt (available in the GitHub repository) to perform tiering. The model was instructed to assign each included study to one of several tiers:
-
(1)
Tier S: real-world, prospective evaluations of a deployed system in a live clinical environment. These studies evaluate LLMs in a randomized, controlled, blinded (if applicable) study on real patient data in a clinically relevant, real-world task. We consider this the most robust tier of evidence, as results directly represent the effect of the LLM intervention on defined outcomes.
-
(2)
Tier I: retrospective or prospective evaluations on real, never-before-seen clinical data. The LLM neither needs be deployed in a live setting nor needs the evaluators to be blinded to the methodology. We consider this to be one of the strongest tiers of evidence available, providing preliminary predictions on how a similarly designed Tier S study may perform.
-
(3)
Tier II: simulated clinical situations, open-ended free-response questions and subjective patient ratings. Data are not taken directly from a real clinical setting and are usually post-processed or synthesized, yet they still represent a task or scenario relevant to clinical practice (for example, simulated patient conversations, common online questions and specially designed open-ended vignettes: ‘what are the best next steps?’). We consider this a medium tier of evidence, as these studies assess clinical competency to some degree, but do not make predictions on how a system would perform in a real clinical setting.
-
(4)
Tier III: board exams, multiple-choice exams and case-studies and vignettes with a clear-cut answer (for example, ‘what’s the diagnosis?’). Typically, performance is measured in accuracy. These studies represent the lowest tier of evidence: they offer little insight into real-world clinical performance aside from knowledge retrieval and synthesis, a task at which LLMs have already been shown to perform robustly.
Screening human validation
Although frontier LLMs are generally recognized to be highly performant in classification and data extraction tasks, we sought to validate the screening and tiering performance of our frontier LLMs to validate our approach and determine robust statistical bounds on the number of each study found.
For inclusion screening validation, we took a random subset of 500 articles (without replacement) from the deduplicated data pull and assigned two independent groups of human screeners (five nonoverlapping screeners each) to assign each study to the inclusion or exclusion group. Additionally, an independent screener provided tiebreaks for studies upon which groups disagreed. Screeners consisted of third- and fourth-year medical students, graduate students and residents. We computed several metrics for agreement between the LLMs. First, we compared the agreement (Cohen’s κ) between the human screener groups. Next, we computed the agreement between the LLM decisions and the tiebroken human data, as well as characterizing various statistics around LLM performance compared to ground truth.
Tiering human validation
For tiering, we took a random subset of 250 articles (without replacement) from the studies deemed to be included by the LLM to be tiered by humans. We again proceeded with two randomly assigned groups of humans (four nonoverlapping screeners each) to tier the studies. Each group conducted a first pass to remove any studies that were mistakenly marked to be included by the model, leaving 234 valid studies to tier. Each human group then assigned each study to a tier. In a similar manner to the inclusion and exclusion criteria, interhuman agreement, human–LLM agreement for each screening group, and the agreement between tiebroken human decisions (serving as the ground truth) and the LLM were computed.
Next, treating the tiebroken human screening decisions as the ‘ground truth’ decision set, we computed the agreement between the LLM and the ground truth set, as well as sensitivity and specificity. We used these values to compute error bounds on the true number of included and excluded studies.
Unsupervised data extraction
For each included study, we extracted a plethora of metadata from the title and abstract of each study using OpenAI’s GPT-5 (reasoning set to ‘high’). Specifically, we extracted (if applicable) the model(s) evaluated, the specialties/subspecialties related to the study, the type of human evaluator (medical student, resident, fellow, attending and so on), the type of dataset used in the study, whether the LLM performance was found to be superior to humans and the type of data source (for example, clinical notes, vignettes, patient messages and so on). The full prompt used for this data extraction is available in our GitHub repository.
Given the diverse range of responses, the set of responses for each metadata field was manually reviewed and an overarching set of categories for each metadata field was curated. For example, ‘GPT-4o, ChatGPT, Gemini’ would all belong to ‘Closed-source general LLM.’ A full list of categories can be found in the GitHub codebase. Next, the free-form data along with the title and abstract were re-input into GPT-5 (reasoning set to ‘minimal’) with instructions to classify the free-form data via the aforementioned category lists. This allowed for a dramatic reduction in the number of unique terms parsed by the LLM and enabled broad categorical analysis. Given that other performance metrics were strong, that this was purely an extraction task and that there was an overwhelming amount of metadata extracted, we opted not to perform human validation on this task.
Statistical analysis
To estimate the range of sensitivity, specificity and Cohen’s κ of the LLM in the inclusion and exclusion phase, we first resampled the human–LLM agreement data with replacements 50,000 times and computed the sensitivity and specificity of each sample. We then formed the 95% CI by taking the 1,250th (2.5th percentile) and 48,750th (97.5th percentile) values from these samples. To estimate the range of true positive, true negative, false positive and false negative values, we first modeled the true posterior prevalence distribution based on the tiebroken human inclusion and exclusion decisions (which we considered to be the ground truth) as a beta distribution with a Jeffreys prior, beta(I + 0.5, E + 0.5), where I is the number of included studies and E is the number of excluded studies. Next, we took each of the 50,000 sensitivity and specificity draws, sampled a prevalence, π, from the beta posterior, and propagated sensitivity, specificity and prevalence to each metric of interest. Finally, we took the 2.5th and 97.5th percentiles of each resulting distribution.
