Human participants and ethics
This work did not involve new recruitment of human research participants. Demographics of previously recruited human participants included in the MXB project are described in ref. 11. Ethical approval was obtained from the Institutional Review Board of the National Institute of Public Health (INSP; approvals CI1479 and CB1470) for the genetic characterization of samples from the 2000 National Health Survey (ENSA 2000). Participants were enrolled through informed consent and extensive community engagement nationwide. National Health Surveys in Mexico have been conducted periodically since 1988, so the population is highly participative and receptive to household visits by INSP staff and fieldwork teams. Sampling and biobank maintenance were carried out by INSP, while genomic data were generated at the Cinvestav Research Center in Mexico. The data have been analyzed jointly, fostering interinstitutional collaboration and local leadership among Mexican researchers and trainees.
MXB dataset
To explore and catalog biomedically relevant variants within Mexican populations, we used the genetic information generated by the MXB Project11. It currently includes data for 6,011 individuals from all 32 states across Mexico recruited as part of the 2000 National Health Survey (ENSA 2000), and genotyped at 1.8 million variants on the MEGA39. The MEGA array provides extensive coverage of clinically relevant variants associated with disease and PGxs, ensuring the representation of diverse populations. Its design incorporates over 500,000 variants linked to clinical research, drawing from major variant annotation databases, including ClinVar, OMIM, the GWAS Catalog, PharmGKB, ACMG, the Clinical Pharmacogenetics Implementation Consortium and gnomAD. This comprehensive integration offers a robust framework for identifying variants with established or potential clinical significance.
Participants in the ENSA 2000 were selected using a probabilistic, multistage, stratified and clustered sampling design to ensure national representativity across 32 states of Mexico. The survey targeted civilian, noninstitutionalized individuals and collected household, health and sociodemographic data through structured interviews conducted by trained personnel. Biological samples, including serum and buffy coats, were obtained from 43,085 individuals aged 20 years or older. This cohort has been used in various epidemiological and genetic studies, offering a comprehensive resource for assessing health determinants at both the individual and population levels. The inclusion of individuals from rural and remote areas further strengthens the dataset’s utility in investigating genetic diversity and its implications for health disparities in Mexico. For more details, see ref. 40.
Samples for the MXB project were selected to maximize both geographic coverage and representation of Indigenous ancestries. The 6,011 genotyped samples are distributed across 898 recruitment sites throughout Mexico, ensuring an average sample size of five to ten individuals per locality, regardless of population density. Each state has, on average, 188 individuals, ranging from 86 to 309. The number of individuals per state is shown in Extended Data Fig. 1. The selection process prioritized individuals who reported speaking an Indigenous language (1,055), followed by random selection until budget limitations were reached. For further details, see ref. 11.
Curation of biomedically relevant variants
We focused on known variants within our dataset that are relevant to human health. To identify these variants, we used data from the following four main databases: ClinVar, GWAS Catalog, PharmGKB and OMIM12,13,14,15. The complete datasets were downloaded directly from their respective websites in March 2023. We parsed these databases to extract biomedically relevant variants, including their variant identifiers, chromosomal locations, genetic positions, effect directions, levels of evidence, clinical significance, drug associations, associated phenotypes and related genes. We then intersected these variants with the 1.8 million variants directly genotyped for the MXB cohort. For a subset of the analysis, and for the MexVar app, variants with a MAF of less than 5% were filtered out using PLINK41 with the MAF option set to 0.05, ensuring that our analysis focused on variants exhibiting meaningful frequency patterns within the Mexican population. After merging and filtering, we retained 42,769 variants. A detailed summary of these selected variants is provided in Supplementary Table 1.
Ancestry and population descriptors
‘Genetic ancestry’, as used in this study, is a statistical construct based on the genetic similarity that an individual shares with a given reference panel of source populations, reflecting their potential ancestors. In contrast, ‘race’ and ‘ethnicity’ are social constructs used to group people based on perceived physical, geographical, cultural or other social characteristics. In the analyses presented here, we exclusively refer to genetic ancestry as described above, except when mentioned otherwise. Notably, an individual’s assigned genetic ancestry is not equivalent to, and does not invalidate, how that individual self-identifies.
In this study, we distinguish between two levels of genetic ancestry: continental and subcontinental.
Continental ancestry refers to broad ancestral groupings based on large-scale worldwide population structure; this study includes Indigenous American, African and European components. These proportions were inferred using global reference panels and represent the primary axes of genetic variation relevant to populations across Mexico.
Subcontinental ancestry, on the other hand, captures the finer-scale structure within the continental component. While continental ancestry reflects large-scale population groupings, subcontinental ancestry refers to the regional genetic differentiation that arises from long-term demographic, cultural and geographic isolation within continental landmasses. For example, within a continental ancestry, such as Indigenous American, there exists substantial genetic heterogeneity between regional populations due to historical separation, founder effects and limited gene flow. Accounting for this substructure is essential for accurately characterizing patterns of genetic variation, since different subcontinental contributions can lead to significant differences in risk allele frequencies with medical implications, even among individuals with similar continental ancestry proportions.
