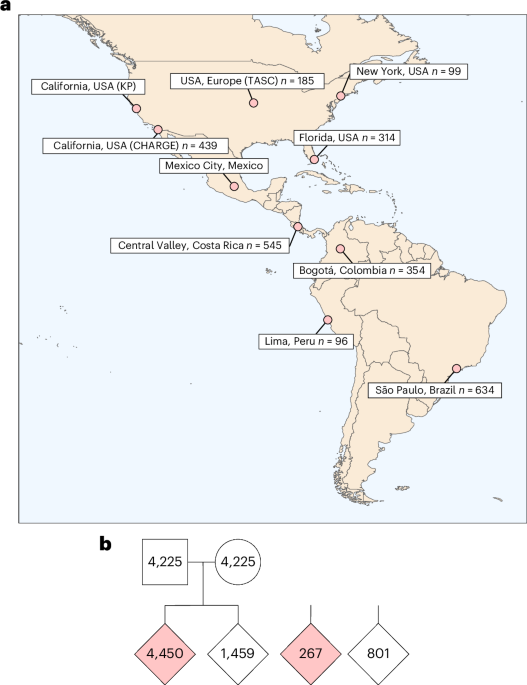
Cohort description
GALA comprises multiple sites from North, Central and South America recruiting AMR participants for studies on the genetic architecture of autism. Study procedures were approved by the institutional review board (IRB) of the Program for the Protection of Human Subjects at Mount Sinai (no. 16-01262). Informed consent was obtained from the parents or legal guardians of all study participants.
Study procedures for participant enrollment were approved by the Program for the Protection of Human Subjects at Mount Sinai (no. 16-01262 for the Seaver Center at Mount Sinai, São Paulo, Brazil, and Bogotá, Colombia; and no. 21-00039 for Peru), the University of California, Davis IRB (no. 226028-22) and the University of Miami IRB (no. 20070193). Two cohorts were collected previously: study procedures for participant enrollment in Costa Rica were approved under the guidelines of the Ministry of Health of Costa Rica, the Ethical Committee of the National Children’s Hospital in San Jose and the IRB at Mount Sinai, as described previously53,54; and The Autism Simplex Collection (TASC), which included an estimated 12% of individuals of Latin American ancestry, was recruited across 13 sites in North America and Europe, as described previously55, with local IRB oversight and all consents reviewed before depositing biospecimens and data to the National Institutes of Health repository.
For clarity, we use ‘ASD’ to refer to individuals who received a clinical diagnosis according to the procedure outlined below and ‘autism’ elsewhere. ASD diagnoses are based on expert clinical evaluations using Diagnostic and Statistical Manual of Mental Disorders, 5th Edition (DSM-5) criteria, incorporating all available data, including standardized assessments. Participants can be any age. Individuals with a known genetic condition (for example, fragile X syndrome) are excluded from analyses. Once a diagnosis of ASD is confirmed, the individual and their parents contribute a sample (blood or saliva) for genetic analyses. If both parents are not available, collection of other biological family members is encouraged (siblings, grandparents, etc.). Participating sites generally also collect additional clinical and family history information.
Description of GALA sites
New York, USA
The Seaver Autism Center for Research and Treatment at the Icahn School of Medicine at Mount Sinai, located in New York City, is the main coordinating site within the GALA Consortium. AMR individuals make up almost 30% of the population of New York City. Affected individuals undergo a full diagnostic ASD workup and receive additional assessments, including a cognitive test, adaptive behavior measure, medical checklist and behavioral checklists. Participating families receive $100 USD in compensation.
São Paulo, Brazil
The Human Genome and Stem Cell Research Center (HUG-CELL) at the Universidade de São Paulo in Brazil has over 20 years of experience in clinical and molecular research in autism, with more than 2,000 families seen. Brazil has a multiethnic admixed population, including African and Amerindian ancestry56. The HUG-CELL conducts research in human and medical genetics of rare diseases, providing genetic counseling services and genetic tests for the population. A team of psychiatrists, psychologists and neurologists completes a formal ASD diagnostic workup prior to obtaining samples for genetic testing from the individual and their family members. Financial compensation for participation is not permitted at this site; however, individuals who meet clinical criteria are offered free fragile X testing.
