Participants
We included East Asian individuals from BBJ and JCTF for the PheWAS. BBJ1, the first cohort of BBJ, recruited ~200,000 individuals with at least one of 47 target diseases from 12 Japanese medical institutions between 2003 and 2007 (refs. 16,23). BBJ2, the second cohort of BBJ, consisted of ~80,000 individuals with at least one of 38 target diseases recruited between 2013 and 2018 independently of BBJ1 (ref. 22). All participants provided written informed consent, approved by ethics committees of the Institute of Medical Sciences, the University of Tokyo and RIKEN Center for Integrative Medical Sciences. Japanese patients with COVID-19 were recruited through JCTF17. European individuals were obtained from UKB, a population-based cohort of ~500,000 individuals aged 40–69 years recruited in the UK18.
Genotyping and imputation
Japanese samples in BBJ1 were genotyped using the Illumina HumanOmniExpressExome BeadChip or a combination of the Illumina HumanOmniExpress and HumanExome BeadChips. We used Eagle (v.2.3) for haplotype phasing and Minimac3 for genotype imputation with a Japanese WGS-based reference panel. BBJ2 participants were genotyped using the Illumina Asian Screening Array-24 v.1.0, and genotypes were phased with SHAPEIT4 and imputed with Minimac4. Quality control of samples and genotypes in BBJ1 and BBJ2 was described elsewhere22. To assess the biobank-scale overview of the haplogroup distribution, we analyzed all the males in BBJ. For other analyses, we restricted to East Asian ancestry individuals defined by principal-component analysis (PCA) to adjust for the effect of population structure. UKB participants were genotyped using the Applied Biosystems UK BiLEVE Axiom Array or the Applied Biosystems UK Biobank Axiom Array. After quality control, genotypes were imputed with the Haplotype Reference Consortium data and the merged UK10K and 1KGP3 reference panels18. We analyzed individuals tagged ‘Caucasian’ in UKB Data-Field 22006 and confirmed European ancestry using PCA.
Y chromosome genotyping and imputation for East Asian people
We used GenomeStudio (v.2.0.5) to genotype MSY SNPs in BBJ. Because GenomeStudio is not optimized to genotype haploids, automated clustering can misclassify a substantial proportion of Y-chromosomal SNPs. Therefore, we manually re-clustered mis-clustered SNPs and stringently excluded those that remained mis-clustered even after manual correction. In total, we obtained 1,117 SNPs for BBJ1 and 4,358 SNPs for BBJ2.
As BBJ1 genotype data covered relatively few SNPs, major haplogroups could not be estimated with sufficient accuracy (Extended Data Fig. 1a). In detail, yhaplo v.2.1.7 (https://github.com/23andMe/yhaplo) incorrectly assigned haplogroup E (a major African haplogroup) to some individuals with haplogroup D in BBJ1. As haplogroups D and E are both classified into the broader DE lineage, individuals lacking key markers of haplogroup D were misassigned to haplogroup E (Extended Data Fig. 1b). Similarly, some haplogroup C individuals were incorrectly assigned to haplogroup F (a haplogroup common in Southeast Asian people, including Vietnamese).
To improve haplogroup estimation accuracy, we performed genotype imputation using the Japanese WGS-based reference panel. We used SHAPEIT5 for BBJ1 and SHAPEIT4 for BBJ2 for haplotype phasing. Because the MSY does not recombine, we set the recombinant rate to zero during pre-phasing. Genotype imputation was then conducted using IMPUTE2 for BBJ1 and BBJ2. Variants with imputation quality Rsq < 0.3 were excluded.
