Multi-omics-based modeling of obesity
We first sought to determine which molecular domains—circulating metabolome and proteome, gut metagenome and dietary intake—were most strongly associated with obesity (operationally defined as excess weight relative to height, as still widely applied) and adiposity (reflecting adipose tissue quantity and distribution) in a well-characterized, cross-sectional cohort (Impaired Glucose Tolerance and Microbiota Study (IGT-microbiota); n = 1,408; Methods, Supplementary Table 1 and Extended Data Fig. 1). This cohort, comprising at-risk individuals without established cardiovascular disease or diagnosed T2D, enables the delineation of preclinical obesity-related signatures that may generalize to populations with more advanced disease.
Using nested ridge regression with 10-fold cross-validation to optimize model regularization, we trained predictive models for BMI, waist-to-hip ratio (WHR), waist circumference and computed tomography-derived visceral and subcutaneous adipose tissue (VAT and SAT) areas. Models were constructed using individual omics layers—circulating metabolites (n = 1,190); proteins (n = 1,462); microbiome features such as gut bacterial species (metagenome-assembled genomes (MAGs) (n = 2,820)); gut microbial modules (GMMs) (n = 117); Kyoto Encyclopedia of Genes and Genomes (KEGG) orthologues (n = 11,411, corresponding to 384 pathways); and dietary variables (including dietary indices, macro-nutrient and micro-nutrient intake and food groups)—and further integrated into a combined multi-omics model (n = 5,420 variables, including metabolome, proteome, metagenome and diet).
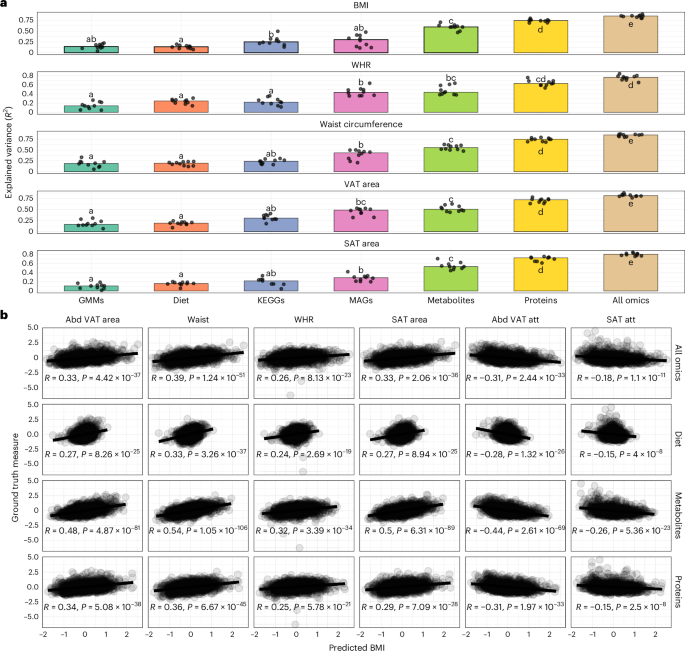
MAGs explained a similar proportion of variance in central adiposity traits (44% for waist circumference and approximately 50% for VAT area, Bonferroni-adjusted P = 1, Wilcoxon rank-sum test against metabolite-based estimates; Fig. 1a, Supplementary Table 2 and Extended Data Fig. 2a), suggesting shared links with visceral fat. However, for BMI, metabolites explained nearly twice the variance captured by MAGs (60% versus 30%, respectively; Fig. 1a and Extended Data Fig. 2b), indicating that the metabolome better represents broader obesity-related processes.
a, Proportion of variance explained (hold-out R2) for traits predicted from single omics layers: GMMs, diet, KEGG orthologues, MAGs, plasma metabolites and proteins or their combination within the IGT-microbiota cohort. Points show the per-fold R2, and bars summarize the median across ridge regression cross-validation folds (n = 10). Letters denote pairwise differences, with bars sharing a letter not differing significantly (two-sided Wilcoxon rank-sum test, Benjamini–Hochberg corrected). Exact P values are in Supplementary Table 2. b, Two-sided Pearson’s correlations between omics-predicted BMI and ground truth adiposity traits. The line represents the linear regression fit, and each point represents 1 individual with a total n = 1,408. Pearson’s r correlation coefficient and the corresponding nominal P value are shown in each panel. Abd VAT, abdominal visceral adipose tissue; Abd VAT att, abdominal visceral adipose tissue and attenuation; att, attenuation.
