In vitro experiments
Biochemical binding assay
Biochemical binding of TLC-2716 and TLC-6665 to LXRα or LXRβ was determined by time-resolved fluorescence resonance energy transfer-based biochemical assays evaluating the concentration-dependent displacement of a N-terminally biotinylated coactivator NCOA3 to recombinant glutathione S-transferase (GST)-tagged LXRα or LXRβ ligand-binding domain (LBD). Briefly, different concentrations of TLC-2716 or TLC-6665 were incubated with an assay buffer (Tris/HCl buffer (pH 6.8) containing 240 mM KCl, 1 µg µl−1 bovine serum albumin and 0.002% Triton X-100) containing 530 µg ml−1 recombinant GST-tagged LXRα/LXRβ LBD, 125 pg µl−1 GST-Tb cryptate (CisBio), 400 nM N-terminally biotinylated NCOA3 (coactivator) peptide, 1 μM 24(S)-25-epoxycholesterol (LXR agonist, only in the NCOA3 assay) and 2.5 ng µl−1 Streptavidin-XL665 (CisBio) for 1 h at 4 °C, following which fluorescence was measured in a VictorX4 multiplate reader (PerkinElmer Life Science) using 340 nm as excitation and 615 and 665 nm as emission wavelengths. Assays were performed in triplicate.
LXR GAL4 reporter transient transfection assays
LXRα and LXRβ activity status was determined via detection of interaction with coactivator and co-repressor proteins in mammalian two-hybrid experiments. HEK293 cells were transiently transfected with full length proteins of LXRα or LXRβ and their respective LBD domains under the pCMV-AD promoter(Stratagene). The cofactors, either the coactivator SRC1 or the co-repressor NCoR, were expressed as fusions to the DNA binding domain of the yeast transcription factor GAL4. Interaction was monitored via activation of a coexpressed Firefly luciferase reporter gene under control of a promoter containing repetitive GAL4 response elements (vector pFRLuc, Stratagene). Four hours after transfection, cells were incubated with a concentration range of TLC-2716 and TLC-6665 for 16 h, after which they were lysed, and luciferase activities were measured sequentially in the same cell extract using a BMG luminometer.
ABCA1 and SREBP1c luciferase reporter assays
HT-29 or HepG2 cells were stably transfected with pGL4 luciferase reporter plasmid containing the ABCA1 or SREBP1C, respectively, promoter region upstream of the Firefly luciferase reporter gene and incubated with a concentration range of TLC-2716 or TLC-6665 for 16 h as described above.
Human primary hepatocyte culture
Human primary Upcyte hepatocytes were cultivated in medium containing high concentrations of glucose (25 mM) and palmitate (100 nM) for 5 days in the presence or absence of TLC-2716, and intracellular lipids were measured by Bodipy staining.
Dysmetabolic rodent models
Ethics statement
The in vivo studies were performed at Synovo in accordance with their bioethical guidelines, which are fully compliant with ethical regulations and internationally accepted principles for the care and use of laboratory animals. Animal housing facilities were maintained at 20–22 °C with 30–50% humidity. Mice were kept on a 12-h light/12-h dark cycle with ad libitum access to food and water.
DIO mice
Eighteen-week-old male C57BL/6 DIO mice (14 weeks on HFD, Research Diets), purchased from The Jackson Laboratory, were dosed with vehicle (5% DMSO and 0.5% Kollidon-30 in 100 mM sodium phosphate buffer) or TLC-2716 (0.3 or 1 mg per kg (body weight)) once daily by oral gavage for 3 weeks.
ZDF rat
Six- to 7-week-old male obese (fa/fa) Zucker rats (ZUCKER-Leprfa, Charles River Laboratories) were fed a HFD for a total of 5 weeks and dosed with vehicle (5% DMSO and 0.5% Kollidon-30 in 100 mM sodium phosphate buffer) or TLC-2716 (0.1 or 1 mg per kg (body weight)) once daily by oral gavage for the last 2 weeks. In a second study, 6- to 7-week-old male ZDF rats (ZDF-Leprfa/Crl) were prefed a high-fat, high-cholesterol diet for 2 weeks and dosed with vehicle or TLC-6665 (5 mg per kg (body weight)) once daily by oral gavage for 4 weeks.
