Ethical aspects
The TCGA, PLCO and NLST cohorts are open-source datasets. Access to PLCO and NLST was applied for via the Cancer Data Access System portal (cdas.cancer.gov). All study steps were performed in accordance with the Declaration of Helsinki. This study was approved by the Ethical committee of the University of Cologne (20-1583), Halle/Cologne/Essen joint 22-1233 (Project FED-PATH/BMBF). Given the retrospective/archive nature of used data, the necessity of obtaining patients’ permission was waived by the ethical committee.
Patient cohorts, H&E-based analysis
Multiple retrospective patient cohorts of primarily resected malignant tumors (LUAD, LUSC, COAD, BRCA and HNSC) were included in the study. Inclusion criteria were the presence of one or more representative H&E slides per tumor and the absence of neoadjuvant therapy. Detailed clinicopathological characteristics of patients are presented in Supplementary Table 1.
LUAD included five subcohorts: the TCGA cohort (source: >30 departments, mostly in the USA; n = 245), the PLCO trial cohort (multi-institutional trial, USA; patient n = 215), the NLST cohort (multi-institutional trial, USA; n = 206), the UKK cohort (part of the national FEDPATH study cohort; n = 561) and the UKE cohort (part of the national FEDPATH study cohort; n = 197). All LUAD subcohorts are resection specimens covering broad ranges of Union for International Cancer Control (UICC) pT/pN stages. The TCGA cohort contains information on CD274 (PD-L1) mRNA expression. The UKK/UKE subcohort cases partially contained information on PD-L1 expression on tumor cells (TPS). Overall survival (OS)/CSS clinical endpoint information was available in all subcohorts. PFS endpoint information was available in part of the cases in the TCGA, NLST and UKK subcohorts. In all analyses, when not stated otherwise, the TCGA cohort was used as the exploration cohort and the merged PLCO/NLST/UKK/UKE cohort as an independent test cohort.
LUSC included five cohorts from same institutional sources as LUAD: TCGA (n = 463), PLCO (n = 103), NLST (n = 86), UKK (n = 415) and UKE (n = 79). All LUSC subcohorts are resection specimens covering broad ranges of UICC pT/pN stages. The TCGA cohort contains information on CD274 (PD-L1) mRNA expression. UKK/UKE subcohort cases partially contained information on PD-L1 expression on tumor cells (TPS). OS/CSS clinical endpoint information was available in all subcohorts. PFS endpoint information was available in part of cases in TCGA, NLST and UKK subcohorts. In all analyses, when not stated otherwise, the TCGA cohort was used as the exploration cohort and the merged PLCO/NLST/UKK/UKE cohort as an independent test cohort.
The COAD collective included four cohorts: TCGA (n = 342), PLCO (n = 645), University Hospital Halle (HAL; part of the national FEDPATH study cohort; n = 183) and UKK (n = 257; only for the MSI prediction use case). All COAD subcohorts are resection specimens covering broad ranges of UICC pT/pN stages. The TCGA, HAL and UKK cohorts contained information on MSI status (TCGA: based on DNA analysis, MSI-high cases were considered a correlate to mismatch-repair deficiency in other cohorts; UKK and HAL: using mismatch-repair deficiency and immunohistochemistry and polymerase chain reaction confirmation in unclear cases). OS/CSS clinical endpoint information was available in all subcohorts. PFS endpoint information was available in the TCGA cohort. In all analyses, when not stated otherwise, the TCGA cohort was used as the exploration cohort and the merged PLCO/HAL cohort as an independent test cohort (prognostic analysis).
BRCA included two subcohorts: TCGA (n = 807) and PLCO (n = 492). All BRCA subcohorts are resection specimens of female patients covering broad ranges of UICC pT/pN stages, hormone receptor status and different histological subtypes. Histological grading was available in the PLCO cohort and not available in the TCGA cohort. OS/CSS clinical endpoint information was available in all subcohorts. PFS endpoint information was available in the TCGA cohort. In all analyses, when not stated otherwise, the TCGA cohort was used as exploration cohort and the PLCO cohort as an independent test cohort.
The HNSC use case included two cohorts: UKK (n = 140) and University Hospital Giessen (only for the HPV-status-prediction use case; n = 152; HPV/p16-positive n = 35 (ref. 29)). The UKK cohort is a resection cohort covering broad ranges of UICC pT/pN stages with available information to the OS clinical endpoint. In all prognostic and correlation analyses, the TCGA LUSC cohort (both squamous cell carcinomas) was used as the exploration cohort and the UKK cohort as an independent test cohort.
Patient cohort, multiplexed tissue data analysis
For the analysis of SPARK using multiplexed tissue characterization data, we used the METABRIC BRCA cohort32 with available imaging mass cytometry data. A total of 625 primary tumors were available for analysis, with one TMA region analyzed in most patients (Fig. 5j). For all patients, information on pT (tumor size), pN, histological grade and receptor status was available (see Extended Data Fig. 8a for details).