Major haplogroup estimation
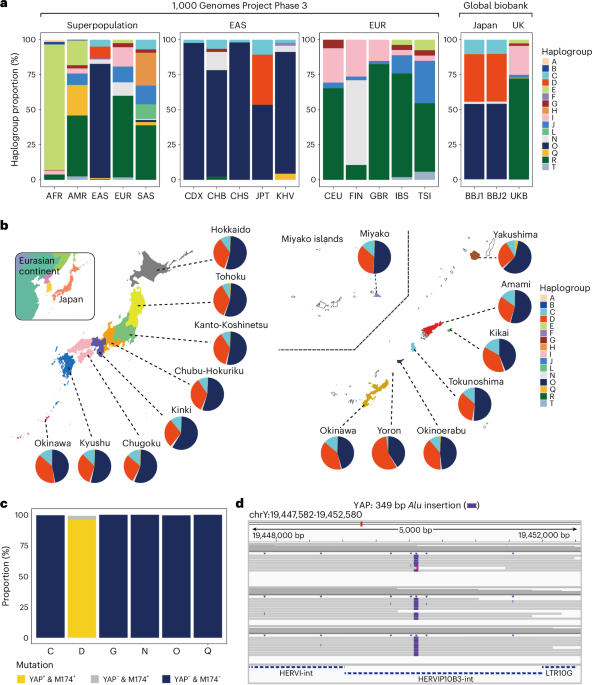
We used yhaplo to estimate Y haplogroups. We validated major haplogroup assignments by comparing them with those obtained from HaploGrouper (https://gitlab.com/bio_anth_decode/haploGrouper) and SNAPPY v.0.2.2 (https://github.com/chrisgene/snappy) (Extended Data Fig. 1c). The phylogeny trees for yhaplo and SNAPPY were based on the International Society of Genetic Genealogy’s Y-SNP trees released in 2015−2016. For HaploGrouper, we used the trees released in 2019−2020 to confirm the consistency of haplogroup estimation across different phylogeny versions. To detect the SINE specific to haplogroup D, we applied MEGAnE v.1.0.1 (https://github.com/shohei-kojima/MEGAnE) to short-read WGS data.
Long-read WGS
DNA samples were sheared to a target size of 20 kb using Megaruptor 3 (Diagenode). SMRTbell libraries were prepared to the SMRTbell Express Template Prep kit 2.0 following the manufacturer’s protocol. Fragments were size-selected using SageELF (Sage Science). Libraries were sequenced on the Sequel II (Pacific Bioscience) system using the Sequel II Binding kit 2.0 and Sequel II Sequencing kit 2.0. Circular consensus sequence (CCS) reads were generated from subreads using SMRT Link (v.9.0.0). CCS reads were aligned to hs37d5 using pbmm2 (v.1.7.0), and resulting alignments were manually inspected using the Integrative Genomics Viewer.
Detection of somatic mutations on the sex chromosomes
For detection of sex-chromosomal somatic mutations from SNP array data of BBJ, we used the MoChA pipeline27. In brief, IDAT genotype intensity data were transformed to VCF files containing log2 R ratio (LRR; total allelic intensity) and B-allele frequency (relative allelic intensity) values. After genotype phasing using SHAPEIT4, allelic imbalances were detected based on B-allele frequency values in the PAR (LOY) or whole chromosome X (LOX and GOX), followed by LRR-based evaluation to determine copy-number changes. Germline trisomy samples were removed from candidate calls. For robustness, we retained only LOX and GOX events with a cell fraction > 5% (ref. 48). For UKB, we used results from a previous analysis8 and excluded samples with events flagged as ‘unknown’.
East Asian meta-analysis of LOY GWAS
We calculated the mean LRR across MSY SNPs as a proxy for LOY28. GWAS for LOY was then conducted in each East Asian cohort using a generalized linear mixed model implemented in SAIGE v.0.43 (https://github.com/weizhouUMICH/SAIGE). We included male samples from the Hospital-based Epidemiologic Research Program at Aichi Cancer Center (HERPACC52; genotyped in the same way as BBJ2), the Japanese COVID-19 vaccine cohort (COVC53; genotyped in the same way as BBJ2) and the Japan Public Health Center-based prospective study (JPHC54; genotyped using the Illumina HumanOmniExpressExome). For TMM, we performed GWAS using regenie v.3.2.5.2 (https://github.com/rgcgithub/regenie). We included age, age2, disease status (only for BBJ), array batch (only for TMM) and the top ten genetic PCs as covariates, and used the leave-one-chromosome-out scheme to avoid proximal contamination. Variants with imputation quality Rsq < 0.3 or minor allele frequency (MAF) < 0.005 were excluded. For 9,062,001 variants, a fixed-effects meta-analysis across East Asian cohorts was conducted using the inverse-variance method implemented in METAL (released on 5 March 2020). We set the genome-wide significance threshold at P < 5.0 × 10−8 and defined genomic loci as regions within 1 Mbp of the lead variants. Our results were compared to previous reports8,39.
