Data sources
This mega-analysis was designed and carried out as a community-wide effort that sought to integrate all available rsfMRI datasets examining the acute effects of classic serotonergic psychedelics in healthy adults. Eleven independently acquired fMRI neuroimaging datasets were ultimately included (Table 1). Access to these datasets was obtained by contacting the first and senior authors of the primary publications for previous acute psychedelic neuroimaging studies.
Our two core inclusion criteria were: (1) rsfMRI acquired during the acute effects of a classic psychedelic and (2) a healthy adult participant sample. To our knowledge, no eligible datasets were excluded for reasons other than feasibility of data sharing. One site—Copenhagen University Hospital (principal investigator P. Fisher)—was unable to contribute due to restrictions related to general data protection regulation. As of our final outreach in August 2024, we were not aware of any additional eligible datasets that were excluded due to lack of contact or nonresponse.
Of the 13 datasets, 12 were DB-RCTs that included a matched placebo condition. Most employed within-subject crossover designs with counterbalanced order of drug and placebo administration. One dataset29 used a fixed-order design without a placebo condition. Table 1 summarizes the dosage, timing, and study design of each dataset. Full details on each dataset can be found in the original publications.
Brain signal preprocessing
All resting-state fMRI data underwent an identical preprocessing protocol using the fMRIprep software v.22.1.154. This standardized, uniform preprocessing protocol was centrally coordinated and executed locally in an automated fashion in each laboratory where the data originated.
For each of the BOLD runs found per subject, our analytical protocol performed the following preprocessing steps. First, a reference volume and its skull-stripped version were generated using a custom methodology of fMRIPrep. The BOLD reference was then coregistered to the T1w reference using bbregister (FreeSurfer55), which implements boundary-based registration. Coregistration was configured with six degrees of freedom. Head motion parameters with respect to the BOLD reference (transformation matrices, and six corresponding rotation and translation parameters) were estimated before any spatiotemporal filtering using McFlirt (FSL56). BOLD runs were slice-time corrected using ‘3dTshift’ from AFNI57. The BOLD timeseries were resampled onto their original, native space by applying the transforms to correct for head motion. We then resampled the ensuing voxelwise BOLD timeseries from individual subject space into standard anatomical reference space, generating preprocessed BOLD runs in the Montreal Neurological Institute (MNI) 152NLin2009cAsym 2-mm space. fMRIPrep then calculated the following confounds automatically based on the preprocessed BOLD: FD, spatial root mean square of the data after temporal differencing (DVARS)58 and the global signal (mean signal within a whole-brain gray matter (GM) mask). FD and DVARS were calculated for each functional run, both using their implementations in Nipype.
All fMRIPrep outputs were generated by each site following fMRIPrep’s standardized quality control protocols. This included verification of: (1) accurate brain extraction; (2) correct tissue segmentation into GM, white matter (WM) and cerebrospinal fluid (CSF); (3) precise coregistration between functional and anatomical images, with alignment confirmed in several planes; (4) appropriate normalization to MNI-space and (5) complete and accurate brain masking. Corresponding checks were guided by fMRIPrep-generated HTML reports and aligned with best practices from the Human Connectome Project. Each site conducted these checks before sharing preprocessed data with the central analysis team. Subjects showing poor registration, segmentation errors or mask artifacts were excluded.
To automatically remove high-motion subjects based on the obtained fMRIPrep QC outputs, we followed past work and excluded subjects which featured >15% of timepoints with FD > 0.5 (refs. 13,14,47,58). A total of six subjects were fully excluded from this study (five psilocybin, one DMT), corresponding to 12 total connectomes. An additional 29 connectomes were removed due to excessive motion at the session level (13 psilocybin, 7 LSD, 1 mescaline, 1 DMT and 7 placebo). Thus, a total of 41 connectomes were removed, resulting in a total of 519 remaining for the analyses.