WSI analysis pipeline, input data types
We developed a computationally efficient pipeline for preprocessing single WSIs as input objects for SPARK (Fig. 1f). This pipeline integrates multilayered information generated by different algorithms, which is efficiently preprocessed and stored as WSI objects (for preprocessing details, see below). The steps include tissue detection, quality control, multiclass tissue segmentation, tumor region mask filtering and single-cell segmentation and classification.
WSI analysis pipeline, quality control and foreground/tissue segmentation
We used our previously developed GrandQC tool48 for tissue detection in WSIs and segmentation of artificially changed regions (seven types of artifacts; using MPP1.5 version), which were excluded from any downstream analysis.
WSI analysis pipeline, multiclass tissue segmentation
Previously developed and extensively validated organ-specific semantic segmentation models were implemented to produce detailed multiclass tissue segmentation maps. Briefly, these models are based on UNet++ decoders with EfficientNet encoders and were trained on large, high-quality, manually annotated datasets. They analyze WSIs in 512 px × 512 px regions at a microns-per-pixel resolution of 1.0, roughly corresponding to ×10 objective magnification. In independent validation using external datasets, these models demonstrated excellent generalization, achieving Dice scores above 0.89 for multiclass segmentation. For details on development and validation, we refer to the original studies4,49.
For lung cancer, the model detects 12 classes4; for colorectal cancer, 11 classes49; and for breast cancer, 12 classes. All models identify tumor-relevant classes (epithelial and stromal compartments, necrotic debris, mucin, tertiary lymphoid structures) as well as numerous benign classes. All WSIs from all cohorts were processed with the models to generate multiclass tissue segmentation masks (example in Fig. 1f). From these masks, all classes other than tumor (epithelial tumor compartment) and tumor stroma were excluded. These two classes underwent minimal postprocessing to remove false-positive tumor and stroma detections, retaining only tumor stroma adjacent to tumor. The filtering methodology for tumor regions was adapted from our recent study12. For the HNSC cohort, given its shared morphology with LUSC (squamous cell carcinoma), the lung cancer model was used for multiclass segmentation. However, this model does not include all benign classes relevant to HNSC (for example, squamous mucosa, specific submucosal glands), and HNSC slides often exhibit numerous mechanical or electrocauterization artifacts. To address these limitations, an experienced board-certified pathologist (Y.T.) annotated the tumor regions to be analyzed. The lung cancer model was then applied to these annotated regions for automatic segmentation.
WSI analysis pipeline, single-cell detection and classification
Previously developed single-cell detection and classification for H&E was used to detect and classify all cells within the filtered tumor region (Fig. 1f). Briefly, this algorithm was developed using a large dataset of manual annotations (cell n = 1,272,506) for 8 cell types (tumor cells, benign epithelial cells, lymphocytes, plasma cells, macrophages, neutrophilic granulocytes and eosinophilic granulocytes) and 8 different tumor types (including all tumor types studied in this work). Macrophage ground truth was generated using immunohistochemistry stain for CD163, which was transferred to H&E images. The model was trained using the HoverNext algorithm50 with a convnextv2_large backbone at microns per pixel 0.5 (approximately ×20 magnification) and demonstrated segmentation/classification performance comparable to or better than recent state-of-the-art algorithms in independent validation51. All WSIs from all cohorts were analyzed using the model (example output in Fig. 1f). Additional postprocessing was applied to remove potential false classifications: (1) all cell detections outside tumor and tumor stroma regions in filtered masks were discarded; (2) very small objects (minimum cell-polygon size in µm2: tumor cells, 15; connective cells, 6; all other cell types, 4) were removed; (3) all ‘benign’ and ‘connective’ detections in the epithelial tumor compartment were relabeled as tumor cells (given the review showing this type of misclassification and robustness of tumor region segmentation for this filtering); and (4) all ‘tumor’ and ‘benign’ detections in the tumor stroma (mostly single cells in isolated regions) were assumed to be false positives and discarded. All filtered cell detections (nuclei polygons with cell-type classification) were saved as GeoJSON files for SPARK input.
Preprocessing of multiplexed data
First, we excluded TMA regions containing only benign tissue or fewer than 10 tumor cells. Compared to WSI analysis, we used a slightly different approach for METABRIC. As only small regions (TMA spots, ~1 mm) were available, we did not use tiling and instead applied SPARK to the entire region. Tissue segmentation masks and single-cell detection/classification masks were prepared as input for the SPARK pipeline. Tissue segmentation masks included the epithelial and stromal tumor compartments and were used by SPARK to identify the affiliation of each single cell.