Consistently, the circulating metabolome provided the most physiologically informative signal for predicting obesity among individual omics layers, particularly in capturing the strongest associations with adiposity-related traits (Fig. 1a,b): metabolite-predicted BMI showed significantly stronger correlations with ground truth measures, such as waist circumference and VAT and SAT area, than BMI estimates derived from the proteome, diet or even the combined multi-omics model (Fig. 1b). These results position the metabolome as a more biologically grounded proxy of obesity-related fat accumulation.
The combined multi-omics model achieved the highest overall predictive performance (median variance explained (VEmed) 0.8 for SAT area to 0.85 for BMI; Fig. 1a and Supplementary Table 2). However, contributions across layers were not additive, reflecting overlapping molecular signals. The second-highest overall predictive performance was observed for the proteome, which explained substantial variance for several traits (for example, VEmed 0.74 for BMI and waist and 0.71–0.74 for VAT and SAT; Fig. 1a and Supplementary Table 2). Nonetheless, its performance did not significantly exceed that of the metabolome for several traits (for example, SAT area; Bonferroni-adjusted P = 0.3), and the associations with central adiposity traits were less pronounced (Fig. 1b). This observation is further supported by recent intervention data, where proteome-predicted BMI remained stable despite reductions in BMI, metabolite-predicted BMI and improvements in metabolic health, suggesting proteome stability at the expense of metabolic responsiveness to intervention10.
Finally, inter-omic comparisons highlighted the broader integrative capacity of the metabolome: metabolites explained up to 76% of the variance of individual proteins (median 35%). In comparison, proteins explained up to 74% of individual metabolites with a similar median of 34% (Extended Data Fig. 2c and Supplementary Table 3). Microbiome gene richness was best explained by metabolites, with a median variance of 61%, compared to 44% for proteins (Extended Data Fig. 2d). Similarly, metabolites outperformed proteins in explaining individual species abundances, reaching a maximum of 82% variance explained for specific MAGs versus a maximum of 51% for proteins (Extended Data Fig. 2e). However, the VEmed for MAGs was similar for both metabolites and proteins (22% and 24%, respectively; Extended Data Fig. 2e).
These results underscore strong covariance across omics layers and highlight the metabolome’s central role as a clinically relevant integrator of host, microbial and dietary signals.
Uncoupling the obesogenic signature from BMI
To improve the parsimony of the model while addressing colinearity, we trained a ridge regression model using the 267 metabolites most stringently associated with BMI (Methods and Supplementary Table 4). The resulting metBMI was highly correlated with the measured BMI (Fig. 2a; Pearson’s r = 0.62, Spearman’s ρ = 0.63, P < 2.2 × 10−16), explaining 39% of BMI variance in the held-out test set of the IGT-microbiota cohort (Extended Data Fig. 3a). Similar results were obtained using least absolute shrinkage and selection operator (LASSO) regression (Methods).
a, Two-sided Pearson’s correlation between ground truth BMI and metBMI (n = 1,408). Each dot represents one individual, colored by metBMI group (sample size per group as described in the legend). Pearson’s coefficient (r) and the corresponding P value are shown. b, Principal component analysis (PCA) of whole plasma metabolome. Each point represents one individual, colored by metBMI group. Large points denote group medoids. Side box plots display metBMI group distributions along PC1 and PC2 (two-sided Kruskal–Wallis derived, n = 1,408 and per metBMI group as described in the top legend; n for normal weight = 313, overweight = 487, obesity = 307, LmetBMI = 147, HmetBMI = 154). Box plots display the median; interquartile range (IQR) with whiskers specify ±1.5× IQR; and plotted points denote outliers. c, Comparisons of z-score-transformed anthropometric, metabolic and lifestyle features across metBMI groups (two-sided Kruskal–Wallis tests with Benjamini–Hochberg adjustment). VAT attenuation is shown as absolute values. n per group and box plot as in b. oGTT, oral glucose tolerance test; FINDRISC, Finnish Diabetes Risk Score; PC, principal component.