To estimate the true counts of the Tier S + I, II and III studies (Tier S was grouped with Tier I due to an extremely small sample size), we constructed a Bayesian hierarchical Dirichlet-multinomial model that jointly inferred the population prevalence of each tier and the LLM’s tier-specific misclassification rates. The model placed Dirichlet priors on the prevalence vector and each row of the confusion matrix and then fitted them to both the 234 ground truth human studies and the 4,609 LLM-assigned totals via a multinomial likelihood. We used Markov chain Monte Carlo sampling to obtain a large number of posterior samples from this distribution, using the 2.5th and 97.5th percentiles to construct our confidence interval.
Specifically, let \(\varphi =({\varphi }_{I},{\varphi }_{II},{\varphi }_{III})\) denote the population-level prevalences of Tiers S + I, II and III and let \(\varTheta \in [0,1{]}^{3\times 3}\) be the LLM confusion matrix whose i‑th row gives the probabilities that a true tier‑i study is labeled as each tier by the model. We assign independent Dirichlet(1, 1, 1) priors to \(\varphi\) and to each row of \(\varTheta\). For the audit, the 3 × 3 table of human–LLM agreements, M, is modeled as row‑wise multinomials:
$$\begin{array}{ll}{M}_{i,\cdot }| {\varTheta }_{i,\cdot } \sim \text{Multinomial}({n}_{i},{\varTheta }_{i,\cdot }), & {n}_{i}=\mathop{\sum }\limits_{j}{M}_{ij}\end{array}$$
For the remaining Ntot = 4,609 studies, we observe only the LLM totals
$${\bf{T}}=({T}_{I},{T}_{II},{T}_{III}).$$
Marginalizing the unknown true labels gives the likelihood:
$$\begin{array}{ll}{\bf{T}}| (\varphi ,\varTheta ) \sim \text{Multinomial}({N}_{\text{tot}},{\boldsymbol{\pi }}), & {\pi }_{j}=\mathop{\sum }\limits_{i=1}^{3}{\varphi }_{i}{\varTheta }_{ij}.\end{array}$$
The posterior can thus be modeled as:
$$\begin{array}{l}p(\varphi ,\varTheta | {\bf{M}},{\bf{T}})\propto \text{Dir}(\varphi ;{\bf{1}})\mathop{\prod }\limits_{i}\text{Dir}({\varTheta }_{i,\cdot };{\bf{1}})\mathop{\prod }\limits_{i}\text{Mult}({{\bf{M}}}_{i,\cdot };{n}_{i},{\varTheta }_{i,\cdot })\\ \text{Mult}({\bf{T}};{N}_{\text{tot}},{\boldsymbol{\pi }})\end{array}$$
We sample from this posterior via Markov chain Monte Carlo, with four NUTS chains (4,000 draws each, 2,000 warm up steps, accept ratio of 0.9). Convergence is confirmed by \(\{\hat{{\rm{R}}}\}\, < \,1.01\) and effective sample sizes >400 for all parameters. The true tier counts are recovered deterministically as \({{\bf{N}}}^{{\rm{S}}}={\varphi }^{{\rm{S}}}{\text{N}}_{\mathrm{tot}}\) for each posterior draw, s; the median of these draws is reported as the point estimate, and the 2.5th and 97.5th percentiles provide 95% credible intervals. All modeling was implemented in PyMCv5.
Comparison of tier counts was performed via the Bonferroni-corrected two-sided t-test.
Comparison of the rates of increase of the number of studies published per month between each tier was performed via pairwise comparisons of the linear-regression slopes using independent two-sample Welch t‑tests on the slope estimates, with degrees of freedom computed by the Welch–Satterthwaite approximation.
Pairwise comparisons between tiers of the proportion of studies where LLMs outperformed humans were conducted with one-sided, two-sample z-tests with Bonferroni correction for multiple comparisons.
The year-over-year rate of change in the proportion of studies where LLMs outperformed humans was analyzed by fitting a binomial (logistic) regression of the binary outperformance indicator on publication year and comparing it to an intercept-only model via a likelihood-ratio test.
Comparison of the proportion of studies where LLMs outperformed humans across levels of experience was performed via pairwise one‑sample z‑tests for proportions, with P values adjusted for multiple comparisons using the Benjamini–Hochberg false discovery rate procedure with a false discovery rate of 5%. We chose false discovery rate over the more conservative Bonferroni correction because, with a large family of pairwise tests, controlling the expected proportion of false discoveries (rather than the probability of any false positive) preserves statistical power and is more appropriate for this exploratory analysis context.
Because this review was designed as a descriptive, bibliometric and methodological mapping of the LLM-in-clinical-medicine literature—rather than an evaluation of the magnitude or direction of treatment effects—we did not conduct a conventional study-level risk of bias assessment. We did not pool effect estimates, quantitatively compare interventions or base recommendations on individual study results; consequently, risk of bias judgments would neither have been comparable across the highly heterogeneous set of included designs (ranging from randomized trials and observational studies to simulation and exam-style evaluations) nor altered our analyses or conclusions. Instead, we captured the aspects of methodological rigor that were relevant to our objectives through a prespecified evidence tier framework (Tiers S/I/II/III, reflecting data realism and study design) and by explicitly quantifying misclassification error in our LLM-assisted screening and tiering pipeline against human-validated labels. For these reasons, a formal study-level risk of bias assessment was considered not applicable and was therefore not performed.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.