GLMs
We used linear models to investigate the influence of genetic background (Indigenous American and African ancestry) and geographic factors on the variation observed in biomedically relevant variants. Before modeling, these variables were standardized in R to reduce scale-related biases. Subsequently, GLMs were used in R42 to analyze the associations between the standardized predictor variables and genetic variations. Coefficients and P values were calculated to determine the statistical significance of each predictor variable.
Local ancestry analysis
Local ancestry inference
To further investigate the influence of ancestry on the incidence of biomedically relevant variants in individuals within the MXB cohort, we used the Gnomix software from ref. 43 to infer local ancestry tracts. We used a k = 4 model, which assumes the presence of four distinct genetic groups. Reference populations were selected to represent the major continental genetic ancestries in Mexico11—African (Afr), European (Eur), East Asian (Eas) and Indigenous American (Ind; Supplementary Fig. 14). This approach allowed us to accurately characterize the genetic contributions from these ancestries within the cohort.
Reference populations were taken from the Human Genome Diversity Project44, the Population Architecture using Genomics and Epidemiology study45, and individuals from the MXB. We used the same number of individuals in each reference population to mitigate potential bias towards a particular ancestry. Sixty individuals were randomly chosen for each of the four populations. For the African component, we selected individuals self-identifying as Bantu, Mandenka and Yoruba; for the European, we selected populations from Western Europe, individuals self-identifying as French, Italian and Orcadians; and for East Asian, we used a combination of individuals identifying as Han and Japanese. For the Indigenous people from the Americas, we integrated genetic information from various groups, including individuals self-identifying as Mixe, Surui, Puno, Zenu and Indigenous populations in Honduras. We also included individuals from the MXB who exhibited more than 98% Indigenous ancestry based on unsupervised ADMIXTURE analysis46.
We used PLINK41 to merge the datasets and retained only the intersecting biallelic variants, excluding triallelic variants and those with genotype missingness <5%. We ran an ADMIXTURE analysis to corroborate the homogeneity of our reference panel (Supplementary Fig. 14). We ran Gnomix with the default parameters, with the exception of setting ‘inference to best’ and ‘phase’ to FALSE.
Local ancestry accuracy
We used Gnomix to estimate local ancestry tracks due to its higher accuracy compared to other programs like RFmix, as stated in the main Gnomix paper43. This paper also evaluates accuracy over time and reports that for ancestries traced back up to 20 generations, Gnomix maintains an accuracy of over 93% on array data. Given that our dataset primarily captures ancestry within the past 16 generations (assuming an average of 30 years per generation and admixture starting approximately 500 years ago), Gnomix is well-suited for our analysis.
We also conducted a simulation to evaluate the impact of local ancestry inference errors on allele frequency estimation. We simulated chromosomes with ancestry proportions reflecting those observed in the Mexican population (Afr = 4, Ind = 65, Eur = 30, Eas = 1). In this simulation, alleles were assigned frequencies specific to each ancestral background (for example, Ind = 0.1). To model errors in local ancestry prediction, we incorporated the confusion matrix derived from the Gnomix results. Using the predicted ancestry, we then estimated asF across a range of MAFs, using a sample size comparable to that of the MXB. Overall, in our simulations, we found that local ancestry inference errors had minimal impact on the estimation of asF (Supplementary Fig. 15). When training the model with our own data, we achieved a mean estimated accuracy of 94.97% across all chromosomes.
Estimation of asF
To estimate asF, we used a customized pipeline approach (Fig. 5) that involved creating a masked Variant Call Format (VCF) file. This approach uses predicted local ancestry inferences, in which variants that do not match the specified ancestry are designated missing. Subsequently, allele frequencies are computed from these ancestry-masked VCF files using VCFtools with the –freq option47. asF are then calculated as the ratio of alleles corresponding to a given ancestry over the total number of alleles for that ancestry, as detailed in the following formula:
$${\mathrm{asF}}_{x}=\frac{p\,\mathrm{alleles}\,\mathrm{in}\,x\,\mathrm{ancestry}}{\mathrm{total}\,\left(p+q\right)\,\mathrm{alleles}\,\mathrm{in}\,x\,\mathrm{ancestry}}$$
where x is the given ancestry.
asFst
To accurately quantify genetic differentiation between the states in Mexico, we calculated asFst using a customized ancestry-specific mask to filter variants for each specific ancestry. We provided a VCF file with missing alleles from other ancestries to VCFTOOLS47 using the –weir-fst-pop argument, performing this analysis separately for European and Indigenous local ancestry masks. This approach allowed us to quantify genetic differentiation between these populations accurately. asFSt values were calculated for all biomedically relevant variants with MAF > 0.05
Clinically relevant and actionable variants
To identify variants with established clinical relevance, we defined a subset referred to as the clinically relevant and actionable variants. This subset was curated to include only variants with strong evidence for medical actionability based on current clinical guidelines and expert consensus. The following criteria were applied:
-
(1)
PGx variants—we included variants annotated in the PharmGKB database at level 1A, 1B, 2A and 2B corresponding to gene–drug associations supported by clinical practice guidelines
-
(2)
Pathogenic variants in medically actionable genes—we identified all pathogenic and likely pathogenic variants without conflicting interpretations reported in the ClinVar database (accessed 30 April 2023) occurring in genes listed in the ACMG SF (v3.2; ref. 48). The guideline list includes 81 genes deemed to be medically actionable, of which 28 are associated with hereditary cancer, and the remaining 53 genes are associated with cardiovascular, metabolic and other genetic conditions for which there are available medical interventions that can prevent or reduce morbidity and mortality due to these conditions.