SD rat
Six- to 7-week-old male SD rats (Charles River Laboratories) were fed a HFD for a total of 5 weeks and dosed with vehicle (5% DMSO and 0.5% Kollidon-30 in 100 mM sodium phosphate buffer) or TLC-2716 (0.1 or 1 mg per kg (body weight)) once daily by oral gavage for the last 3 weeks.
Choline-deficient HFD rat
Six- to 8-week-old male Wistar rats (Charles River Laboratories) were fed an L-amino acid rodent diet with 60 kcal% fat with no added choline and 0.1% methionine (Research Diets, A06071302) for a total of 12 weeks. After 4 weeks of diet feeding, animals started receiving sodium nitrite injections (25 mg per kg (body weight) intraperitonially) three times a week for the remainder of the study. After 6 weeks of choline-deficient HFD feeding, animals were dosed with vehicle (5% DMSO and 0.5% Kollidon-30 in 100 mM sodium phosphate buffer) or TLC-6665 (5 mg per kg (body weight)) once daily by oral gavage for the last 6 weeks.
Humanized liver mice
Human liver chimeric PXB mice were purchased from PheonexBio, and in-life procedures were performed at InterVivo Solution in accordance with their bioethical guidelines, which are fully compliant with ethical regulations and internationally accepted principles for the care and use of laboratory animals. Briefly, animals were assigned to two groups (n = 5 male mice per group), vehicle (5% DMSO and 0.5% Kollidon-30 in 100 mM sodium phosphate buffer) or TLC-2716 (1 mg per kg (body weight) daily by oral gavage) for 8 days, after which animals were humanely killed by cardiac puncture, and plasma and tissues were collected for lipid and gene expression endpoints as described below.
Histopathology on liver tissues
Liver tissue was dipped into ice-cold PBS for 1 min and then fixed in 4% paraformaldehyde in PBS overnight for a maximum of 16 h. Afterward, samples were transferred to cold 70% ethanol/PBS to avoid prolonged fixation and embedded in paraffin before sectioning. Sections were stained with picrosirius red to visualize collagen.
Plasma parameter measurement
Plasma samples were analyzed for levels of alanine aminotransferase, aspartate aminotransferase, cholesterol and triglycerides using respons 910 (Diasys Diagnostics Systems), as per the manufacturer’s instructions.
Liver parameter measurements
Collected frozen liver samples were ground to a fine powder with a pestle and mortar under liquid nitrogen. An aliquot was subjected to lipid isolation using organic solvents (hexane and isopropanol), and, after extraction, triglyceride and cholesterol levels were quantified using commercially available kits (FUJIFILM Wako Chemicals Europe). An additional aliquot was used for collagen determination after acid hydrolysis (Total Collagen Assay, Quickzyme Biosciences).
Two-step hyperinsulinemic–euglycemic clamp
The study was performed at Physiogenex S.A.S. in accordance with ethical regulations, Guide for the Care and Use of Laboratory Animals (revised 1996 and 2011, 2010/63/EU) and French laws.
Twenty-four-week-old DIO mice (Jackson Laboratories) were fed a HFD (Research Diets, D12492) for 18 weeks and dosed with vehicle (5% DMSO and 0.5% hydroxypropyl methylcellulose in phosphate-buffered saline), TLC-6665 (5 mg per kg (body weight)) or pioglitazone (30 mg per kg (body weight)) once daily by oral gavage for 4 weeks. Animals were subjected to a two-stage hyperinsulinemic–euglycemic clamp procedure after a 6-h fast and 2 h after the last dose. Briefly, animals received a bolus of D-[3-3H]glucose (30 μCi per mouse) followed by D-[3-3H]glucose (30 μCi min−1 kg−1) infusion (2 μl min−1) for up to 210 min. Insulin was simultaneously infused at 8 mU kg−1 min−1 for the first 100 min and at 18 mU kg−1 min−1 for the last 110 min. Blood glucose was measured from the tip of the tail every 10 min by a glucometer. The glucose infusion rate was adjusted according to blood glucose levels until a first euglycemic steady state was reached (from ~70 to 100 min of infusion) and similarly adjusted until a second euglycemic steady-state was reached (from ~150 to 210 min of infusion). During both steady states, blood (5 μl) was collected frequently from the tail tip for 3H-radioactivity measurements and calculations of glucose flux.