To capture the metabolic signature of obesity across the BMI spectrum, we extracted metBMI residuals for each participant, adjusted for age, sex and BMI. Individuals with disproportionately high (> +2.5) or low (< −2.5) residuals were classified as HmetBMI and LmetBMI, respectively, each representing approximately 10% of the cohort. These groups exhibited distinct metabolomic profiles (P = 1.2 × 10−7, post hoc Wilcoxon rank-sum test; Fig. 2b). LmetBMI individuals clustered with those of normal weight, whereas HmetBMI individuals clustered with those with obesity, despite similar BMI ranges (range, LmetBMI: 18.98–46.27 kg m−2, HmetBMI: 20.59–39.92 kg m−2, P = 0.28, Wilcoxon rank-sum-test; Extended Data Fig. 3b–d) and similar broad clinical characteristics (for example, age, sex, fasting glucose and blood pressure; Fig. 2c).
HmetBMI individuals exhibited hallmarks of metabolic dysfunction, including higher WHR, more severe VAT area and attenuation, elevated triglycerides, insulin resistance (Homeostatic Model Assessment of Insulin Resistance (HOMA-IR)), inflammation (C-reactive protein (CRP)), poorer adherence to an anti-inflammatory diet (Anti-Inflammatory Diet Index (AIDI))19 and reduced gut microbiome gene richness compared to LmetBMI (Fig. 2c and Supplementary Table 5). These patterns were consistent across sex and BMI class, highlighting that metBMI captures metabolic risk independent of body size (Supplementary Tables 5 and 6).
Some differences between the HmetBMI and LmetBMI, however, were sex specific: lower physical activity was more pronounced in males, and elevated inflammation and poor adherence to an anti-inflammatory diet were more evident in females (Supplementary Table 6), despite balanced model training and the independence of metBMI residuals from BMI and sex (Methods). Crucially, key discriminators, such as lower gut microbiome gene richness, more pronounced VAT attenuation, insulin resistance and insulin hypersecretion, were consistently observed in HmetBMI across both sexes and BMI classes (Supplementary Tables 5 and 6), emphasizing the unique contribution of hyperinsulinemia, insulin resistance and impaired glucose uptake/utilization in metabolic obesity beyond actual BMI.
These findings were replicated in the independent Swedish Cardiopulmonary Bioimage Study (SCAPIS) cohort (n = 466; Supplementary Table 7), where metBMI and BMI remained strongly correlated (r = 0.72, ρ = 0.71, P < 2.2 × 10−16, out-of-sample R2 = 0.52; Extended Data Fig. 4a,b). This cohort had a more balanced sex distribution but was slightly older and showed higher disease burden than the IGT-microbiota cohort. Notably, it included a three-fold higher prevalence of metabolic syndrome, 11% with newly diagnosed T2D at screening and more severe dyslipidemia, despite more intensive treatment with lipid-lowering agents, thus suggesting a further progression of metabolic dysfunction (Supplementary Tables 7 and 8). Within SCAPIS, HmetBMI individuals had slightly higher ground truth BMI than LmetBMI (27.5 kg m−2 versus 26.2 kg m−2) but a markedly higher metBMI than the LmetBMI (median 31 kg m−2 versus 23 kg m−2) and a more adverse cardiometabolic profile, including elevated triglyceride–glucose (TyG) index and fasting glucose and a higher prevalence of incident T2D (Extended Data Fig. 4b and Supplementary Table 8).