Phenome-wide association studies
According to ICD-10 codes, we defined cases and controls for 55 binary phenotypes in BBJ and UKB. In brief, cases were individuals affected by the target phenotypes at the time of enrollment and blood collection. From the controls, we excluded individuals with phenotypes classified into the same categories as the target phenotypes. Detailed descriptions of the binary phenotypes are provided in Supplementary Table 6.
We analyzed 35 quantitative phenotypes registered in BBJ and UKB. BBJ collected baseline clinical information through interviews and medical record reviews using a standardized questionnaire. Among the quantitative traits, age, height and weight were obtained from self-reported questionnaires, whereas laboratory measurements were retrieved from routine clinical records. We applied appropriate trait-specific transformations (z-score, z-score of log-transformed values and rank-based inverse-normal transformation)55. Detailed descriptions of trait transformations are provided in Supplementary Table 5.
We performed logistic regression (binary phenotypes) and linear regression (quantitative phenotypes) analyses of haplogroups and LOY, adjusting for age, age2, smoking and drinking status (unavailable in JCTF), and top ten autosomal genetic PCs. We additionally adjusted for BMI for phenotypes other than anthropometric traits. For the haplogroup-based PheWAS, we compared each target haplogroup with all other haplogroups in the corresponding population. For East Asians, we performed a fixed-effects meta-analysis across BBJ1 and BBJ2 using meta (v.7.0-0).
Longitudinal follow-up analysis of incident disease
We evaluated prospective associations between baseline LOY status and incident diseases occurring after recruitment in BBJ and UKB. Among the BBJ target diseases, we focused on four diseases showing significant positive associations with LOY in our PheWAS. Incident cases of T2D, dyslipidemia, asthma and COPD were identified from follow-up records. HRs and 95% CIs were estimated using Cox proportional hazards models adjusted for age, age2, BMI, drinking status and smoking status.
Replication analysis
We used 19,351 TMM samples to validate the associations between Y chromosome variation and T2D in East Asian people. TMM is a population-based prospective cohort that enrolled participants from Miyagi and Iwate Prefectures in the Tohoku region, the northeastern part of Japan19. T2D was defined based on a self-reported history of diabetes in questionnaires, excluding individuals under 40 years of age, as described elsewhere56. Genotyping was conducted using a custom SNP array for the Japanese population (Japonica Array v.2). Y haplogroups were assigned by retaining individuals whose haplogroup estimates were concordant between HaploGrouper and SNAPPY (haplogroup C, n = 1,597; D, n = 7,133; O, n = 10,440). LOY was estimated following the reported method9. We calculated the mean LRR across MSY SNPs and defined LOY dichotomously (Supplementary Table 1). GWAS was conducted using imputed genotype data, with details of imputation and quality control described elsewhere56. After imputation, variants with an imputation INFO score < 0.3 or MAF < 0.005 were excluded. Quality control of participants was performed with the following exclusion criteria: (1) outliers from East Asian ancestry clustering based on projection PCA using 1KGP3 samples; (2) genotype call rate of <95%; and (3) missing phenotype or covariate information. Leukocyte telomere length was estimated from WGS data for 3,252 TMM samples using TelSeq v.0.0.2 (https://github.com/zd1/telseq), regressed on sequencing batch and depth to correct for technical variation, and then inverse-normal transformed.
We performed replication in FinnGen, a Finnish biobank, to validate the negative LOY–T2D association in European people. Details of phenotyping, genotyping and quality control are described elsewhere29. We analyzed 137,612 males and detected LOY using the MoChA pipeline (Supplementary Table 1). Logistic regression models included age, age², BMI, smoking status and the top ten genetic PCs as covariates.
