Cohorts and patients
This study was approved by the Dana-Farber/Harvard Cancer Center institutional review board (no. 21-127) in accordance with the Declaration of Helsinki. The PANGEA project is based on a cohort of patients with precursor conditions for MM identified at the DFCI for which longitudinal follow-up data, including clinical and biological variables, were collected and curated between 25 March 2021 and 21 October 2024. The primary hypothesis of the PANGEA project was an improvement of prediction accuracy compared to the previous SMM stratification models determined by the inclusion of new features (that is, dynamic biomarker) of the individual clinical profile. Among this cohort, 1,031 patients diagnosed with SMM were included as a training cohort in this study. PANGEA is a long-term cohort study at the DFCI, and all eligible patients with SMM were included for model training. To our knowledge, this is the largest cohort used for characterizing the transition from SMM to MM. Model validation is based on five independent cohorts of patients with SMM from six international centers. Cohort 1 included 380 and 105 cases from the National and Kapodistrian University of Athens (Athens, Greece) and University College London (London, UK), respectively; cohort 2 included 447 cases from the Heidelberg University Hospital (UKHD, Heidelberg, Germany); cohort 3 included 240 cases from the University of Navarra (Pamplona, Spain); cohort 4 included 67 cases from the University of Milan (Milan, Italy); and cohort 5 included 74 cases from the University Hospital of Würzburg (Würzburg, Germany). The recruitment of the training cohort and validation cohort 1 was approved by the Dana-Farber/Harvard Cancer Center institutional review board (no. 21-127). In accordance with ethical guidelines, our study was granted a waiver of informed consent by the institutional review board because the information collected on this protocol was retrospective and, thus, involved no more than minimal risk to the included patients. For validation cohort 2, written informed consent was obtained from all patients individually. Approval of the cohort recruitment was granted in ethics approval S-578/2023 by the Heidelberg ethics committee. For validation cohort 3, the study protocol, including recruitment and informed consent form, was approved by the ethics committee of the University of Navarra (no. 2017.134), and informed consent was obtained from all participants. For validation cohort 4, data were acquired within a protocol approved by the institutional review board of Milan (no. 419 on 30 August 2021), and all patients signed an informed consent. For validation cohort 5, informed consent was obtained based on a local ethics vote (no. 08/21) from Würzburg University.
Clinical annotation
For the training cohort, we collected baseline characteristics of patients at the date of diagnosis of SMM, including age, race, ethnicity and sex (self-reported), height and immunofixation isotype. We collected follow-up data with a median of two visits per year starting from the date of diagnosis of SMM until the date any of the following events occurred first: progression to active MM defined by SLiM-CRAB criteria, last follow-up visit, start of precursor treatment or death. Charts were manually reviewed by a team of expert clinical data annotators to identify any evidence of MM as defined by SLiM-CRAB criteria throughout follow-up and to ensure that any transition to MM was accurately dated. According to current standard of care, patients who met SMM criteria prior to clear SLiM-CRAB confirmation were classified and managed as SMM. In the training cohort, 114 patients (49% of 231 progressors) progressed to overt MM during follow-up based on SLiM criteria only (BMPC >60%, FLC ratio >100 with absolute involved FLC >100 mg l−1 and/or at least two magnetic resonance imaging focal lesions >5 mm). To rule out any substantial misclassification bias in our training cohort, we examined the 2-year progression rates stratified by the IMWG 20/2/20 risk category, which were as follows: 5.1% (95% CI: 3.1–7.1%), 18.6% (95% CI: 12.6–24.2%) and 41.9% (95% CI: 28.9–52.5%) for low, intermediate and high risk, respectively. These rates are similar to those reported by Mateos et al.3 in 2020 in their 20/2/20 validation study (6.2%, 17.9% and 44.2%, for low-risk, intermediate-risk and high-risk patients with SMM, respectively). Follow-up data included patient information relevant for the diagnosis and follow-up of MM and precursor conditions, including the following blood/serum values: total protein, IgA, IgM, IgG, κ and λ FLCs, sFLC ratio, calcium, creatinine, albumin, hemoglobin, lactate dehydrogenase, β-2 microglobulin and M-protein(s) concentration. Other collected variables include imaging, weight and therapy (including bisphosphonate use). Data from all BM biopsies annotated for patients during this follow-up and extracted BMPC and FISH findings, when available, were collected. FISH data were structured into one of four categories: positive, negative, not tested or unavailable. The following aberrations were captured: translocations t(4;14), t(6;14), t(11;14), t(14;16), t(14;20) and t(14;18), −17/17p deletion, 6q deletion, 11q22 deletion, 1q gain, 8q24/MYC rearrangements, −13/13q deletion, +3/+7 hyperdiploid, +9/+15 hyperdiploid, trisomy 4, trisomy 12 and trisomy 18. Study data were collected and managed using Research Electronic Data Capture (REDCap) electronic data capture tools hosted at the DFCI34,35. REDCap is a secure, web-based software platform designed to support data capture for research studies, providing (1) an intuitive interface for validated data capture; (2) audit trails for tracking data manipulation and export procedures; (3) automated export procedures for seamless data downloads to common statistical packages; and (4) procedures for data integration and interoperability with external sources.