Clinical risk stratification and intervention response using metBMI and its residuals
To evaluate the predictive utility of metBMI, we tested its ability to classify six cardiometabolic outcomes in the SCAPIS cohort using logistic regression adjusted for age and sex (Methods). For each outcome, we compared three models: one with BMI, one with metBMI and a nested model including both. Likelihood ratio tests (LRTs) assessed whether metBMI added explanatory power beyond BMI in the nested model. MetBMI yielded the strongest predictive performance for metabolic syndrome (MetS), metabolic dysfunction-associated steatotic liver disease (MASLD), combined impaired fasting and postprandial glucose (Combined Glucose Intolerance and Type 2 Diabetes (CGI-T2D)) and screen-detected T2D (Fig. 3a). In metBMI-only models, the predicted odds ratios per 1-s.d. metBMI increase were substantial and statistically significant (MetS: odds ratio = 5.36 (95% confidence interval: 3.88–7.66, P = 2.6 × 10−22); MASLD: odds ratio = 4.95 (95% confidence interval: 3.36–7.65, P = 2.3 × 10−14); CGI-T2D: odds ratio = 2.40 (95% confidence interval: 1.88–3.11, P = 6.9 × 10−12); screen-detected T2D: odds ratio = 2.6 (95% confidence interval: 1.83–3.77, P = 2.7 × 10−7)). Nested models demonstrated a significantly improved fit compared to BMI alone (Fig. 3a), suggesting that metBMI captures additional disease signals. However, neither BMI nor metBMI predicted subclinical atherosclerosis (Coronary Artery Calcium (CAC) score and carotid plaque presence; P > 0.3 for LRTs).
a, Forest plot for six cross-sectional outcomes in the SCAPIS cohort (CAC score, carotid plaque, MetS, MASLD, CGI-T2D and screen-detected T2D). Data are presented as odds ratio estimated (center points) with 95% confidence intervals (horizontal bars), with lower and higher confidence interval limits from multivariable logistic regression per 1-s.d. increase in the predictor (BMI, metBMI or both in the nested model). The dashed line marks odds ratio = 1. P values are derived from two-sided Wald tests for BMI/metBMI. For the nested model, P is derived from an LRT versus BMI-only model. Sample sizes per outcome: CAC score (n = 212), carotid plaque (n = 268), MetS (n = 163), MASLD (n = 78), CGI-T2D (n = 136) and T2D (n = 52). b, Two-sided Spearmanʼs correlation for metBMI residuals with BMI loss 12 months after bariatric surgery (n = 75), with its corresponding P value. Each dot represents one individual, and the dashed line represents the linear regression. c, Two-sided partial Spearmanʼs correlation between metBMI residuals and all available circulating metabolites, proteins and clinical chemistry, corrected for age, sex and BMI in the IGT-microbiota cohort (n = 1,408). Positive correlations are in pink; negative correlations are in blue. Metabolites with variance explained >20% (ref. 32) or predominantly predicted by the microbiome11 are highlighted in green. Only Benjamini–Hochberg-adjusted significant correlations are shown (q < 0.05). ApoA1, apolipoprotein A1; TG, triglycerides.
The associations remained robust after adjusting for traditional risk factors (lipids, glucose, blood pressure, WHR and statin use). MetBMI remained a strong and independent predictor of MetS (odds ratio = 2.12, 95% confidence interval: 1.43–3.24, P = 3.1 × 10−4), MASLD (odds ratio = 4.24, 95% confidence interval: 2.69–6.95, P = 2.1 × 10−9) and CGI-T2D (odds ratio = 1.76, 95% confidence interval: 1.28–2.43, P = 5.0 × 10−4) risk (Extended Data Fig. 5a); continued to add predictive value over BMI in nested models for MetS (LRT P = 0.0005) and CGI-T2D (LRT P = 1.6 × 10−6); and, unexpectedly, reduced carotid plaque burden (LRT P = 0.017) (Extended Data Fig. 5a).