We also extracted blood vessel areas from the mask and binarized this information. These vessel areas were filled and then converted into disjoint polygons. Along with the cell objects and tissue segmentation mask, a list of polygons for vessel objects (as an additional subregion within the stroma) was also used as SPARK input. All epithelial tumor cell classes were merged into a single class, and all single-cell detections (nuclear polygons) for a total of 14 cell classes were transformed into GeoJSON masks for use as SPARK input (Fig. 6j).
SPARK agentic workflow, modular structure
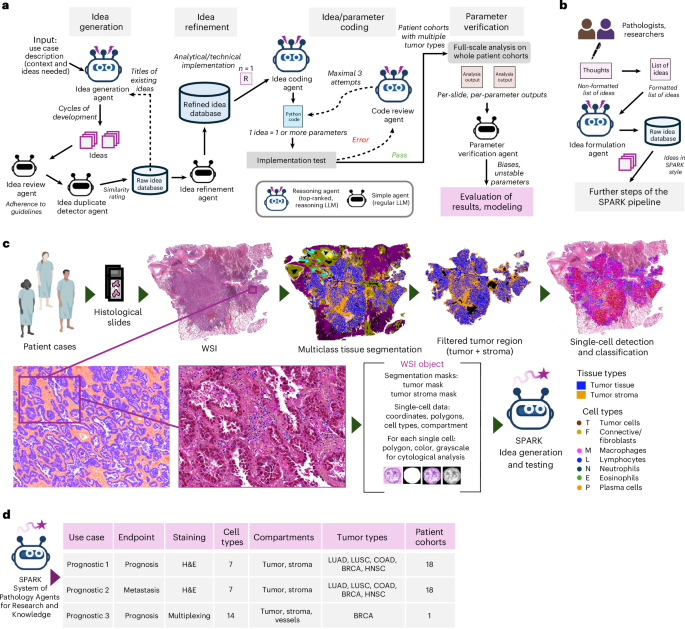
We developed SPARK as a modular agentic workflow, where the output of each step serves as the input for subsequent steps (Fig. 1c). SPARK comprises four main agentic modules: idea generation, idea refinement, idea/parameter coding and parameter verification. An additional module was implemented to enable human interaction and human-triggered idea generation (Fig. 1d).
SPARK idea generation pipeline
The principle is outlined in Fig. 1c. Three agents are involved in this task. The main agent (Idea Generation Agent (IGA)) receives a task description that includes the clinical context, goal (ideas with prognostic or other relevant implications; Fig. 2a), available data, output formatting (JSON), restrictions and other pertinent information. The IGA generates ideas in predefined cycles. To make the idea generation process more systematic, we define three parameters to guide these cycles: the number of cells involved in an idea, the ‘quality’ of the idea and the number of ideas generated per cycle (Fig. 2a). After each cycle, the Idea Review Agent evaluates the generated ideas for adherence to the provided guidelines (task description), and all qualified ideas are then checked by the Idea Duplicate Detector Agent (IDDA). The IDDA compares each generated idea against every original idea already stored in the ‘Raw Idea Database’. It uses a composite similarity score, based on predefined metrics (0–10), to determine whether an idea is a duplicate and should be discarded. All ideas with a similarity score >8 against any idea in the database are declined as duplicates. All original ideas saved in the Raw Idea Database receive unique identifiers (idea_uid). Starting from cycle 2, all original idea titles from the Raw Idea Database are provided to the IGA to reduce the number of duplicates among newly generated ideas.
SPARK idea refinement pipeline
During this step, the Idea Refinement Agent takes one idea at a time from the Raw Idea Database and, using the given task description (including information on available data, coding structure of the WSI pipeline and the structure and coding nomenclature of WSI object features—that is, EvalObj, described in the next section on preprocessing), provides a detailed, step-by-step analytical and technical implementation of the parameter(s) defined in the idea. In other words, it specifies a clearly defined process from the WSI object to the parameter outputs for that WSI (Fig. 1c). All refined ideas, now containing the added technical implementation, are saved in the Refined Idea Database (JSON file) to be sampled for coding in the next step.
Preprocessing of WSI objects for SPARK
We preprocess and preload the data before passing it to the coding agentic module. These steps produce a precomputed, well-structured backbone of information that (1) reduces computation time per parameter (during coding and deployment at later stages) and (2) improves the quality and reliability of agentic coding. For each WSI, we collect and package information from three sources into a WSI object (Fig. 1f): the WSI itself, a tissue segmentation mask (tumor and tumor stroma compartments) and a GeoJSON file containing cell detection/classification information (as outlined above).