Directional causality of the association between T2D and LOY
We conducted GWAS of binary-LOY and T2D separately in male samples from BBJ1 and BBJ2, using SAIGE and including age, age2 and the top ten autosomal genetic PCs as covariates. A Two sample MR analysis was then performed using TwoSampleMR v.0.6.2 (https://github.com/MRCIEU/TwoSampleMR). As genetic instruments, we obtained genome-wide significant variants detected in the BBJ1 GWAS (T2D, 60 SNPs; LOY, 23 SNPs) to evaluate causal relationships. For associations between PRS and traits, we used PRS-CSx33 to construct PRS for T2D and LOY based on the BBJ1 GWAS results.
Haplogroup risk score construction and assessment of prediction accuracy
Given the high LD structure of the Y chromosome, we constructed haplogroup risk scores (HgRS) for T2D and height by weighting haplogroups according to the BBJ1 PheWAS results. Haplogroups with Pnominal < 0.05 were assigned their corresponding effect sizes, whereas those with Pnominal ≥ 0.05 were assigned a weight of zero.
To adjust for autosomal genetic effects on T2D and height, we prepared autosomal PRS using GWAS meta-analysis resources from the DIAGRAM Consortium21 (T2D) and the GIANT Consortium57 (height). We used PRS-CSx33 to construct a Bayesian autosomal PRS of T2D using non-palindromic SNPs with MAF > 0.01 in the DIAGRAM East Asian GWAS meta-analysis. For height, we used the East Asian PRS provided by the GIANT Consortium.
To further adjust for X chromosomal genetic effects, we constructed PRS using a clumping and thresholding method58. We performed sex-stratified GWAS of T2D and height in BBJ1 males and females using SAIGE, and meta-analyzed the male and female GWAS results with METAL. X chromosomal PRS was then constructed using a clumping and thresholding implemented in plink2 (v.2.00a3.7). We set an LD r2 threshold of 0.1 and an LD window of 250 kb for clumping, and evaluated 13 P value thresholds of PRS. For each trait, we adopted the PRS with the highest R2 among the 13 candidates (T2D, P = 1 × 10−6; height, P = 0.05).
Incorporating somatic mutations, HgRS and autosomal and X chromosomal PRS into prediction models, we performed association analyses with T2D and height, adjusting for the same covariates as in the PheWAS. We then used lmtest (v.0.9.40) to perform likelihood ratio tests between models with and without somatic mutations or HgRS to assess predictive accuracy.
Single-cell eQTL analysis of Y haplogroups
We previously constructed a single-cell eQTL resource of PBMCs, as described elsewhere14. This dataset included 146 Japanese males (haplogroup C, n = 17; D, n = 42; O, n = 83), of whom 64 were patients with COVID-19. We evaluated differentially expressed genes (DEGs) among haplogroups using a pseudobulk approach. Pseudobulk matrices were created by aggregating gene counts for each cell type (CD4+ T cells, CD8+ T cells, NK cells, B cells, monocytes and dendritic cells) within each individual. Haplogroup effects on gene expression were evaluated using a linear regression model with age, COVID-19 status, experimental reagent, LOY, top two genetic PCs and top 15 expression PCs as covariates. Genes with an expression rate >2.5% in each cell type were included in the analysis and were considered significant if they satisfied the Bonferroni-corrected threshold.
Metabolomics and proteomics analysis
We used targeted high-throughput NMR metabolomics from Nightingale Health (biomarker quantification v.2020) to measure 249 circulating lipid and metabolite biomarkers from 54,250 BBJ1 serum samples. Metabolome data were quality controlled to remove technical variation using ukbnmr v.2.2 (https://github.com/sritchie73/ukbnmr), yielding 325 markers. After excluding duplicates, we extracted independent East Asian samples (51,612 individuals) based on genotype PCA. All metabolites were inverse rank-normalized before association tests.
We measured circulating protein levels using the Olink Explore 3072 platform in 2,700 BBJ1 individuals across three batches. Normalized Protein eXpression (NPX) values were bridge-normalized using OlinkAnalyze and then inverse rank-normalized. A total of 2,071 independent East Asian individuals were extracted using genotype PCA.