For the validation cohorts, we extracted the targeted outcomes, time to progression, censoring or death and the biological data required by PANGEA-SMM analysis at initial and follow-up visits.
Defining dynamic/evolving biomarkers
For each of the four biomarkers in Table 2 we defined binary (0/1) dynamic variables indicating if the biomarker has increased/decreased in a way that markedly elevates the risk of progression to MM, beyond simply knowing the current biomarker value. We considered seven different candidate definitions of these binary dynamic variables and various thresholds. Candidate definitions were as follows:
-
(1)
The biomarker has increased by at least X% compared to any of the previous values in the past Y months.
-
(2)
The biomarker has increased by at least X (absolute increase) compared to any previous value in the past Y months.
-
(3)
The biomarker has increased by at least X% compared to the previous value.
-
(4)
The biomarker has increased by at least X (absolute increase) compared to the previous value.
-
(5)
The biomarker has increased by at least X (absolute increase) compared to the previous value and is at least as high as 90% of the maximum of all previous values.
-
(6)
The average change (slope, based on ordinary least squares regression) of the biomarker over the past Y months is greater than X.
-
(7)
The average change (slope) of the biomarker over the last K observations is greater than X.
The complete list of candidate thresholds (with X, Y and K values) tested can be found in Supplementary Table 12. For dynamic hemoglobin, we considered decreases (not increases) in definitions 1−7.
To determine the definition of each biomarker’s dynamic feature, we used a systematic grid search to evaluate the improvement from adding each candidate binary feature to a ‘basic’ model including only current biomarkers.
The baseline model was a multivariate Cox regression with time-varying biomarkers trained only with four biomarkers (that is, latest values of M-protein, involved/uninvolved sFLC ratio, creatinine and BMPC). Then, each candidate time-varying dynamic indicator variable was added one at a time as a predictor in the baseline model. We computed the model’s C-statistic by five-fold cross-validation (using only the training dataset). The optimal candidate dynamic definition for each biomarker was the definition that gave the greatest increase in C-statistic over the baseline model.
Training PANGEA-SMM models
The PANGEA-SMM models are two multivariate Cox regression models with time-varying predictors, namely the ‘BM’ and ‘no-BM’ models. Both include effects for three biomarkers (M-protein, log involved/uninvolved FLC ratio and log creatinine) and age as well as dynamic M-protein trend, dynamic involved/uninvolved sFLC ratio trend, dynamic creatinine trend and dynamic hemoglobin trend. Other demographic variables, including race, ethnicity and sex, were not included as input variables in the model because a previous study demonstrated that including them did not improve predictions of disease progression28. The dynamic biomarkers take values of 0 or 1 and can vary over time. The BM model also includes BMPC as a predictor, whereas the no-BM model does not and can be used when recent BMPC is not available. The dynamic variables are defined to be zero (not missing) when patient history is not available, so the models can be used without patient history (see ‘Handling of missing biomarker history’ in the Methods section for discussion of an alternative approach). The models were estimated using the survival36 package (version 3.7-0) in R37 (version 4.4.2) and output risk scores defined as the probability of progressing to MM within 2 years of the latest visit. Death was treated as a censoring event and not a competing risk, due to rare frequency in our training and validation cohorts (<5%)38.
We also assessed whether cytogenetic markers measured by FISH improve the predictions of the PANGEA-SMM BM model. Due to sample size limitations, we analyzed each FISH probe separately, adding it as a single new predictor in the PANGEA-SMM BM Cox model.