Bogotá, Colombia
The Centro de Investigaciones Genéticas en Enfermedades Humanas (CIGEn) at the Universidad de los Andes in Bogotá, Colombia, in close collaboration with the Instituto Colombiano del Sistema Nervioso, Clínica Montserrat, focuses on unraveling the prevalence and characteristics of autism within the Colombian population. Through ASD referrals, the impact of CIGEn extends beyond Bogotá, reaching out to other cities throughout Colombia (Medellín, Cali, Armenia, Pereira, Bucaramanga, Cartagena, Barranquilla and Santa Marta), with the aim of including families from diverse backgrounds. Financial compensation is not offered for participation.
Mexico City, Mexico
The Children’s Psychiatric Hospital ‘Juan N. Navarro’ (HPIJNN), which is part of the Psychiatric Care Services of the Mexican Government’s Ministry of Health, provides professional care for minors with mental health, psychiatric and behavioral problems. As the largest teaching center in child and adolescent psychiatry in Mexico, it performs diverse biomedical and clinical research activities. One of the main lines of research focuses on autism, in collaboration with the Genetics Department at the National Institute of Psychiatry Ramón de la Fuente Muñíz (INPRFM). The samples from Mexico are being sequenced and were not included in the current analyses. Financial compensation is not provided at this site, in accordance with ethics committee requirements.
Lima, Peru
The Centro Ann Sullivan del Perú is a non-profit center in Lima, Peru, that serves individuals with varying abilities and their families. The center specializes in helping individuals with ASD. GALA investigators from the Seaver Autism Center (M.P.T. and A.K.) traveled to Lima to perform 40 psychiatric evaluations, aid in ASD diagnostics and collect blood samples from individuals with ASD and their families. Behavioral surveys were carried out for all participants, and ASD and attention-deficit/hyperactivity disorder diagnoses were made using DSM-5 criteria. Financial compensation was not offered; instead, participating individuals received their clinical evaluation results.
California, USA (CHARGE)
The Childhood Autism Risks from Genetics and the Environment (CHARGE) cohort is a population-based case−control study collected in California at the University of California, Davis, Center for Children’s Environmental Health laboratories with the intent of addressing the impact of environmental exposures on risk57.
Florida, USA
The John P. Hussman Institute for Human Genomics at the University of Miami, located in Miami, Florida, recruits families through clinical referrals and lay organizations, providing services to families with ASD. Upwards of 70% of the Miami population identifies as AMR. The diagnostic workup included the Autism Diagnostic Interview-Revised (ADI-R) and assessment of adaptive behavior. Discrepancies between ADI-R and clinical findings were resolved using additional clinical measures, including the Autism Diagnostic Observation Schedule (ADOS).
Central Valley, Costa Rica
The founder population of the Central Valley of Costa Rica (CVCR) originated at the end of the 16th century from the intermarriage of 86 Spanish families and Indigenous Americans. The population was geographically isolated until the late 19th century; therefore, the current inhabitants are estimated to descend from fewer than 1,000 founders58. A genetic study on autism in the CVCR was initiated in 2003, and affected individuals were ascertained using the translated Spanish versions of the ADI-R and the ADOS as well as assessment of intellectual abilities and adaptive behavior53.
USA and Europe (TASC)
TASC was a collaboration among 13 sites in North America and Western Europe funded by the National Alliance for Autism Research, now Autism Speaks, and the National Institute of Mental Health. As detailed previously55, more than 1,700 individuals with ASD confirmed with extensive prospective assessment, as well as additional family members including parents, completed this study. Individuals within this study were sequenced, and those who were of AMR ancestry were included in these analyses.