In an independent bariatric surgery cohort20 (n = 75; Methods), baseline metBMI residuals were inversely correlated with BMI loss/reduction at 12 months (r = −0.30, P = 0.008; Fig. 3b), despite no significant difference in baseline or follow-up BMI between HmetBMI and LmetBMI (Extended Data Fig. 5b). As expected, a higher BMI was associated with greater absolute BMI loss (Extended Data Fig. 5c). These findings highlight a dissociation between BMI and metBMI: whereas higher BMI predicts greater weight loss, higher metBMI residuals predict poorer response, suggesting that metBMI captures aspects of metabolic resistance to intervention that are not reflected in BMI alone.
Together, these findings establish metBMI and its residuals as biomarkers of a metabolically adverse obesogenic signature, capturing risk and intervention response beyond BMI and other traditional risk factors.
Characterizing clinical and multi-omics signatures of metBMI residuals
Next, we assessed how metBMI residuals relate to metabolic, anthropometric and omics data to identify the biological features behind the metabolic obesogenic signature. These residuals, orthogonal to BMI, age and sex, correlated more strongly with VAT attenuation, an imaging proxy for adipose tissue lipid content and fibrosis21, than with VAT area or liver attenuation, both indicators of ectopic fat. Additionally, metBMI residuals correlated more strongly than BMI with insulin resistance, β-cell-linked insulin hypersecretion (Homeostatic Model Assessment of β cell function (HOMA-B), fasting insulin) and impaired glucose tolerance (Extended Data Fig. 6a). Mediation analysis revealed that metBMI residuals mediated 38% of the effects of VAT attenuation (that is, adipose tissue architecture) on β cell function (HOMA-B; bootstrap 95% confidence interval: 0.28–0.51, P < 2 × 10−16), supporting their role in inter-organ metabolic regulation.
In line with these results, metBMI residuals positively associated with steroidal metabolites implicated in insulin resistance and cardiometabolic disease (for example, metabolomic lactone sulfate22 and cortolone glucuronide) as well as with glutamate and inversely with glutamine. The balance between these two amino acids, previously identified as a marker of adipose tissue dysfunction23, is highly predicted by the microbiome in our cohort (Supplementary Table 3). Other metabolites positively associated with metBMI residuals included branched-chain and aromatic amino acids as well as several phosphoinositol and phosphatidylethanolamine species. Inverse correlations included phosphatidylcholines, acetyl-carnitines, gut and diet-derived carotene diols and cinnamoylglycine11 (Fig. 3c and Supplementary Table 9).
MetBMI residuals were also associated with proteome features involved in insulin responsiveness and energy regulation across central, hepatic and adipose tissues. Positively correlated proteins included oxytocin, carboxylesterase 1 (ref. 24), leptin25 and asialoglycoprotein receptor 1, the latter reported to impair hepatic cholesterol clearance, thereby elevating circulating lipids26. In agreement, metBMI residuals were inversely correlated with insulin-like growth factor binding protein 2, whose deficiency exacerbates hepatic steatosis and worsens MASLD phenotypes27.
To assess heritability, we tested polygenic risk scores (PRSs) related to insulin secretion, adipose tissue distribution, circulating lipids and ectopic fat accumulation28,29,30: although each PRS correlated with its respective trait, neither metBMI nor its residuals was significantly captured by any PRS (Extended Data Fig. 6b).
These findings indicate that metBMI residuals reflect a non-genetic, acquired metabolic signature characterized by ectopic fat accumulation, hepatic and adipose tissue dysfunction and altered insulin signaling across omics. This aligns with the Twin Cycle Hypothesis31, whereby, depending on a personal fat threshold, liver and pancreatic interactions contribute to the individual pathogenesis of insulin resistance and metabolic disease, independent of BMI-defined obesity and across the entire BMI range.