Toxicity study
CD-1 mice
A 26-week Good Laboratory Practice (GLP) toxicology study was conducted at Inotiv according to the protocol, Inotiv’s Standard Operating Procedures and in compliance with the current US Food and Drug Administration GLP Regulations for Non-Clinical Studies (21 CFR Part 58). Animal housing facilities were maintained at 20–26 °C with 30–70% humidity. Mice were kept on a 12-h light/12-h dark cycle with ad libitum access to food and water. Briefly, the safety profile of TLC-2716 was evaluated in CD-1 mice (Charles River Laboratories) administered orally once daily at 0 (vehicle: 100 mM sodium phosphate puffer, 0.5% polyvinylpyrrolidone (PVP; pH 7.4) and 3% DMSO), 15, 60 and 120 mg per kg (body weight) per day for 26 weeks. This terminal study included comprehensive assessments of toxicokinetics and clinical laboratory parameters (hematology, clinical chemistry and urinalysis). Animals were monitored daily for clinical signs, with regular measurements of body weight, food consumption and vital signs. Necropsy and detailed tissue analysis were performed at study termination.
NHPs
A 4-week GLP toxicology study was conducted in cynomolgus monkeys (Guangzhou Xiangguan Biotech) to evaluate the safety profile of TLC-2716 administered orally once daily at 1, 5 and 15 mg per kg (body weight) per day. This terminal study included comprehensive assessments of toxicokinetics and clinical laboratory parameters (hematology, clinical chemistry and urinalysis). Animals were monitored daily for clinical signs, with regular measurements of body weight, food consumption and vital signs. Necropsy and detailed tissue analysis were performed at study termination. All procedures involving animals were reviewed and approved by the Institutional Animal Care and Use Committee and were conducted in accordance with international guidelines for the care and use of laboratory animals.
RNA isolation for real-time quantitative PCR with reverse transcription
Target organs (liver and intestine) were subjected to gene expression analysis by real-time quantitative PCR with reverse transcription (RT–qPCR). Aliquots of liver and ileum were snap frozen in liquid nitrogen and subsequently homogenized with a pestle and mortar under liquid nitrogen. Aliquots of the homogenized tissues were processed for RNA extraction and cDNA synthesis. Aliquots from the reverse-transcribed samples were used for the detection of specific mRNA transcripts of different genes of interest by RT–qPCR using commercially available mouse cDNA sequence-specific PCR primers and Taqman probes. Resulting cycle threshold (Ct) values were normalized to the housekeeping gene Tbp measured in the same sample. Results are plotted as fold change compared with vehicle controls. All primers or product codes for RT–qPCR are indicated in Supplementary Table 1.