California, USA (Kaiser Permanente)
The Autism Research Program (ARP) at the Kaiser Permanente Northern California (KPNC) Division of Research was established in 2002 by Senior Research Scientist Lisa Croen. The program focuses on research identifying genetic and environmental factors associated with autism and understanding patterns of detection, diagnosis and utilization of health services for individuals with ASD across the lifespan. The ARP created the Autism Family Biobank, a repository including genetic, medical and environmental information from more than 1,000 individuals with ASD and their two biological parents, who donated blood or saliva between 2015 and 2017. This collection is representative of the diverse population served by KPNC, an integrated healthcare system. The samples from Kaiser Permanente are being sequenced and were not included in the current analyses. Participants receive $15 USD per biospecimen, and families receive an additional $15 USD upon completion of the parent surveys.
Ancestry determination and sample-level quality control
Latin American samples analyzed in the current freeze include (1) GALA participants (some published in Fu et al.4); (2) non-overlapping AMR samples in the ASC and SPARK19 reported in Fu et al.4; and (3) additional AMR samples from the new release of SPARK (iWESv2). The current freeze includes trio data from 14,359 AMR samples, including 4,450 affected individuals (609 from GALA and 3,841 from ASC and the SPARK releases) and 1,459 typically developing siblings and case−control data from 267 cases and 801 controls.
To assign ancestry to each case, we followed an approach modeled after the pipeline used by gnomAD (https://gnomad.broadinstitute.org/news/2021-09-using-the-gnomad-ancestry-principal-components-analysis-loadings-and-random-forest-classifier-on-your-dataset/). Specifically, each of three jointly called datasets, derived from unpublished GALA sequencing, Fu et al. and SPARK (iWESv2), was merged with the Human Genome Diversity Project (HGDP) + 1000 Genomes Project (1KG) subset of gnomAD22, and principal component analysis (PCA) was performed in the joint dataset after they had been restricted to 5,000 ancestry-informative single-nucleotide polymorphisms59. A random forest classifier was trained on the HGDP + 1KG reference samples using the first 10 principal components and used to assign superpopulation/continental ancestry to individuals in our dataset. AMR ancestry classification was based on the predicted ancestry label assigned by the random forest model. Non-AMR cases included any individuals with ASD in ASC or SPARK releases who did not meet our criteria for genetically inferred AMR ancestry (28,818 parents, 13,030 probands and 4,749 typically developing siblings).
Hail 0.2 was used to process the SPARK (iWESv2) and unpublished GALA joint-genotyped variant call files (VCFs). Multiallelic sites were split; variants were annotated using the Variant Effect Predictor (VEP)60; and low-complexity regions (https://github.com/lh3/varcmp/blob/master/scripts/LCR-hs38.bed.gz) were removed. Hail’s pc_relate() function was used to confirm reported pedigrees and identify duplicate samples within and between datasets, which were removed. Sex was imputed using the impute_sex() function, and genotype filters were applied as described in previous methodology6 to generate working datasets (Extended Data Fig. 1).
De novo variants
Previously published de novo calls were extracted from Supplementary Table 20 from Fu et al.4. For the unpublished GALA and SPARK (iWESv2) datasets, de novo variants were called using the my_de_novo_v16() function (https://discuss.hail.is/t/de-novo-calls-on-hemizygous-x-variants/2357/19) with variant frequencies from the non-neuro subset of gnomAD exomes version 2.1.1 as priors. Potential de novo variants were dropped if they were present at a frequency greater than 0.1% within the non-neuro subset of gnomAD version 2.1.1, gnomAD version 3.1.2, in any subpopulation of these gnomAD datasets or the dataset in which they were called. Variants were further excluded if they had ‘ExcessHet’ in the Filters field, exhibited a proband allele balance < 0.3 or demonstrated a depth ratio < 0.3. Only ‘HIGH’-confidence or ‘MEDIUM’-confidence variants were kept, with the MEDIUM-confidence calls limited to a maximum allele count in the dataset of 1. A single variant per person per gene was chosen, giving preference to variants with more damaging consequences. Samples were finally excluded if the count of coding de novo variants was significantly greater than expected.