Because the coding agent (and later, parameter deployment for analyzing large cohorts) is designed to operate on smaller tumor regions (for example, 1 mm × 1 mm) and the generated parameters focus on single-cell analysis, regions and cells are the central objects processed as described below. For the standard setup (use cases 1 and 2: autonomous idea generation), in the first step, we read the information from the GeoJSON, producing a list of cell objects. Each cell object represents a single cell, with the following information initially stored in the GeoJSON: the cell polygon (coordinates in the WSI pixel coordinate system) and the cell type. We map each cell’s centroid to the tissue mask, classifying the cell as belonging to the epithelial or stromal compartment. Additionally, we attach a small image—extracted from the WSI and containing the cell nucleus—to each cell. Using the bounding boxes of the cell polygons, we create R-trees for fast spatial lookup.
For each cell, the following attributes are available and accessible for parameter coding by SPARK:
‘index’: index in the list of cells
‘cell_class’: cell type
‘polygon’: shapely polygon of the cell nucleus (pixel coordinates in the original WSI)
‘CentroidPix’: centroid of the cell nucleus (pixel coordinates in the WSI)
‘in_mask_set’: a set that describes in which tissue type(s) the cell is located (single entry: ‘Tumor’ or ‘Stroma’)
‘ImageColor’: small RGB Pillow image that contains the cell nucleus with 2 px padding around the bounding box of the cell
‘ImageGrey’: grayscale image (np.ndarray) that contains the cell nucleus (same region as ImageColor)
‘ImageMask’: numpy.ndarray with True/False values, cell nucleus mask, corresponding to polygon, ImageColor and ImageMask
In the second step, we split the WSI into square regions (‘patches’) with a side length of 1 mm. This region size was empirically chosen, based on consensus among participating expert pathologists, as the minimal spatial unit within a tumor region that could have prognostic or other clinical implications (roughly corresponding to a subclone definition, where certain features may influence clinical endpoints).
For each patch, we generate a list of cells within it, using the centroids of their polygons for unique mapping. Cells may be assigned to multiple patches if their centroid lies exactly on a boundary—this occurs in ~ 5 out of 100,000 cells and has no practical implications.
Each patch is also assigned a cropped version of the tissue mask, downscaled by a factor of 16 relative to the WSI for computational efficiency. This mask can be used to analyze the size and shape of tumor and stroma areas within the patch. If an additional comparison between the mask and the cell positions is required—such as evaluating tumor cells near the tumor–stroma boundary—the coding agent should use the provided function downscale_point_into_local_area(point_xy, patch_id). This function adjusts for both pixel offsets (entire WSI versus local/patch coordinates of the cropped image) and scaling differences.
Using the mask, we label patches as significant if they contain at least 10% tumor area; only these patches are analyzed. In the final function library, each idea is computed using the function calculate_features_fun(Eval_Obj, sign_indices, cell_ids_in_patches, patchwise_tissue_masks, output_file_NEW).
The Eval_Obj (= collection of patches) contains the Eval_Obj.single_cell_list and the function Eval_Obj.downscale_point_into_local_area(point_xy, patch_id). The three parameters—sign_indices, cell_ids_in_patches and patchwise_tissue_masks—are lists encoding patchwise information. Each patch is assigned a unique index (for reproducibility and for comparing outputs from different functions) that contains a list of cell indices mapping to the cell object list and is linked to a cropped, scaled version of the tissue mask. Our pipeline also supports larger patch sizes, which can be selected by the coding agent (for example, 5 mm or more). The coding agent can choose the patch mode best suited for the task (a fixed choice per feature) by setting patch_mode = ‘standard’ or patch_mode = ‘large’. However, all but a few features operate with 1 mm2 patches; we restrict the analysis to these.
SPARK idea/parameter coding pipeline
A single idea typically contains one or more parameters necessary for its implementation. The Idea Coding Agent (ICA) sequentially samples ideas—one idea per iteration—from the Refined Idea Database. Based on the detailed task description specifying how the Python code snippet for the parameter(s) should be structured (that is, a Python function integrated into the WSI analysis pipeline at a predefined position), the ICA generates a code snippet that executes the idea on a patch using precomputed structures. Each code snippet includes, at the end of the file, a commented section with a textual description of the parameters and the underlying measurements. For each idea, the code snippet is tested for functionality using a prepared test case (10 selected patches from a TCGA slide) as part of the WSI analysis pipeline (Fig. 1c).
If the analysis run fails due to a code error, the initial task description, the generated Python snippet and the stdout output containing the error description are provided to the Code Review Agent (CRA). The CRA may review and optimize the code up to three times (Fig. 1c) before declaring the idea as ‘failed’. If the code is functional but exceeds a runtime threshold of 60 s (for 10 test patches, only execution of idea-snippet measured), a timeout error is raised, and the snippet is passed to the CRA with the explicit task of finding a faster implementation.