HLO models of steatohepatitis
HLOs were generated as previously described63. Briefly, human iPS cells were differentiated into foregut progenitor cells using a published protocol44. To initiate organoid formation, foregut cells were resuspended in Matrigel at a final concentration of 750,000 cells per ml. Fifty-microliter droplets of the cell–Matrigel mixture were plated and cultured in Advanced DMEM/F12 (Gibco) supplemented with B27, N2, 10 mM HEPES, 1% GlutaMAX, 1% penicillin–streptomycin, 5 ng ml−1 FGF2, 10 ng ml−1 VEGF, 20 ng ml−1 EGF, 3 μM CHIR99021, 0.5 μM A83-01 and 50 µg ml−1 ascorbic acid for 4 days. The medium was then replaced with the same basal formulation containing 2 μM retinoic acid and cultured for an additional 4 days. Finally, the organoids were maintained in hepatocyte culture medium (Lonza) for 6 days to promote hepatic maturation. To induce a steatohepatitis-like phenotype, HLOs were gently retrieved from the Matrigel and washed with PBS. HLOs were then exposed to 300 μM sodium oleate (Sigma) in hepatocyte culture medium for 3 days using ultra-low attachment six-well plates (Corning). For quantification of intracellular lipid accumulation, sHLOs were rinsed three times with prewarmed PBS and stained with 2 μM BODIPY 493/503 (Thermo Fisher Scientific) for neutral lipid detection, along with NucBlue Live ReadyProbes (Thermo Fisher Scientific) for nuclear counterstaining. Fluorescent images were acquired using a Keyence BZ-X710 automated fluorescence microscope. Lipid droplet volume was quantified using the Hybrid Cell Count application (Keyence) and normalized to nuclear signal intensity. For RNA sequencing, total RNA was extracted from iPS cell-derived HLOs using an RNeasy Plus Mini kit (Qiagen) following the manufacturer’s protocol. RNA quality were assessed using a Fragment Analyzer (Advanced Analytical), and only samples with an RNA Quality Number of ≥8.0 were used for library preparation.
GCKR single-nucleotide polymorphism genotyping
Genotypes of GCKR variants were determined using an Illumina Infinium Global Diversity Array with the Enhanced PGx platform (Illumina) according to the manufacturer’s instructions. Genomic DNA was extracted from undifferentiated iPS cells using a QIAamp DNA Mini kit (Qiagen). Genotyping was performed at Cincinnati Children’s Hospital Medical Center Genomics Sequencing Facility using standard Infinium HTS assay protocols. Quality control steps included removal of samples with a call rate of <98%, excess heterozygosity or sex mismatch. Single-nucleotide polymorphisms (SNPs) with a call rate of <95%, Hardy–Weinberg equilibrium P of <1 × 10−6 or minor allele frequency of <1% were excluded from downstream analysis. Genomic positions were mapped to the GRCh38 (hg38) human reference genome.
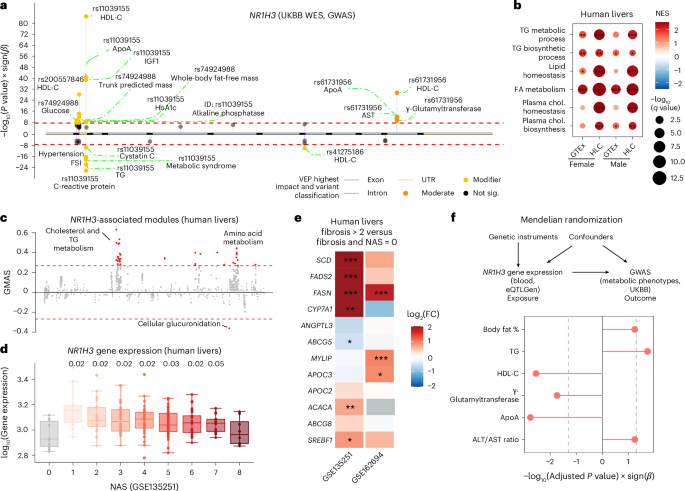
Human genetic studies
We analyzed metabolic-related phenotypic traits from the UKBB30,31 under Application Number 48020 by focusing on participants of European ancestry only (based on the UKBB return dataset 2442). A GWAS was performed using REGENIE. Step 1 involved the estimation of population structure using genotyping arrays (UKBB field 22418). Step 2 calculated genetic variant–phenotype associations based on the WES data (UKBB field 23159) using the following covariates: the first ten genetic PCs, age, sex and age–sex interaction. Gene-based testing (burden test) was performed using REGENIE step 2 by collapsing single-variant test statistics into gene-level results. Variants were grouped into functional sets defined by mask objects, where each mask represents a labeled variant category that combines different types of annotations.