Notably, no allele frequency threshold was applied to this subset, as many clinically relevant variants are rare but nonetheless important for diagnosis or therapeutic decisions.
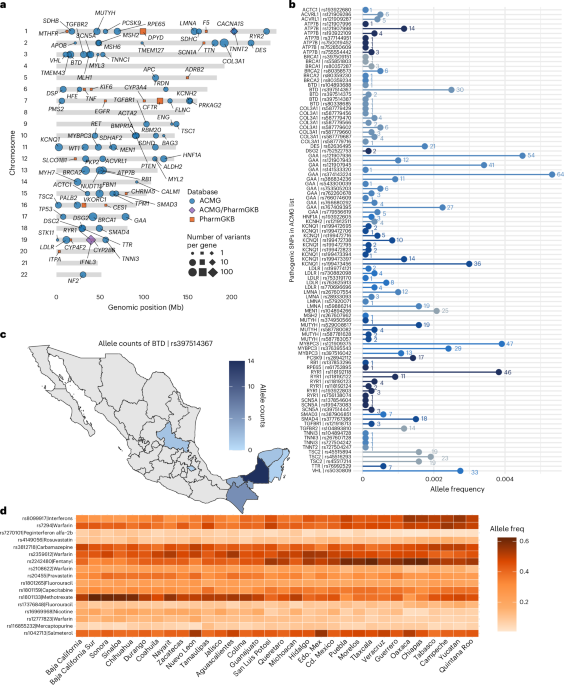
This curated subset was used to illustrate the ancestry-specific distribution of clinically actionable variants within the MXB cohort. Summary statistics—including the number of variants, genes and distribution across ancestries—are provided in Fig. 1 and Extended Data Fig. 3.
PharmGKB annotations
We obtained variant annotations from the PharmGKB, where each variant can be associated with multiple annotations. These annotations are categorized into six levels of evidence (4, 3, 2B, 2A, 1B and 1A; Supplementary Fig. 16), reflecting the strength of the evidence supporting the association with a particular drug response, with level 1A representing the highest evidence and level 4 the lowest. When a variant was associated with multiple drugs at different levels of evidence, we considered the drug with the highest level of evidence. Specifically, we focused on annotations with a level of evidence starting from 2B, which denotes variant-drug combinations supported by a moderate level of evidence and requires support from at least two independent publications.
For SNP rs4149056, we analyzed the top five drugs associated with it. The data for this analysis were retrieved from the PharmGKB database on 11 September 2023. For each drug, we also determined the number of studies supporting the association, providing insights into the robustness of the identified relationships (more details are available at https://www.pharmgkb.org/variant/PA166154579/variantAnnotation).
Integration of allele frequencies of the MXB on the Shiny app
In this paper, we introduce MexVar, a robust, user-friendly web application leveraging the Shiny framework in R49. This graphical user interface serves as a dynamic platform for querying allele frequencies nationwide in Mexico. Our analysis relied on a comprehensive dataset from the MXB, which encompasses allele frequency distributions of biomedically relevant variants across all 32 states of Mexico and comprises 42,769 variants. Detailed information regarding this dataset is provided in Supplementary Table 1. Maps displayed here and in the MexVar app are generated using mxmaps38.
The application allows users to view both genome-wide and asF, thus providing an in-depth understanding of each variant’s impact relative to the ancestry. In the ancestry selection module, users can choose from five options—‘all’ (for nonancestry-specific data) and four ancestry-specific categories (European, Indigenous, African and East Asian). Detailed methodology for ancestry-specific analysis is mentioned in ‘Local ancestry analysis’.
Two mandatory inputs for the MexVar application are the rsID of a given SNP and the desired genetic ancestry used to calculate allele frequencies. Additionally, users can customize the application’s esthetic elements, such as color schemes and titles, to suit their preferences. The application processes these inputs in real time to display an output that includes a dynamic map of Mexico, where each state is color-coded according to the selected SNP allele frequency. Moreover, users can browse an additional tab within the application to explore detailed information about the SNP, including its presence in the app, the originating database, associated phenotype, gene, risk allele (if mentioned in the database) and relevant publications. Users can also download the frequency table for offline analysis, facilitating deeper investigation and data exploration.
The design is adaptable and scalable, allowing for the integration of additional data as it becomes available, such as whole-genome sequencing. This inherent flexibility facilitates ongoing improvements and the expansion of analytical scope in line with evolving datasets.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.