SPARK parameter verification pipeline
After successful coding, parameters are ready to be implemented on the whole cohort using our developed WSI analysis pipeline. Each snippet is converted into a callable function, resulting in one comma-separated values file per WSI and idea, containing regional measurements (analyzed patches = rows) for all parameters coded for that idea. An additional agentic module processes these comma-separated values files post-analysis to perform further verification of parameters before they are used in any downstream analysis concerning clinical endpoints. This review includes both non-agentic and agentic components. The non-agentic component excludes all parameters with >80% (empirical threshold) of measurements equal to 0 or NA (missing). The agentic component reviews the idea description, the produced code snippet and the included comment describing exactly what the parameters measure and excludes parameters that are non-normalized (for example, raw cell counts without area normalization), non-numeric parameters or parameters with any significant deviations. The result of this workflow stage is two lists: excluded parameters and verified parameters. Verified parameters can then be used in further downstream analysis.
SPARK human-triggered idea generation
We developed an adaptation of SPARK for human-triggered idea generation (Fig. 1d). For this, we created an additional module consisting of two agents. The List Formatting Agent takes notes provided by human pathologists and transforms them into a more structured idea formulation. The Idea Formulation Agent converts these ideas into a structured SPARK-style JSON format, resulting in a Raw Idea Database that can then be passed through the Idea Refinement and Idea/Parameter Coding steps, similar to SPARK-generated ideas. This enables rapid implementation of ideas and deliberate exploration of cohort data by human experts, while skipping the autonomous idea generation stage.
Automatized invasion front versus tumor center detection
All human pathologists’ inputs are encoded using the above WSI preprocessing framework. The patch size remains the same as previously defined, 1 mm (patch_mode = ‘standard’). The experimental component involving human pathologists (Fig. 5a) revealed a request for separate analysis of the tumor core/center and the invasion front. To accommodate this specific request from the pathologists (expressed by P1), we labeled the patches as belonging either to the tumor core (center) or to the invasion front, using our highly detailed multiclass tissue segmentation masks, which also include benign tissue areas, following the approach described below.
Given the benign area (coherent regions at least 500,000µm2 in area, then inflated by 1,000 µ into any unlabeled/background regions) as a reference, tumor within a distance of 800 µ is labeled as an invasion front; the remainder is labeled as core. A patch is then classified as either front or core depending on which tissue label is dominant (that is, has the highest number of mask pixels within the patch). Due to the frequent intermixing of tumor and stroma, using the benign tissue as a reference proved to be the most robust and reliable way of defining these tumor layers. However, this approach naturally depends on the invasion front being captured in the WSI—which is not always the case (sometimes only the tumor core is present). In such cases, analysis of parameters involving invasion-front measurements is not possible, and therefore these cases had to be excluded.
Agentic framework and task definition
We follow a typical agent–task–crew–flow–tool paradigm within the crewAI framework. All tools are implemented as extra-agentic in the accompanying pipeline code to reduce the token usage required to accomplish the task. Task formulation for agents is crucial for proper execution by the agent. All formulations were iteratively fine-tuned during the development phase until the desired output was achieved, and all edge cases that could lead to potential failures were properly addressed. For agent descriptions and task formulations, we refer to the code (Code Availability; agents, /config/agents.yaml; tasks, /config/tasks.yaml). Empirically, agentic memory within a task or crew in our workflow setup proved to be non-beneficial, generated substantial additional costs and was therefore disabled at all stages.
LLM selection
Selection of the LLMs as specific agents was motivated by extensive initial tests during development and the following considerations. First, idea generation is the most critical stage, where substantial reasoning efforts are required. Therefore, we selected the best available LLM with strong reasoning capabilities at the time of the experiments (January–February 2025), o1 (full version) from OpenAI. Second, given the significant costs of o1 (15 € per 1 million input tokens, 60 € per 1 million output tokens), we chose a simpler model (o3-mini from OpenAI) for tasks that do not involve reasoning and require high output volumes (in tokens). This includes, for example, the Idea Review Agent and IDDA in the idea generation workflow. This choice proved to be highly effective in terms of both accuracy and cost. Third, the LLM for the ICA needed to be highly capable, given the relative complexity of the task (integration into the existing WSI analysis pipeline, handling multiple objects and their features) and the resulting parameters. For this purpose, we selected Claude Sonnet 3.5 (Anthropic) as the LLM of choice. This decision, consistent with benchmarking results on general coding tasks at the time of the experiments, was based exclusively on performance analysis through direct comparison of GPT-4o, o3-mini, o1 and Claude Sonnet 3.5 in coding 30 ideas, followed by manual review of the generated snippets. The results revealed a significantly better understanding of the task and higher-quality implementation by Claude compared to o1, whereas GPT-4o and o3-mini performed markedly worse.