Inherited variants
Starting with the same working datasets as for de novo calling, counts of transmitted and non-transmitted alleles were generated using Hail’s transmission_disequilibrium_test() function. Variants were filtered out if they were marked ‘ExcessHet’ by GATK4 or had allele frequencies greater than 0.01% within their own dataset, within the non-neuro subset of gnomAD version 2.1.1, gnomAD version 3.1.2 or within any subpopulation of these gnomAD datasets. Variants with an allele count > 6 in the total parents of the dataset were excluded as well. Hard filtering was applied according to GATK recommendations (https://gatk.broadinstitute.org/hc/en-us/articles/360035890471-Hard-filtering-germline-short-variants). Final counts of transmitted and non-transmitted alleles were produced for PTV, MisB, MisA (1 ≤ MPC < 2) and synonymous variants.
Case−control variants
Probands within incomplete trios were identified from the ASC and GALA cohorts and matched using the top 10 principal components (‘Ancestry determination and sample-level quality control’) with non-psychiatric, unrelated controls from BioMe at a ratio of three controls to one case (3:1). Incomplete trios from SPARK (iWESv2) were removed. To ensure genome build standardization between these two cohorts, CRAM files from ASD cases were unmapped using GATK4 (ref. 61) and then remapped to a different version of the hg38 reference genome (https://biobank.ndph.ox.ac.uk/ukb/refer.cgi?id=838) using GATK3.5. Single-nucleotide variants (SNVs) and insertions/deletions (indels) were joint-genotyped across cases using the Haplotypecaller of GATK4. Like for the trio dataset processing, Hail 0.2 (https://hail.is) was used to process the joint-genotyped VCF file. The identity_by_descent() function of Hail was used to test for relatedness, which resulted in the removal of 13 cases. Sex was imputed for every sample using the impute_sex() function of Hail and cross-checked with metadata provided by all sites to ensure sample concordance.
As was done for the previous datasets4, multiallelic sites were split, variants were annotated using the VEP and low-complexity regions were removed. Variants were removed if they had an allele count ≥ 2 in the entire case−control dataset as well as an allele count ≥ 5 in the non-psychiatric subset of gnomAD version 2.1.1. Genotype calls were filtered to genotype quality > 25 and allele balance > 0.3. For case−control coverage harmonization, variants in high coverage, defined as a call rate ≥ 90%, were kept. To perform case−control matching, we excluded one case that was an outlier in the distribution of the number of synonymous variants. Finally, 267 cases were matched to 801 controls by sex and the first 10 principal components using the match_on function of the R package optmatch62.
CNV analysis
De novo CNVs called in Fu et al.4 coming from AMR samples were extracted (1,861 probands and 680 unaffected siblings). Trio and case−control datasets were analyzed separately, and GATK-gCNV24 was used to detect CNVs. First, raw CRAM files were compressed into read counts that covered the annotated exons to serve as input data. Then, a PCA-based approach that combines density and distance-based clustering was employed on the observed read counts to organize batches of samples for parallel processing. GATK-gCNV was run on cohort mode analysis for 200 samples within the cluster identified through PCA, and the remaining samples were subjected to GATK-gCNV analysis using the case mode, with models specific to the cohort (368 probands and 29 typically developing siblings). For quality control, CNV calls were processed according to Fu et al.4 methodology; CNVs were retained if they had an allele frequency < 1% that spanned more than two captured exons. For homozygous deletions, the quality score threshold was set to the lesser of 400 or 10 times the number of intervals. For heterozygous deletions, the quality score threshold was set to the lesser of 100 or 10 times the number of intervals. For duplications, the quality score threshold was set to the lesser of 50 or four times the number of intervals. For sample-level quality control, samples were retained if the number of raw, autosomal CNV calls detected by GATK-gCNV did not exceed 200 and if the number of calls with quality score ≥ 20 did not exceed 35. After quality control, 291 probands, 25 typically developing siblings, 209 cases and 735 controls remained.
A gene was considered impacted by a deletion if at least 10% of its non-redundant exons were overlapped by the deletion. For a duplication, a gene was considered impacted if at least 75% of its non-redundant exons were overlapped. Additionally, CNVs were annotated against a list of 79 curated genomic disorder loci (see Supplementary Table 10 in Fu et al.4), and a CNV call was classified as a genomic disorder CNV if it shared at least 50% reciprocal overlap with an annotated genomic disorder.