Brain signal denoising
Following preprocessing, we applied four distinct pipelines to remove physiological, scanner-related and motion-related artifacts from the neuroimaging data, yielding four different sets of denoised brain signal volumes and, therefore, four sets of results. These pipelines included: (1 and 2) the anatomical CompCor (aCompCor) approach of Behzadi et al.59, with and without GSR, and (3 and 4) the ‘aggressive’ ICA-AROMA approach of Pruim et al.60, also with and without GSR. Pipelines 1 and 2 (aCompCor) are reported in the main text, whereas Pipelines 3 and 4 (ICA-AROMA) are included as a more stringent secondary approach in the Supplemental GitHub Repository at https://github.com/banilo/BOLD_psychedelics_consortium.
For the aCompCor-based pipelines, with and without GSR, a set of physiological regressors were extracted to allow for latent factor component-based noise correction59. Principal components were estimated after high-pass filtering the preprocessed BOLD timeseries using a discrete cosine filter (128-s cut-off). Probabilistic masks were generated in anatomical space for CSF and WM, and combined for an additional CSF + WM mask. Unlike the original implementation of aCompCor, which erodes the nuisance masks in BOLD space by contours of two voxels, we employed an anatomically informed refinement whereby a dilated GM mask, extracted from FreeSurfer’s ‘aseg’ segmentation, was subtracted to avoid partial volume contamination54. This GM-removed CSF + WM mask was resampled into BOLD space and binarized at a 0.99 threshold. Components were extracted separately from each of CSF, WM and CSF–WM combined masks, with the top k components retained such that their timeseries cumulatively explained 50% of the variance within each mask. The remaining components were discarded from further analysis. These aCompCor-derived regressors were then used to denoise the BOLD signal with and without GSR.
In contrast, the ICA-AROMA-based pipelines, with and without GSR, involved applying ICA as a basis to classify and remove motion-related artifacts automatically. This was performed on MNI-space BOLD volumes that were first smoothed using a 6-mm FWHM Gaussian kernel (smoothing was applied only for ICA-based noise-versus-no-noise classification, not for final analysis). The fMRI timeseries data were decomposed into spatially independent components (ICs), which were then distinguished as signal or noise by a supervised pattern-learning algorithm using three features: (1) temporal correlation with motion parameters, (2) high-frequency spectral content and (3) spatial overlap with CSF or brain edges60. ICA-AROMA includes both ‘nonaggressive’ and ‘aggressive’ variants; we implemented the aggressive variant, which regresses only the noise ICs, thereby providing a more stringent denoising procedure compared to aCompCor. In contrast, the nonaggressive approach would have retained shared variance between signal and noise components, potentially leaving residual artifacts. By employing aggressive ICA-AROMA, we aimed to maximize noise removal; thus, using this approach as a conservative approach to compare with aCompCor. ICA-AROMA was applied with and without GSR.
Following denoising, all FC matrices were assessed using a standardized procedure. Each matrix was plotted using identical color scales and thresholds to ensure consistent evaluation across participants and studies. As part of our protocol, we checked and confirmed the presence of canonical resting-state network structure—specifically that within-network connectivity was visibly higher than between-network connectivity for key neural systems such as the default mode, VIS and SMN networks. Matrices were also assessed in terms of the direction and relative magnitude of drug-induced changes and were confirmed to be consistent with those reported in the original studies for each dataset.
Regional and (sub)network parcellations
To facilitate comparability to other studies and aid neuroscientific interpretation, we extracted the denoised BOLD timeseries from cortical and subcortical regions using field-standard publicly available atlases. Cortical timeseries were extracted using the Schaefer et al.53 local–global parcellation (100 region resolution and were grouped into large-scale subnetworks based on the 17 network assignments of Yeo et al.61). Subcortical timeseries were extracted using the Tian et al.51 subcortical parcellation. The ‘S2’ parcellation was used for all nonthalamus subcortical regions (24 parcels total) and the ‘S3’ parcellation was used for the thalamus (14 parcels total), given the special interest in the latter in psychedelic research. CEREB timeseries were extracted using the Buckner et al.52 cerebellar parcellation (17 parcels in total).