Open-source LLM selection and deployment
The critical steps (idea generation and idea coding) were reproduced using open-source, local LLMs. We deploy them using either an AI server (4× NVIDIA A100 80 Gb; ‘S’) or a consumer-grade laptop (Apple MacBook Pro 64 Gb RAM, M1 Max CPU; ‘L’). We choose seven models: four generalist models (model/deployment: gpt-oss-120b/‘S’, gpt-os-20b/‘S’, distilled DeepSeekR1-Llama3-70b/‘S’ and Qwen3-32b/‘L’) and three LLMs fine-tuned on the medical domain (Meditron-70b/‘S’, MedGemma-27b/‘L’ and BioMistral-7b/‘L’). ‘S’ models were deployed using vllm v.0.11.0, and ‘L’ models were deployed using ollama v.0.12.3.
Parameter decorrelation
We performed decorrelation of SPARK-generated parameters before any downstream analysis. This process resulted in the removal of a subset of parameters that showed strong correlations with others. First, interparameter relationships were quantified at the region level (technical correlation: that is, two parameters effectively measuring the same feature) by computing pairwise Pearson correlation coefficients and associated P values for all parameters. For each parameter, the most strongly correlated counterparts were identified and tabulated together with their correlation coefficients. Second, parameters exhibiting high correlation (above a defined threshold; Results) were considered redundant. These were iteratively removed to retain a subset of minimally correlated parameters. The selection process was guided by correlation network visualizations to ensure comprehensive coverage of relevant parameters while reducing multicollinearity. This procedure resulted in highly correlated parameters forming connected clusters. From each such cluster, only one parameter was preserved and the others were discarded.
Case-level aggregation
Prognostic analysis requires case-level aggregation of single-parameter values. All generated parameters, including SPARK parameters and parameters from experiments involving human pathologists, have multiple regional measurements for a single WSI and (in many cohorts) multiple WSIs per case. For each parameter, we compute three derivatives aggregated over all measurements and all available WSIs: minimal, mean and maximal values. For use case 3 (analysis of multiplexed data), for a limited number of cases, several TMA spots were available. Parameter values were aggregated to the case level using averages.
Multiple hypothesis testing considerations
As deployment of autonomous idea generation with SPARK results in numerous parameters, we apply false discovery rate (FDR) correction using the Benjamini–Hochberg procedure, accounting for the number of tests performed, in cases where downstream analysis involves simultaneous testing of multiple parameters (for example, correlation analysis, prognostic analysis for all use cases). For Cox models, we apply separate correction for each prognostic endpoint (both categorical and continuous approaches).
Training of predictive models
An XGBoost classifier (v. 2.0) was employed for a binary classification task using pathomics features and clinical parameters (only for MSI prediction). We selected XGBoost for state-of-the-art performance on tabular data, interpretability (compared to deep learning methods), regularization and excellent handling of imbalanced datasets and error correction capabilities in consecutive decision trees (compared to the Random Forest approach). The model was implemented via the scikit-learn-compatible XGBClassifier interface, using a binary logistic regression objective and the area under the ROC curve (AUROC) as the evaluation metric.
Hyperparameter tuning was performed with Optuna, employing tree-structured Parzen estimator sampling and a fixed random seed to ensure reproducibility. The optimization process maximized fivefold stratified cross-validation ROC-AUC over 60 trials. The hyperparameter search space used reasonable defaults and was consistent across all trained models: n_estimators: 300–2,000, learning_rate: 1 × 10−3 to 0.2, max_depth: 3–10, min_child_weight: 1 × 10−2 to 10.0, subsample: 0.5–1.0, colsample_bytree: 0.5–1.0, min_split_loss (gamma): 0.0–5.0, reg_alpha: 1 × 10−8 to 10.0, reg_lambda: 1 × 10−8 to 10.0. Class imbalance was addressed by automatically computing and applying scale_pos_weight as the ratio of negative to positive samples.
Model selection employed fivefold stratified cross-validation with shuffling to ensure balanced representation of target classes across folds. For each fold, early stopping with a patience of 50 rounds was applied to prevent overfitting, monitoring validation AUC. After hyperparameter optimization, the final model was trained on the full training dataset using the optimal parameters. A validation split (20% of the training data) was generated via stratified sampling for early stopping during the final model fitting. Model performance was evaluated using ROC-AUC on independent test datasets.
Model interpretability was enhanced using SHAP values. A background dataset of 100 randomly sampled training instances was used to compute SHAP values for all test samples, and parameter importance was quantified as the mean absolute SHAP value.
RNA expression data extraction and preprocessing
We use normalized mRNA expression data for CD274 (PD-L1) for LUAD and LUSC. This data was downloaded from the TCGA portal of the Broad Institute (http://gdac.broadinstitute.org).