Genetic association analyses
TADA4,27 was performed for three types of inheritance classes: de novo (PTV, MisB, MisA, deletion (DEL) and duplication (DUP)), inherited (PTV, MisB and MisA) and case−control (PTV, MisB, MisA, DEL and DUP) variation. CNVs resulting from non-allelic homologous recombination (NAHR) were excluded, and only CNVs impacting fewer than nine constrained genes were retained (LOEUF < 0.6) (Supplementary Tables 13–19).
Bayes factors were constructed separately for each variant class (PTV, MisA, MisB, DEL and DUP) as described, accounting for sample size and directly using relative risk priors from Fu et al. directly (see Supplementary Table 8 in Fu et al.4). Previously published mutation rates were adjusted to align with the observed variant counts in unaffected siblings for each variant type in the dataset4.
Expected versus observed mutations in GALA
As noted in the main text, for top genes in GALA that were also significant in FuCOMP, we compared the observed numbers of variants in the GALA cohort with the expected number of variants derived from TADA analysis in FuCOMP. Although observed and expected counts may vary at the individual gene level, as expected for ultra-rare events, the overall observed and expected totals across all genes are well matched, supporting the consistency of signal with expectation (Extended Data Table 1).
Clinical genetics analyses
In addition to VarSome described in the main text, we also ran Neptune32, which uses databases of previously identified variants to call P/LP variants in a set of target genes; we took a similar approach to a recent study10 carried out in the All Of Us Research Program by focusing on 73 actionable ACMG genes63. Of the 12,162 variants in these genes among the 4,450 family-based AMR cases, Neptune provided a classification for 8,501 (69.9%); this compares to 28,262 variants among the 13,030 non-AMR family-based cases, of which 20,750 (73.4%) were classified by Neptune. In AMR participants, 136 variants were classified as P/LP, representing 1.12% (136/12,162) of all variants in these genes and 1.60% (136/8,501) of all classified variants. In non-AMR participants, 344 variants were classified as P/LP, representing 1.22% (344/28,262) of all variants and 1.66% (344/20,750) of all classified variants. Examining the results from the perspective of the participants, in AMR we observed 2.73 variants in these genes per individual, of which 1.91 per individual could be classified by Neptune, and 0.031 per individual were classified as P/LP. The corresponding numbers were 2.17 variants, 1.59 Neptune classified variants and 0.026 P/LP variants per non-AMR individual. The results show that, on the variant level, the differences in AMR versus non-AMR participants trace in part to a reduced ability of Neptune to classify non-AMR variants (Extended Data Fig. 6 and Supplementary Table 11). However, as also noted above, there are more variants per AMR participant (both total and Neptune classified), leading to an apparent lessening of impact in terms of P/LP variants per individual.
ACMG interpretation of variants
As noted above, for genetic association analyses, the TADA framework was limited to autosomal genes with available mutation rates and LOEUF scores (n = 18,128 genes) and considered only missense variants with an MPC score ≥ 1. By contrast, the clinical interpretation of variants included all autosomal or X-linked protein-truncating, synonymous and missense variants, regardless of gene annotation. In addition to applying the allele frequency cutoff of 0.1% (‘De novo variants’), X-linked variants were subjected to an allele frequency cutoff of 0.1% in the male non-psychiatric subsets of gnomAD versions 2.1.1 and 3.1.2 and their subpopulations. This resulted in 20,571 de novo variants being included for clinical genetics annotation. Inherited variant analysis was restricted to a list of well-established X-linked genes implicated in autism and/or intellectual disability (Supplementary Table 9) and subjected to the same allele frequency cutoff.