Handling of missing biomarker history
One important feature of the PANGEA-SMM models is that they are simple to use even when biomarker histories are unavailable. This is achieved by defining the trajectory variables to be zero when the biomarker histories are not available.
We considered an alternative approach to handle missing biomarker histories using a flexible framework that switches between dynamic and static submodels depending on data availability. The dynamic submodels were versions of the PANGEA-SMM models that were trained only on the subset of the training data in which all biomarker histories are observed (3,805 observations on 717 patients for the BM model; 3,912 observations on 733 patients for the no-BM model). The static submodels were simplified versions of the PANGEA-SMM models that excluded trajectory variables and were trained on the full training dataset. For each patient, risk predictions were generated from the dynamic submodels when biomarker histories were available and from the static submodels otherwise. This approach produced highly similar risk predictions to PANGEA-SMM (correlations of 0.98 for BM models and 0.97 for no-BM models) and nearly identical predictive accuracy (concordance: 0.8334 versus 0.8403 for BM models and 0.8083 versus 0.8103 for no-BM models) in the training cohort. Given these minimal differences, we chose to keep the simpler PANGEA-SMM strategy, which sets trajectory variables to zero when biomarker histories are not available.
Validating PANGEA-SMM models
We evaluated the ranking accuracy, risk stratification and calibration of the PANGEA-SMM models and rolling 20/2/20 on each validation cohort. ‘Rolling 20/2/20’ refers to the low−intermediate−high risk categories based on Lakshman et al.2, computed using the patient’s latest biomarker measurements22. We also evaluated the two PANGEA-SMM models when risk predictions use only the latest biomarker information (that is, all dynamic variables set to zero), in order to assess predictive performance when patient history is not available. Overall ranking accuracy was assessed for each model by generalized C-statistics39 including all serial observations for each patient. Dynamic ranking accuracy for each model was assessed by computing C-statistics based only on each patient’s most recent visit (and biomarker trends) at 0.1, 1, 2, 3, 4 or 5 years after baseline. C-statistics were pooled across validation cohorts using the random-effects meta-analysis technique of Debray et al.40. C-statistic differences (PANGEA-SMM BM minus rolling 20/2/20) were pooled across validation cohorts using the random-effects meta-analysis technique of Raudenbush41, with standard errors for the differences based on the cohort-specific standard errors of the C-statistics and the correlation between the PANGEA-SMM BM and 20/2/20 C-statistics estimated in the training cohort via the bootstrap.
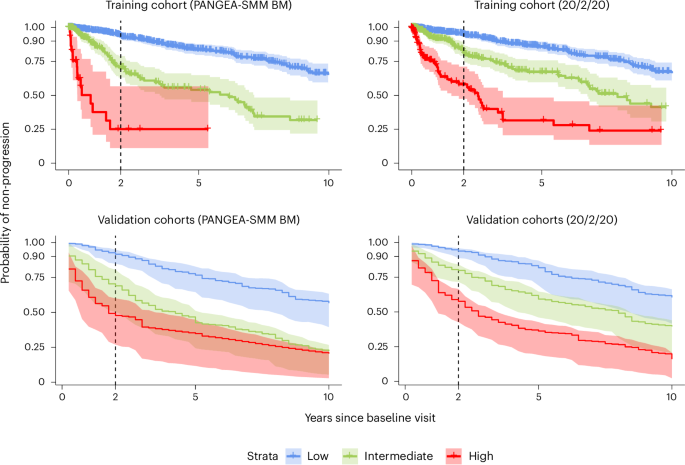
Although PANGEA-SMM produces personalized, continuous 2-year risk scores (that is, progression probabilities between 0% and 100%), we also assessed its ability to stratify patients into low-risk, intermediate-risk and high-risk groups. This stratification was based on PANGEA-SMM’s predicted risk of progression within 2 years, with ‘low’ risk patients having less than 10% predicted risk, ‘intermediate’ risk patients having between 10% and 40% predicted risk and ‘high’ risk patients having greater than 40% predicted risk. The thresholds were selected based on their clinical relevance and practical applicability while also ensuring that the resulting subgroups were sufficiently large within the training cohort to allow for meaningful analysis. These thresholds were defined prior to examining the relative proportions of the three groups in the validation cohorts, in order to preserve the integrity of the validation process. We then computed Kaplan−Meier progression curves stratified by these risk groups and compared results to 20/2/20. We pooled the progression curves for each risk group across cohorts using the random-effects meta-analysis method of Combescure et al.42 with a continuity correction of 0.05.