Prognostic analysis
During prognostic analysis, we always use one cohort per tumor type for exploration purposes (identifying relevant parameters and thresholds), and all other cohorts serve as independent testing cohorts. We implement two types of analysis for each parameter: treating the parameter as a continuous variable, and dichotomization using the optimal threshold. To determine the optimal threshold, we examine a range of quantiles for parameter values across the entire exploration cohort. We select the quantile range Q20–Q80 to reduce the likelihood of outlier-driven results due to small subgroups. Univariate and multivariate Cox analyses with FDR-adjusted P values are performed, as well as Kaplan–Meier estimates with the log-rank test, primarily for visual inspection. We test the proportional hazard assumption for each parameter in univariate (P < 0.01) and multivariate (relaxed to P < 0.05) models. All parameters failing the proportional hazard assumption were additionally tested using a spline-based Cox proportional hazard model with time-varying covariates for their prognostic significance. However, as we observed that the majority of such significant parameters were already included in classical Cox analysis (as another case-level aggregation derivate), we concentrate only on parameters that passed the proportional hazard assumption test in all experiments.
For dichotomization-based analyses, we perform additional decorrelation of parameters using the following principle: we compare each parameter against every other parameter and compute the Jaccard score for the overlap of low- and high-risk groups. We then apply the same network analysis described above to identify highly correlated clusters and retain only one parameter per cluster (Jaccard score threshold: 0.85).
We also compute the C-index for both the baseline models (including only clinicopathological variables) and all multivariate models and report the ΔC-index (difference between the two) along with the likelihood ratio test P value and calibration slopes.
Batch effect detection and automatized parameter review
Given multicentric test cohorts, we analyze parameters for potential batch effects in prognostic and temporal analysis. For every (parameter, project) combination, we generate the number of cases with data, number of cases without data, mean, standard deviation, median, unscaled median absolute deviation (MAD), 25th/50th/75th percentiles and range. To test for cross-project distributional shifts, we applied a Kruskal–Wallis test on the case-level proportions (reporting H and P) and derived the epsilon-squared effect size. Robust dispersion was summarized by the global median and global MAD across all projects, and heterogeneity was captured as (1) the maximum absolute difference between project-specific medians and (2) the same difference normalized by the global MAD. The latter was the most useful parameter. All prognostic parameters were additionally reviewed manually. For temporal evolution parameters, we used automatic filtering based on the MAD-normalized intermedian difference. At that, values 1–2 indicate possible mild batch imbalance (which can be because of cohort-level biases or mild batch effects that are inherent and inevitable to pathology images and must be tolerated well by robust parameters). Values >2 indicate a clear cohort-dependent shift and were excluded from analysis. During manual review, all the parameters excluded analyze non-normalized pixel values (colors, intensities) responsible for batch effects.
To identify all such parameters, we repurpose our parameter verification SPARK module to an additional parameter analysis module for quick parameter review at scale (available as part of the GitHub repository) and produce a list of parameters that are potentially vulnerable to batch effects and should be implemented with care (Supplementary Table 3).
Temporal evolution of the tumor, binary risk maps of SPARK parameters
Our hypothesis is that the temporal evolution of a tumor can be inferred from static data (WSIs) using SPARK parameters that effectively capture the morphological layers of subclonal evolution occurring at deeper, molecular–genetic levels (Fig. 6a). To work with such layers, we spatially map our parameters as areas of high or low risk (Fig. 6a) using the following approach. We first take all SPARK parameters associated with aggressiveness from the previous analysis (that is, those significant in univariate Cox analysis as either dichotomized or continuous variables, after decorrelation; Fig. 4). For purity, for each tumor type, we exclude all parameters potentially affected by batch effects in at least one cohort for purity (using the statistical approach described above; threshold). For each parameter (and for each tumor type separately), we determine thresholds using quantiles of parameter values for the entire cohort (Q5–Q95, with steps of 10 between Q10 and Q90). For each Q-cut-off, we then binarize the tumor area in all cohort cases into low-risk (0) and high-risk (1) regions. As our parameters include both positively associated (higher values = more aggressive) and negatively associated (higher values = less aggressive) measures of aggressiveness, we incorporate this directional information from the previous analysis (Fig. 4). In all binary tumor maps, a value of ‘1’ consistently corresponds to high risk, regardless of parameter direction. Next, for each Q-threshold, we perform univariate Cox analysis for the proportion of tumor area in the high-risk category to identify the most significant Q-threshold (based on the P value of the univariate analysis). This optimal threshold is then used to binarize all tumor cases and generate high-/low-risk maps for temporal evolution analysis (Fig. 6a). If a parameter was not measurable in the region (the reason is always that the target cell(s) are not available), we uniformly mapped it as low risk (that is, effectively leading to filtering such regions from pairwise effect analysis).
Temporal evolution of the tumor, global timing
Based on the generated binary maps of pathology features, we assume that high-risk features occupying a larger tumor area appear earlier. We classify all features as early, mid-term or late based on the number of tumors in the cohort exhibiting a specific feature in the corresponding global timing state. For finer granularity, we also define two additional categories—early-mid and mid-late—when the number of tumors in the cohort is roughly the same for the two corresponding categories (Δ < 10 tumors).