Microbiome features of the obesogenic signature
Given the links between host metabolism and the gut microbiome15,17, we examined how metBMI and its residuals relate to gut microbiome diversity, ecological structure, composition and function. MetBMI and its residuals were more strongly and negatively correlated with gene richness than BMI (ρ = −0.19, −0.24 and −0.3 for BMI, metBMI residuals and metBMI, respectively; P < 2.2 × 10−16 for all correlations and false discovery rate (FDR) < 0.05, adjusted for age and sex as well as BMI where appropriate; Extended Data Fig. 7). In multivariable models, the addition of metBMI eliminated the significant correlation of gene richness and 359 metabolic, dietary and inflammatory markers, including BMI, HOMA-IR, MetS, WHR, CRP, renal function, leptin and dietary variables (Supplementary Table 10), highlighting metBMI as a concise summary of inter-organ and inter-organismal interactions. Notably, the gene richness of individuals with normal weight but high residuals (HmetBMI) was as low as that of individuals with obesity in the LmetBMI group (P = 0.06; Fig. 4a,b), indicating that erosion of microbiome diversity accelerates with metabolically adverse adiposity.
a,b, Microbial gene richness for individuals with lower and higher predicted metBMI within BMI classes (a) and metBMI groups across BMI classes (b), assessed using two-sided Wilcoxon rank-sum tests. Sample sizes: BMI 18.0–24.9 kg m−2: LmetBMI n = 34, HmetBMI n = 45; BMI 25–29.9 kg m−2: LmetBMI n = 67, HmetBMI n = 68; BMI ≥ 30 kg m−2: LmetBMI n = 46, HmetBMI n = 41. c, PCoA of gut microbial communities (Aitchison distance) in the IGT cohort (n = 1,408), colored by metBMI group: green, normal weight (n = 313); taupe, overweight (n = 487); purple, obesity (n = 307); light green, LmetBMI (n = 147); light purple, HmetBMI (n = 154). Large dots indicate group medoids. Variance explained by metBMI group and P values from one-sided PERMANOVA are shown. Side box plots depict group distributions across the first and second principal coordinates (two-sided Kruskal–Wallis test). In a–c, box plots show median (center line), IQR (box), whiskers to the most extreme points within 1.5× IQR and outliers as points. d, Top 50 differentially abundant bacterial species overlapping in all obesity measures. Left: feature contributions to effect size (darker = increase). Right: associations with obesity measures, adjusted for other measures; signed effect size indicated by marker color (green, increased; violet, decreased). Asterisks mark features not confounded by other measures; circles indicate confounded features. **q < 0.01; ***q < 0.001. Full data are in Supplementary Table 12. W., with.
Beyond gene richness, HmetBMI and LmetBMI groups exhibited distinct microbiome community structures. Principal coordinate analysis (PCoA) revealed clear compositional separation and clustering of HmetBMI with obesity and LmetBMI with normal weight (Fig. 4c), consistent with the observed metabolome patterns (Fig. 2b). These differences extended to ecological order, as indicated by network analyses. We observed low similarity between the two clusterings and denser, more modular consortia in LmetBMI, with a greater degree of eigenvector centrality (P = 0.000009 and P = 0.0000081, respectively, adjusted Rand index = 0.0001), indicating a larger number of interactions between nodes, anchored by Christensenellales (for example, Phil1 sp001940855) and Methanobrevibacter smithii (Extended Data Fig. 8a and Supplementary Table 11). HmetBMI networks were sparser and centered around taxa linked to metabolic dysfunction (for example, Blautia, Bacteroides, Flavonifractor, Erysipeloclostridium ramosum and Ruminococcus gnavus), which exhibited more negative interactions with health-related taxa, such as Faecalibacterium and Eubacterium (Extended Data Fig. 8b and Supplementary Table 11).