Summary statistics from the FinnGen33 study were downloaded from https://www.finngen.fi/en/access_results, and those for the Million Veteran Program34 database were accessed through dbGaP under accession number phs002453.v1.p1 through the AgingX project (ID 10143). The significant (P < 5 × 10−8) associated phenotypic traits of genetic variants within candidate genes in each human population were further extracted. Fine-mapping results were downloaded from https://www.finucanelab.org/data (ref. 64).
Mendelian randomization analysis
Significant cis-eQTLs of LXRα (NR1H3) and LXRβ (NR1H2) in liver or blood were obtained from the GTEx (version 8, 208 individuals)36 or eQTLGen (31,684 individuals)65. Linkage disequilibrium clumping for SNPs was estimated using Plink (genetics.binaRies R package, window size 1 Mb, minor allele frequency ≥ 1%) on the 1000 Genomes reference panel66. Only one independent SNP was found within NR1H3 using the GTEx liver cis-eQTLs, whereas eight independent SNPs were identified using the eQTLGen blood cis-eQTLs, suggesting that the eQTLGen blood cis-eQTLs are more reliable for Mendelian randomization analysis. Therefore, significant cis-eQTLs of LXR in blood extracted from eQTLGen were applied as exposures in the Mendelian randomization analysis. GWAS summary statistics for plasma lipid-related traits were derived from WES data in the UKBB and served as outcomes. The causal effects of NR1H3 and NR1H2 gene expression on lipid traits were estimated by Inverse Variance Weighted approach using the TwoSampleMR67,68,69,70 R package (v0.6.2).
Coexpression gene set analysis
Liver gene expression datasets were obtained from GTEx36 and the Human Liver Cohort37. Correlation coefficients between each gene and LXRα (NR1H3) or LXRβ (NR1H2) were calculated by Pearson correlation. Genes were ranked by correlation coefficient, and gene sets were extracted using the msigdbr R package (version 7.5.1)71. Gene set enrichment analysis was performed using the clusterProfiler R package (4.12.6)72.
RNA-sequencing analysis
RNA was sequenced by BGI with the DNBSEQ platform. The quality of the reads was then verified using FastQC (version 0.11.9). Low-quality reads were removed, and no trimming was needed. Alignment was performed against the human genome (GRCh38, v113) following the STAR (version 2.73a) manual guidelines73. Normalized effective library sizes were calculated by trimmed mean of M values. The voom function of the Limma R package (version 3.60.0)74 was applied to transform gene counts for linear modeling with precision weights. The differential expression analysis for the transcriptome was performed using the R package Limma (version 3.60.0)74. Genes were ranked by log2 (fold change), and gene sets were extracted using the msigdbr R package (version 7.5.1)71. Gene set enrichment analysis was performed using the clusterProfiler R package (version 4.12.6)72.
Phase 1 clinical trial
Study oversight
The study protocol was approved by the Northern B Health and Disability Ethics Committee (2022 FULL 12858). This study is registered at ClinicalTrials.gov (registration: NCT05483998) and was conducted at a single site in New Zealand (Auckland Clinical Research) from 27 July 2022 to 18 June 2023, in accordance with relevant local regulatory policies. Written informed consent was obtained before enrollment.
Study design
The randomized, placebo-controlled phase 1 study included SAD and MAD cohorts. In the SAD cohorts, healthy participants were treated with single oral doses of TLC-2716 (0.5, 2, 6, 12 and 20 mg) or placebo, and in the MAD cohorts, participants received once-daily oral doses of TLC-2716 (0.5, 2, 6 and 12 mg) or placebo for 14 days. For each cohort, eight participants were randomized to receive TLC-2716, and two participants were randomized to receive placebo; study medications were administered in a fed state within 5 min of completing a standardized breakfast. In an additional MAD cohort, eight participants received TLC-2716 (6 mg) after an overnight fast to evaluate the effects of fasted versus fed dosing on the PKs of TLC-2716. Participants were confined in the study center until 72 h following the last dose of study drug, and all participants returned to the study center 14 days after the last dose of study drug for a safety follow-up visit. To maximize participant safety in SAD cohorts, safety and tolerability of the study drug through day 2 were assessed in two sentinel participants (one randomized to TLC-2716 and one randomized to placebo) per cohort before dosing of the remaining participants in each cohort. In addition, TLC-2716 dose escalation was preceded by a review of safety, PK and pharmacodynamic data by a safety review committee consisting of the principal investigator and sponsor representatives.