We conducted linear regression analyses of metabolomic and proteomic biomarkers with haplogroups and LOY, adjusting for age, age2, smoking and drinking status, dyslipidemia treatment, and top ten genetic PCs. To account for sample overlap, LOY individuals were excluded from the T2D analysis and those with T2D from the LOY analysis. As BBJ1 NMR metabolomics data were generated in two batches (47,355 and 4,257 individuals) with different genotyping protocols, association analyses were performed per batch and combined using inverse-variance fixed-effects meta-analysis (the metagen function of the meta package).
Single-cell analyses of LOY
We integrated three scRNA-seq datasets14,35,36 and analyzed 1,899,675 PBMCs with UMI counts ≥1,500 from 840 male samples. Cells were annotated into seven major cell types (CD4+ T cells, CD8+ T cells, other T cells, NK cells, B cells, monocytes and dendritic cells) based on original annotations. LOY cells were defined as those lacking MSY expression.
For the pancreas, we analyzed two single-islet datasets: one scRNA-seq-based43 and one multiome-based44, retaining 86,027 cells with ≥1,500 UMIs from 25 male donors aged ≥30 years. For the lung, we utilized a scRNA-seq dataset including multiple respiratory diseases47. Because sex and age were not available for all samples, male-derived samples were defined as those ≥100 cells and <50% LOY cells. In total, 1,067,910 cells with ≥1,500 UMIs from 266 male-derived samples were analyzed.
For four COVID-19 samples from OASIS, we profiled single-nucleus chromatin accessibility and gene expression using the Chromium Next GEM Single Cell Multiome ATAC kit (10x Genomics). PBMCs from three asthma patients were obtained at the University of Osaka Hospital, and single-cell gene expression was profiled using the Chromium GEM-X Single Cell 5’ GEM kit (10x Genomics). All libraries were sequenced on the Illumina NovaSeqX Plus (PE150). Sequencing data were processed with Cell Ranger ARC (v.2.0.2) or Cell Ranger (v.9.0.1) using the GRCh38 reference genome. For RNA data, cells were retained if they had ≥1,500 UMIs, <99th percentile of UMI counts, ≤20% mitochondrial reads and ≤10% hemoglobin reads. For ATAC data, cells with <1,000 or >250,000 fragments or transcription start site enrichment <3 were excluded. Doublets were removed using ArchR v.1.0.3 (https://github.com/GreenleafLab/ArchR) or scds v.1.22.0 (https://github.com/kostkalab/scds), and cell types were annotated with Azimuth v.0.5.0 (https://github.com/satijalab/azimuth).
DEGs between LOY and normal cells were identified using a negative binomial model implemented in lme4 (v.1.1.32), with an offset term for UMI count and random effects for sample (only for the pancreatic analysis). Genes outside the MSY were included if they were expressed in >10% of cells. Significant DEGs were defined as those with adjusted P < 0.05 (80 downregulated DEGs in asthma monocytes) or 1.0 × 10−5 (73 downregulated DEGs in pancreatic β cells). Pathway enrichment of downregulated DEGs was evaluated using enrichGO in clusterProfiler (v.4.10.1) (OrgDb = ‘org.Hs.eg.db’; ont = ‘BP’; pvalueCutoff = 0.05; pAdjustMethod = ‘BH’).
We applied SCENIC+ (v.1.0a2)38 to investigate LOY effects on gene regulatory networks. Here, LOY cells were defined as those lacking detectable Y chromosome gene expression in both RNA and ATAC modalities. We focused on direct eRegulons (TF-target sets with motif support). Per-cell eRegulon enrichment scores were computed from both TF-target gene expression and the accessibility of associated enhancers using the score_eRegulons function (auc_threshold = 0.05; normalize = TRUE). Differences between LOY and non-LOY cells were assessed using two-sided Wilcoxon rank-sum tests at the single-cell level. For each TF, we also calculated the mean enrichment score difference (LOY – non-LOY) based on TF-target gene expression.