Species-level modeling, adjusted for medication and mutually controlling for BMI, VAT area and attenuation, identified 774 taxa associated with metBMI residuals (Supplementary Table 12 and Extended Data Fig. 9a,b). Of the 104 species shared with other adiposity metrics, 100 were primarily driven by metBMI residuals (Fig. 4d), with R. gnavus being the only species enriched across all traits and correlated with impaired glucose tolerance and the TyG index (Fig. 4d and Extended Data Fig. 9c). To exclude that changes in microbiome composition at the species level were secondary to decreasing microbiome richness, we adjusted for the latter. We observed that 45 taxa remained significantly associated with metBMI residuals, most notably Faecalibacterium prausnitzii and Oscillospiraceae (decreased) and oral/aerotolerant species (Streptococcus anginosus, Streptococcus mitis, Gemella and Granulicatella), which increased with metBMI residuals (Extended Data Fig. 9d). These species associated with low-grade inflammation and shifts in fatty acid, bile acid and environmental exposures, such as the plasticizer methyladipate (Supplementary Table 13). Although oral taxa tracked with proton pump inhibitor (PPI) levels, their enrichment with increasing residuals was independent of medication, suggesting parallel ecological changes created by drugs18 and metabolic injury.
Functionally, 57 GMMs associated with metBMI residuals independently of BMI or other adiposity traits (Supplementary Table 14). Residuals were marked by reduced butyrate production, mannose/glycerol utilization and increased trimethylamine production from γ-butyrobetaine and methanogenesis from trimethylamine. Even after adjusting for gene richness, two hydrogenotrophic processes remained significant along metBMI residuals—decreased methanogenesis from carbon dioxide and increased homoacetogenesis—indicating a shift in microbial carbon dioxide and hydrogen utilization, converted to acetate in HmetBMI or dissipated to methane in LmetBMI.
Together, these data suggest that metBMI residuals reflect a microbiome signature characterized by reduced diversity, altered network structure and functional shifts toward pro-inflammatory and atherogenesis-associated metabolism, capturing aspects of metabolic disruption not explained by BMI alone.
Metabolite-mediated microbiome–phenotype interactions
Gut bacteria substantially influence the circulating metabolome11, as also seen in our study (26% of inter-individual metabolite variance explained by MAGs in median; Supplementary Table 3 and Extended Data Fig. 2c) and in SCAPIS (27% variance explained)32. Given the strong covariance in metabolome and microbiome compositions, we postulated that metabolites driving the underlying metBMI signature might be closely related to the microbiome. We generated a clinically tractable signature by applying recursive feature elimination (RFE) and LASSO across 10 resamples, retaining 66 metabolites that best captured metBMI residuals (Supplementary Table 15). This reduced panel explained 38.6% of BMI variance, similar to the performance of the full 267-metabolite model (40%) and markedly more than a model comprising age, sex, triglycerides, high-density lipoprotein (HDL), low-density lipoprotein (LDL), total cholesterol and insulin (26%).
For 61 of 66 metabolites, microbial species accounted for more variance than diet or host genetics (FDR < 0.05; Fig. 5a,b and Supplementary Table 15). Of these, metabolites enriched with metBMI residuals included multiple sphingomyelins, ceramides and the microbial fatty acid derivative cis-3,4-methyleneheptanoylcarnitine, previously linked to insulin resistance and T2D33. Conversely, lower metBMI residuals were associated with 3β-hydroxy-5-cholestenoate, N-acetylglycine, indolepropionate and carotene diols, the latter two being diet-dependent bacterial metabolites with protective effects against cardiovascular risk and T2D34,35 (Fig. 5b, Extended Data Fig. 10a and Supplementary Table 15). Building on the correlations between bacterial species specific to metBMI residuals and the selected metabolites (absolute ρ > 0.1, FDR < 0.05; Extended Data Fig. 10b), we explored how bacteria may influence host phenotypes by conducting bidirectional mediation analyses among microbiome species, metabolites and clinical traits.
a, Donut plot showing microbially determined metabolites11 (orange) and metabolites with more than 20% variance explained (green), across our cohort and external cohorts32. Superpathways of these metabolites are displayed above and labeled with their proportions relative to all measured metabolites. b, Bar plot showing the median variance explained (%VE in ten models) by bacterial species for the top predictive metabolites of metBMI. Metabolite labels are colored by direction of effect on metBMI (green for lower metBMI, orange for higher metBMI). Bar colors denote superpathways. The horizontal dashed line marks the 20% VE threshold32. c, Alluvial plot of significant mediation paths (q < 0.05) between microbiome features (left) and phenotypes (right) via metabolites (middle), excluding reverse mediations. Curved lines indicate mediation effects, colored according to microbiome features. Left-side bars indicate taxonomic or functional group membership. ALAT, alanine aminotransferase; IMAT, intermuscular adipose tissue; PA, physical activity.