FC calculation
Interregional FC was calculated as the Pearson’s product–moment correlation r between all parcels. Between-network FC corresponds to FC between parcels of different networks as defined based on the 17-network assignments of Yeo et al.61, whereas within-network FC corresponds to FC between parcels of the same network.
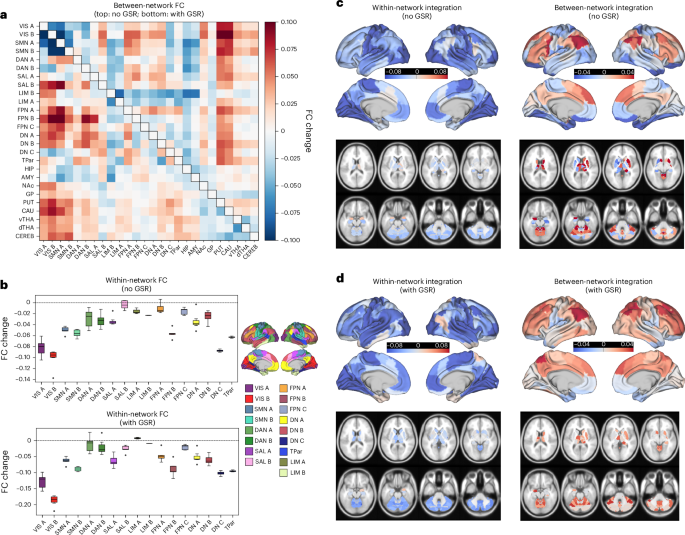
As visualized in Fig. 1b,d below, we additionally computed parcel-wise measures that we refer to as ‘within-network integration’ and ‘between-network integration’, following Siegel et al.50. Defined here, ‘within-network integration’ corresponds to the mean FC of each parcel with all parcels within its Yeo–Schaefer network, whereas ‘between-network integration’ corresponds to the mean FC of each parcel with all parcels outside of its network. In this sense, it is a variant of previous global FC or ‘global brain connectivity’ approaches (for example, refs. 20,22).
Bayesian mega-analytic framework
We carried out a Bayesian mega-analysis to fully quantify the uncertainty of the different sources of variation at play, as a natural choice of method62,63,64. This step of our analysis workflow enabled the discovery and probabilistic characterization of the common neural features underlying psychedelic drug effects as captured by human brain imaging.
Motivations and advantages
Most previous neuroscience studies on psychedelic drugs attempted to draw sharp boundaries for regional drug effects using classical null hypothesis testing and P value thresholds. To overcome these limitations, the present work adopts Bayesian principles as a formal estimation framework to quantify the degree of change in functional coupling links under the psychedelic state. In contrast to frequentist post hoc estimations, the Bayesian regime provides a direct quantification of uncertainty around model parameters by appropriately handling all considered sources of variation. This enables more careful estimation of even very subtle drug effects. As an increasing trend, Bayesian mega-analysis has been used in medicine in high-stake areas such as to re-evaluate drug effects across several RCTs65.
Bayesian hierarchical regression provides several key advantages relevant to the present aims: (1) it enables principled treatment of uncertainty by deriving full Bayesian posterior distributions for all measures, (2) it allows for continuous statements with explicitly quantified degrees of confidence (for example, ‘How sure are we that these two drugs lead to similar changes in FC?’) and (3) it does not require correction for multiple comparisons when models are independently specified, which is a key advantage as we can run one separate Bayesian model per brain feature66. More broadly, although frequentist models are designed to support binary decision-making under controlled assumptions (for example, rejecting a null hypothesis), Bayesian models are better suited for probabilistic reasoning about effect magnitudes and uncertainty. Rather than making dichotomous claims of presence or absence, our approach provides a graded, uncertainty-aware estimate of drug-related FC changes across diverse datasets. This distinction in modeling goals is especially relevant in the context of heterogeneous multisite neuroimaging data, where quantifying between-study variation is as important as identifying consistent effects.