GTEx analysis into LOY in various tissues
To evaluate LOY in GTEx blood samples, we estimated relative Y coverage as the ratio of read coverage of the Y chromosome to that of the autosomes. We applied the DepthOfCoverage function in gatk (v.4.2.0.0) to processed WGS data41 and corrected batch effects from sequencing technology. Gene expression values (transcripts per million) were used to calculate Y expression scores42. For each sample, we computed the ratio of the mean expression of seven Y-linked genes (RPS4Y1, DDX3Y, KDM5D, USP9Y, EIF1AY, UTY and ZFY) to the expression of housekeeping genes.
Statistics and reproducibility
No statistical methods were used to predetermine sample sizes. Data distribution was assumed to be normal, but this was not formally tested. No data were excluded from analyses. Randomization and blinding were not required for the study design.
Ethics statement
Patients and control subjects in FinnGen provided informed consent for biobank research, based on the Finnish Biobank Act. Alternatively, separate research cohorts, collected before the Finnish Biobank Act came into effect (September 2013) and start of FinnGen (August 2017), were collected based on study-specific consents and later transferred to the Finnish biobanks after approval by Fimea (Finnish Medicines Agency), the National Supervisory Authority for Welfare and Health. Recruitment protocols followed the biobank protocols approved by Fimea. The Coordinating Ethics Committee of the Hospital District of Helsinki and Uusimaa (HUS) statement number for the FinnGen study is no. HUS/990/2017. The FinnGen study is approved by Finnish Institute for Health and Welfare (permit numbers THL/2031/6.02.00/2017, THL/1101/5.05.00/2017, THL/341/6.02.00/2018, THL/2222/6.02.00/2018, THL/283/6.02.00/2019, THL/1721/5.05.00/2019 and THL/1524/5.05.00/2020), Digital and Population Data Service Agency (permit numbers VRK43431/2017-3, VRK/6909/2018-3 and VRK/4415/2019-3), the Social Insurance Institution (permit numbers KELA 58/522/2017, KELA 131/522/2018, KELA 70/522/2019, KELA 98/522/2019, KELA 134/522/2019, KELA 138/522/2019, KELA 2/522/2020 and KELA 16/522/2020), Findata (permit numbers THL/2364/14.02/2020, THL/4055/14.06.00/2020,THL/3433/14.06.00/2020, THL/4432/14.06/2020, THL/5189/14.06/2020, THL/5894/14.06.00/2020, THL/6619/14.06.00/2020, THL/209/14.06.00/2021, THL/688/14.06.00/2021, THL/1284/14.06.00/2021, THL/1965/14.06.00/2021, THL/5546/14.02.00/2020, THL/2658/14.06.00/2021 and THL/4235/14.06.00/202), Statistics Finland (permit numbers TK-53-1041-17 and TK/143/07.03.00/2020 (earlier TK-53-90-20) TK/1735/07.03.00/2021 and TK/3112/07.03.00/2021) and Finnish Registry for Kidney Diseases permission/extract from the meeting minutes on 4 July 2019. The Biobank Access Decisions for FinnGen samples and data utilized in FinnGen Data Freeze 9 include: THL Biobank BB2017_55, BB2017_111, BB2018_19, BB_2018_34, BB_2018_67, BB2018_71, BB2019_7, BB2019_8, BB2019_26, BB2020_1, Finnish Red Cross Blood Service Biobank 7.12.2017, Helsinki Biobank HUS/359/2017, HUS/248/2020, Auria Biobank AB17-5154 and amendment #1 (17 August 2020), AB20-5926 and amendment #1 (23 April 2020) and its modification (22 Sep 2021), Biobank Borealis of Northern Finland_2017_1013, Biobank of Eastern Finland 1186/2018 and amendment 22 § /2020, Finnish Clinical Biobank Tampere MH0004 and amendments (21.02.2020 and 06.10.2020), Central Finland Biobank 1-2017, and Terveystalo Biobank STB 2018001 and amendment 25 August 2020. The UKBB analyses were conducted using applications 7089, 9905 and 21552.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.