Temporal evolution of the tumor, temporal sequence analysis
The aim of this analysis is to identify pairs of parameters where parameter A precedes parameter B—that is, parameter A creates a field effect necessary for the acquisition of parameter B (Fig. 6a). To establish the temporal relationship between two or more feature layers from binary maps, we use the concepts (filters) of support, directionality, conditional necessity and sufficiency, applying them to each tumor case in the cohort. Most filter values are empirical, chosen to retain the most significant temporal associations while avoiding issues related to multiple hypothesis testing, yet without losing biological relevance.
For two parameters A and B in a tumor, there are four different states concerning overlap of low- and high-risk regions: 00, 01, 10 and 11. We set a moderately conservative support level for n11 of 30 regions within a single tumor necessary to start analysis for AB pairs.
The directionality filter is intended to identify an asymmetric relationship between parameter: that is, if A precedes B, there should be directional asymmetry for A versus B and not vice versa. We calculate directional ratios: dir_AB = n10/n01 (A without B versus B without A) and dir_BA = n01/n10. A directional relationship was established if (1) the absolute difference |n01 − n10| ≥ 12, ensuring meaningful asymmetry, and (2) either dir_AB > 5 or dir_BA > 5, indicating that one parameter occurs substantially more often without the other. The direction with the higher ratio determined the orientation (A → B or B → A) for subsequent analyses.
Conditional necessity estimates whether A is required whenever B appears: that is, when A happens, whether B usually follows. Conditional necessity was calculated as CN = n11/(n01 + n11). We set the CN filter threshold at 0.85 (if CN ≤ 0.85 ⇒ A not necessary for B; if CN > 0.85 ⇒ filter pass).
Sufficiency evaluated whether the presence of parameter A is sufficient to predict the presence of parameter B. Sufficiency was calculated as Suff = n11/(n10 + n11), representing the proportion of regions with both parameters present among all regions where parameter A is present. We exclude pairs with low sufficiency (Suff < 0.2) due to statistical (multiple hypothesis testing) and biological (uncertain effect) considerations and with high sufficiency (Suff > 0.8), suggesting strong correlation rather than temporal connection.
All pairs that survive all four filters and with sufficiency in range 0.2 ≤ Suff ≤ 0.8 are included in the temporal sequence analysis. In this, we identify chains of three parameters (A → B → C) to decipher the evolutionary sequence of events within the tumor. We consider only chains that occur in >20 tumors and those associated with tumor aggressiveness. For the latter, we use the following approach. For each of the identified chains, we calculate two types of metrics for patient subcohorts with and without this chain: (1) frequency ratio of events, pN1/pN0 and of UICC pathological Stage III–IV / Stage I–II; and (2) relative risk ratio at 36 and 60 months of event for OS, CSS and PFS endpoints. We set rather aggressive filtering to leave only the most significant chains for analysis (1) a pathological variables ratio threshold of at least 5.0 and (2) a relative risk ratio threshold of at least 2.0. We include chains in the downstream analysis if at least two of any ratios are over the threshold for all tumors except colorectal cancer, where, similar to prognostic analysis, the number of significant parameters is very high and chains with >4 significant parameters are taken into consideration.
Statistics and reproducibility
No statistical method was used to predetermine sample size (all available patients were included). No data were excluded from the analyses. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Software
Python 3.9 or later was used for most experiments. For development of agentic workflows, we used an open-source framework (crewai v.0.86.0) and the crewai-tools package (v.0.17.0), with disabled telemetry mode. For operations on GeoJSONs and processing of polygons, the following libraries were used: scipy (v.1.15.2), rasterio (v.1.4.3), shapely (v.2.1.1), rtree (v.1.4.0), py-opencv (v.4.11.0) and numpy (v.2.2.5). Seaborn (v.0.13.2) and matplotlib (v.3.9.2) were used for plotting. Lifelines (v.0.30.0) was used for Kaplan–Meier analysis. Pandas (v.2.2.3) was used for management of databases. Networkx (v.3.4.2) was used for decorrelation. Scikit-learn (v.1.6.1) was used for dataset management and metrics calculation in predictive modeling.
Hardware
Development and implementation of the SPARK agentic workflow was performed on a consumer-grade MacBook Pro (M1 Ultra, RAM 64 Gb). Analysis of cohorts (quality control, segmentation, single cell analysis, filtering, SPARK parameter generation) was performed in parallelized mode on an AI server equipped with 4× NVIDIA A100 80 Gb and using the RAMSES HPC of University Hospital Cologne (multiple NVIDIA H100 92 Gb). For our large cohort of WSIs, parameters were evaluated on the RAMSES HPC, with 120 GB and 12 CPUs reserved per case. Predictive modeling and temporal evolution analysis were performed using above-mentioned MacBook Pro.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.