Among the 116 microbiome-to-phenotype pathways mediated by metabolites, bacteria from the Oscillospiraceae family (for example, uncharacterized taxa in NK3B98, UMGS902 and UMGS1865) and Christensenellales exerted protective effects via anti-inflammatory and lipid-based metabolites. For example, 1-(1-enyl-palmitoyl)-2-linoleoyl-GPC (P-16:0/18:2)36 mediated the impact of Oscillospiraceae on VAT attenuation, improved circulating lipid profiles and lower metBMI. Similarly, cinnamoglycine, a metabolite associated with microbial diversity15, carotene diols and palmitoyl sphingomyelin (d18:1/16:0), connected several Clostridia species, Christensenellales and the lysine degradation pathway of the microbiome, involved in butyrate production, with reduced WHR, improved insulin sensitivity and lower liver fat (Fig. 5c and Supplementary Tables 16–18). By contrast, bacterial species linked to higher adiposity markers and metBMI residuals, such as R. gnavus and aerotolerant/oral bacteria, exerted effects through depletion of these protective metabolites, reported reduced with escalating cardiometabolic and vascular disease17 (Fig. 5c).
Notably, 186 reverse linkages (phenotype-to-microbiome) were identified, implicating systemic inflammation (for example, CRP), dietary vitamin B6 and lipid traits in shaping microbial functions. These effects were direct (147 linkages), mediated by metabolites (seven linkages) or a combination of both (32 linkages) and were associated with functional shifts, including increased triacylglycerol and glutamine degradation and reduced dissimilatory nitrate reduction (Supplementary Table 18).
These findings demonstrate that metBMI residuals capture a bidirectional host–microbiome axis, suggesting that circulating metabolites may not only serve as functional proxies for microbiome composition but also mediate the effects of bacterial species on metabolic risk phenotypes. Disruptions in these microbiome–metabolome interactions may contribute to the metabolic dysfunction observed in subclinical adiposity-driven changes along the BMI spectrum, independent of obesity-defining thresholds (Fig. 6). This putative mechanistic link also explains the superior risk stratification of metBMI over BMI.
Light blue circle: deep phenotyping in the IGT-microbiota cohort (n = 1,408), including metabolomics, proteomics, metagenomics, diet and clinical profiling, enabled development of metBMI using ridge regression. MetBMI outperformed other omics-based and multi-omics models in capturing central adiposity, explaining over 50% of BMI variance in an external cohort (n = 466). In a surgical cohort (n = 75), higher metBMI residuals, adjusted for age, sex and BMI, were associated with approximately 30% less weight loss after 1 year. Light green circle: metBMI residuals identified individuals with metabolically adverse obesity, marked by greater VAT area and more severe attenuation, and mediated the relationship between adipose tissue characteristics and insulin hypersecretion. Light taupe circle: these residuals were linked to reduced gut microbial gene richness, altered ecological networks and enrichment of R. gnavus and aerotolerant/oral bacteria. Functional shifts included increased nitrate respiration and homoacetogenesis, alongside a reduction in methanogenesis. Light red circle: recursive feature selection and bidirectional mediation analyses identified 116 microbiome → phenotype and 186 phenotype → microbiome paths, primarily mediated by 66 circulating metabolites. This reveals a bidirectional, metabolite-centered interface between the gut microbiome and host metabolism, providing insights into the heterogeneity of obesity and its clinical manifestations. Figure created with BioRender.com. CT, computed tomography.