Interpretation
Our analytical approach aimed to answer a distinct question: ‘How certain are we that a particular psychedelic drug in a certain brain region leads to a drug-induced change in functional coupling, and how strong is this effect?’ Accordingly, our models estimate the complete shape of the effect uncertainty of regional drug effects in the form of principled Bayesian posterior distributions. That is, our study did not ask ‘Is there a strict categorical difference in inter-regional or inter-network FC between drug and placebo states?’ Rather, we sought to directly quantify the population uncertainty distributions of functional coupling effects in relation to psychedelic drug influence, rather than focusing exclusively on mean FC differences. This modeling approach thus allows for probabilistic insight into the presence and strength of psychedelic effects across brain circuits, as well as their variability across drugs and studies. A fully specified generative model of the neural responses further opens the possibility to identify graded or overlapping shifts in brain network organization, consistent with the idea that drug-induced mental states may differ in degree rather than in kind67.
Model specification
The random-effects probability model with parameters that vary by study and by drug, using a subject-wise MRI-derived brain measure as outcome and the experimental condition as input variable, had the following form:
Full Bayesian model specification
Hyperpriors
$${\mathrm{Hyper}}_{\alpha }\sim \mathrm{normal}\left(\mu =0,\sigma =1\right)$$
$${\mathrm{Hyper}}_{\beta }\sim \mathrm{normal}\left(\mu =0,\sigma =1\right)$$
Priors
$${\alpha }_{\mathrm{study}\_j}\sim \mathrm{normal}\left(\mu ={\mathrm{hyper}}_{\alpha },\sigma =1\right)$$
$${\beta }_{\mathrm{drug}\_k}\sim \mathrm{normal}\left(\mu ={\mathrm{hyper}}_{\beta },\sigma =1\right)$$
Likelihood of linear model
$${y}_{\mathrm{conn\_link}}=\alpha \left[\mathrm{studies}\right]+{\beta }_{\mathrm{condition}}\left[\mathrm{drugs}\right]\times \mathrm{drug\_condition}$$
where \(\alpha\) denotes the intercept parameter corresponding to each study j out of 11 total studies (datasets), bundled by the hyperprior \({\mathrm{hyper}}_{\alpha }\), \(\beta\) denotes the slope parameter explaining the presence of the psychedelic state in connectivity differences corresponding to drug k out of the 5 overall drugs, bundled by the across-drug hyperprior \({\mathrm{hyper}}_{\beta }\), \(\mathrm{drug\_condition}\) denotes whether the observed resting-state scan was acquired during the placebo or drug-influenced state and \({y}_{\mathrm{conn\_link}}\) captures a region-region functional coupling strength, initially computed by Pearson’s correlation across the duration of a given resting-state brain scan in a particular participant, and then rescaled by Fisher z transformation (arctanh()) as outcome for this model estimation. As such, this model was inputted as many datapoints as we had resting-state scans (four-dimensional timeseries) from all 11 studies.
We approximated the model’s joint posterior parameter distribution using Markov chain Monte Carlo sampling using the PyMC3 software. After 5,000 tuning steps, the sampler had converged to the stationary distribution. Subsequently, we drew 10,000 unbiased samples from the joint posterior distribution over all parameters in the model. We relied on performance metrics that are standards in Bayesian model estimation: (1) \(\hat{R}\) to evaluate the quality of model fit for the model parameters and (2) effective sample size to index the efficiency of model convergence. A range of explanations for the brain–behavior relation between FC links and drug condition were browsed through by obtaining several plausible sets of model parameters, including their similarities and divergences, that could have generated the observed data.
For the purpose of validation of our modeling findings, we have computed three separate Markov chain Monte Carlo chains with different random initialization seeds but otherwise with the same data and same model specification. In other words, we have checked for the robustness of our model fit by rerunning the sampling process and testing whether it converges to the same final solution of posterior parameter distributions, that is, it reiterates the original modeling result.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.