The commercially available VarSome package31 was used to evaluate the clinical impact of both de novo variants and X-linked inherited variation in the selected genes. Given the large number of variants, a batch environment was used, which limited the parameters that could be optimized for each gene. Additionally, as ACMG guidelines30 consider patient phenotype, the focus was placed on genes for which there was a reported relationship with an autism phenotype (Autism Spectrum Disorder, Autism and Autistic Behavior) and/or with a broader NDD phenotype (including the three autism terms as well as Intellectual Disability, Global Developmental Delay, Seizure, Epileptic Encephalopathy and Complex Neurodevelopmental Disorder), without knowing the full spectrum of non-autism phenotypes in the participants. Hence, the results presented here (Supplementary Tables 10–12), although based on a more transparent algorithm, should not be considered fully compliant with ACMG classification guidelines.
The api.batch_lookup function in VarSome was used to obtain germline variant-level information related to ACMG classification, nucleotide substitution and amino acid substitution, along with pathogenicity predictions. When possible, transcripts with the most severe coding impact were selected. Otherwise, the MANE Select transcript, longest canonical transcript, MANE Plus transcript, longest transcript or RefSeq transcript was chosen in that order by default.
For de novo variation, variant lists containing unique sets of variants found in each sex and zygosity were annotated. Inheritance in VarSome was set to ‘Confirmed De Novo’. Output from each list was returned in separate JSON files, which were then read into R for downstream processing into tab-separated tables. Inherited variation was examined in a similar manner; however, inheritance was set to the parent of origin of the variant.
To extend these analyses further, we used Neptune32, examining 73 ACMG actionable genes analyzed in All Of Us10. The VIP database used for annotation in Neptune was downloaded from https://gitlab.com/bcm-hgsc/neptune in VCF format, and all variants were lifted over63 from GRCh37 to GRCh38. Clinical significance annotations were parsed from the INFO field, and variants classified as Pathogenic/Likely Pathogenic, Uncertain significance and Benign/Likely Benign were noted. All rare variants in probands, regardless of mode of inheritance, were used in these analyses. Of the 73 genes, Venner et al.10 annotated only biallelic variants as P/LP in three recessive genes (MUTYH, ATP3B and KCNQ1) and only a specific variant as P/LP in HFE; we did not observe P/LP variants in these four genes, so no additional corrections were made.
Inclusion and ethics statement
This study was conducted in accordance with Nature Portfolio’s guidelines on inclusion and ethics in global research. The research was designed to include participants of diverse ancestries, with the goal of improving representation in autism genetics research. Study protocols were approved by the IRBs at all participating sites, including the Program for the Protection of Human Subjects at the Icahn School of Medicine at Mount Sinai (GCO no. 14-1082(0001)) as well as the local IRBs in Brazil, Colombia, Peru, Mexico, Kaiser Permanente and the CHARGE study (see ‘Cohort description’). Written informed consent was obtained from all participants or from parents or legal guardians where necessary. Data collection adhered to relevant ethical and cultural standards, and compensation for participation varied by site as described above. Collaborations between institutions in the United States and Latin America were established to ensure equitable contributions across sites. Local investigators in Brazil, Colombia, Mexico and Perú were involved in data collection and authorship.
Sex was recorded based on self-report at enrollment and confirmed with genetic information. Both male and female participants were included; however, sex-stratified analyses were not conducted, as the primary focus of this study was on de novo and rare variant burden across ancestry groups rather than sex differences. Participant ages varied by cohort, with probands typically enrolled during childhood or adolescence and parents as adults.
Statistics and reproducibility
All statistical analyses were performed using R (version 4.3.3), Hail (version 2.0) and Python (version 3.8). Statistical methods are described in detail in the relevant sections of Methods. Two-sided tests were used throughout unless otherwise specified. Multiple hypothesis testing was corrected using the Benjamini–Hochberg FDR procedure or Bonferroni correction as appropriate. Sample sizes were determined by the number of available participants meeting inclusion criteria in the ASC, GALA and SPARK cohorts; no statistical method was used to predetermine sample size. All available samples passing relatedness and quality control thresholds were included in the analyses. No data were otherwise excluded from the analyses.
Because this study involved secondary analysis of existing human genomic data, randomization and blinding were not applicable. The investigators were not blinded to sample status during analyses. Scripts for computational analyses performed were deposited in a GitHub repository (https://github.com/buxbaum-lab/GALA) to ensure reproducibility. Key results were independently replicated using validation datasets as described.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.