We also evaluated the dynamic value of PANGEA-SMM and 20/2/20 high-risk status as predictors of progression to MM within 2 years. This was done using standard inverse probability of censoring estimates of the time-dependent PPVs and NPVs43,44 based on each patient’s most recent visit at 0.1, 1, 2, 3, 4 or 5 years after baseline. The time-dependent PPVs and NPVs were pooled across cohorts using the random-effects meta-analysis method of Leeflang et al.45. The overall PPVs and NPVs and their differences (PANGEA-SMM BM minus 20/2/20) were pooled across validation cohorts using the random-effects meta-analysis technique of Raudenbush41, with standard errors based on the cohort-specific and time-specific standard errors of the predictive values and the correlation between these cohort-specific and time-specific predictive values estimated in the training cohort via the bootstrap.
Finally, we assessed calibration of the PANGEA-SMM models, which refers to the level of agreement between the predicted and observed progression rates. For each entire validation cohort and various subcohorts (stratified by low versus high biomarkers), we computed (1) the average 2-year PANGEA risk of progression and (2) the actual 2-year rate of progression (based on Kaplan−Meier analysis). We compared these progression rates to 20/2/20 (based on the reported 2-year progression rates in Mateos et al.3).
Validation results with less frequent observations
We constructed an alternative version of validation cohort 1 in which each patient has, at most, one observation per year. This was achieved by keeping only the first visit per year for each patient, starting from baseline. For example, for a patient who originally had visits at 1.2, 1.4, 2.7, 3.1 and 3.9 years after baseline, in this alternate version of the dataset we would keep only the visits at 1.2, 2.7 and 3.1 years after baseline. Time to final censoring or progression was kept the same.
Supplementary Tables 13–15 and Supplementary Figs. 1 and 2 below describe the descriptive statistics and model performance for this ‘low frequency’ version of validation cohort 1. Despite the median time between observations more than doubling (from 5.5 months to 12.8 months; Supplementary Table 13), the comparisons between PANGEA-SMM and 20/2/20 in terms of ranking accuracy, predictive value and calibration remain similar to the original results.
Open-science validation application
We developed an open-access web application to evaluate the performance of PANGEA-SMM and alternative models on our training data as well as a subset of the validation cohorts. Using the application, users can specify a dataset (DFCI or Greek/UK) and subpopulation of interest (for example, female patients at the DFCI over age 60 with sFLC ratio >20) and see the ranking accuracy, calibration and predictive value of PANGEA-SMM compared to 20/2/20. This application allows users to compare the performance of PANGEA-SMM (BM and no-BM models) with 20/2/20 in flexible and detailed populations, facilitating both decision-making about appropriate populations to use each model and future research on risk models for MM.
Clinical calculator
To easily use PANGEA-SMM in the clinic, we developed an open-access web application. The application allows entering the individual’s values (M-protein concentration, sFLC ratio, creatinine, hemoglobin and ±BMPC) along with dates of measurement. Based on the filled-in information, PANGEA-SMM automatically calculates the evolving dynamic biomarkers when past values are available and identifies the relevant model for analyzing the patient’s data (no-BM and BM). Accordingly, PANGEA-SMM determines the personalized risk of progression to MM for the patient and classifies them into groups of low, intermediate or high risk of progression to MM by comparing their personalized risk to the thresholds described above.
Statistics and reproducibility
This study was a retrospective observational analysis of longitudinal cohorts of patients with SMM assembled from participating institutions. No statistical method was used to predetermine sample size; sample sizes were determined by data availability, and the training cohort represents, to our knowledge, the largest assembled to date for modeling progression from SMM to MM. No data were excluded from the analyses beyond standard cohort eligibility criteria described in Methods.
The study involved no experimental interventions, and patients were managed according to standard of care at their respective institutions; therefore, randomization and blinding were not applicable. All statistical analyses were conducted using prespecified validation and modeling procedures, as detailed in Methods. Model development was performed in a single training cohort, and reproducibility and robustness were assessed through independent external validation across five international cohorts using predefined performance metrics.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.