Inclusion and exclusion criteria
In brief, eligible study participants were healthy, nonsmoking men and women between 18 and 55 years of age and with a BMI from 19 to 35 kg m−2, inclusive at screening. All participants had an estimated glomerular filtration rate of ≥80 ml min−1, normal liver biochemistry (total bilirubin 1.0- to 1.5-fold the upper limit of normal was permitted in participants with Gilbert’s syndrome) and 12-lead electrocardiograms and screening laboratory evaluations (for example, hematology, chemistry and urinalysis) that were normal or considered to have no clinical importance by the investigator. In the MAD cohorts, an attempt was made to enroll participants with TG of ≥150 mg dl−1 and/or LDL-C of ≥130 mg dl−1 to enable preliminary assessment of the lipid-lowering benefits of TLC-2716. Key exclusion criteria included women who were pregnant or lactating, TG of ≥500 mg dl−1, LDL-C of ≥190 mg dl−1, the presence of serious active medical or psychiatric illness, excessive alcohol consumption (defined as greater than 21 units per week for men and 14 units per week for women), substance abuse or recent receipt of an investigational compound. Participants who had taken any prescription or over-the-counter medications, including herbal products, within 28 days before the start of study drug dosing, except vitamins, acetaminophen, ibuprofen and/or hormonal contraceptives, were excluded. A complete list of inclusion and exclusion criteria is available from the authors.
Safety, PK and pharmacodynamic assessments
Primary outcome
Safety and tolerability assessments, including AE monitoring, laboratory tests, physical examinations and electrocardiogram evaluations, were performed throughout the study. AEs were graded according to the Common Terminology Criteria for Adverse Events Grading Scale v5.0. Intensive PK sampling over 72 h after dosing on day 1 (in SAD and MAD cohorts) and day 14 (in MAD cohorts) was conducted, and TLC-2716 plasma concentrations were determined using a validated liquid chromatography–tandem mass spectrometry assay. PK parameters were estimated via noncompartmental methods using Pheonix WinNonlin 6.2.1 and 8.3.4 (Certara).
Secondary outcome
Plasma lipid parameters, including ApoB, were evaluated by NMR LipoProfile (Labcorp). RC was calculated as the difference between TC and the sum of direct LDL-C and HDL-C. Unless indicated, lipid parameters were measured before dosing following an overnight fast. Plasma ApoC3 and ANGPTL3 were evaluated by ELISA (360biolabs), and the impact of TLC-2716 on peripheral RCT was evaluated by assessing changes in the expression of ABCA1 and ABCG1 in peripheral blood mononuclear cells, before dosing and 4 h after dosing on day 14 (Gnomix).
Statistical analyses
Due to its exploratory nature, no formal power or sample size calculations were used to determine the sample size for this study. Empirically, the sample size was selected to adequately characterize the safety, PKs and pharmacodynamics of TLC-2716. Given the skewed distribution of plasma lipid data, we applied the Hodges–Lehmann estimator in deriving placebo-adjusted percentage change from baseline values in lipid parameters. This method estimates the median (and 95% confidence interval) of all possible pairwise differences between treatment groups and addresses the potential impact of skewed data or outliers. For analyses of changes from baseline, two-tailed Wilcoxon signed-rank tests (for comparisons within groups) and two-tailed Mann–Whitney U-tests (for comparisons between groups) were used, which were adjusted using the BH adjustment method. In this clinical trial study, sex is self-reported. Due to the small sample size of each group, sex is not considered in the analysis.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.