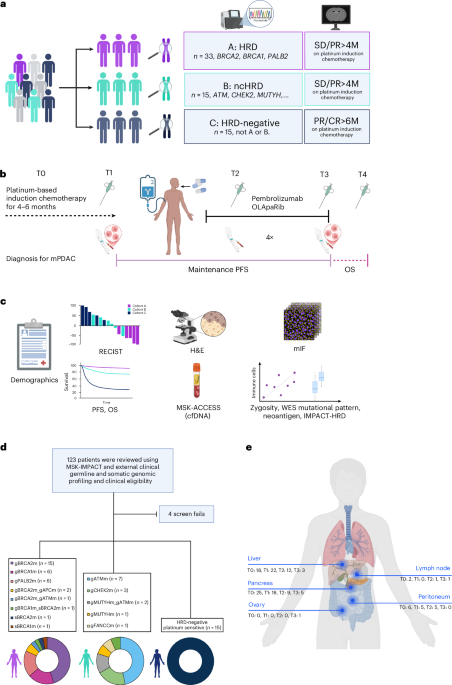
Study design
The POLAR trial enrolled n = 63 participants into three cohorts based on molecular classification and their PC response to platinum-based chemotherapy. Cohort A included participants with pathogenic germline or somatic mutations in HRD genes (BRCA2, BRCA1 and PALB2), without progressive disease after more than 4 months of platinum-based chemotherapy; cohort B (ncHRD) included participants with pathogenic germline or somatic mutations in ncHRD genes (ATM, BAP1, BARD1, BLM, BRIP1, CHEK2, FAM175A, FANCA, FANCC, NBN, RAD50, RAD51, RTEL1 and MUTYH), also without progressive disease over 4 months of platinum-based chemotherapy; and cohort C included participants with platinum sensitivity for at least 6 months (12 cycles of modified FOLFIRINOX) but without HRD or ncHRD gene mutations (Fig. 1a).
a, Cohort stratification. Participants with mPC were prospectively assigned to one of three cohorts based on germline or somatic mutation profiling and clinical response to platinum-based chemotherapy: cohort A (HRD, n = 33): core HRR gene mutations (BRCA1, BRCA2 and PALB2); cohort B (ncHRD, n = 15): non-core HRR mutations (for example, ATM, CHEK2 and MUTYH); and cohort C (HRD negative, n = 15): no HRD-associated mutations but ≥6 months of platinum sensitivity. b, Trial timeline. Schematic overview of treatment course: platinum-based induction chemotherapy (T0–T1), maintenance with pembrolizumab and olaparib, with PFS (T1–T3), on-treatment biopsy (T2) and OS tracked throughout. c, Correlative endpoints. Integrated clinical and translational analyses included demographics, RECIST version 1.1 response, PFS/OS, ctDNA profiling (MSK-ACCESS), WES for mutational patterns, zygosity, neoantigens and TME assessment by H&E and mIF. d, Screening and cohort allocation. Among 123 patients screened using MSK-IMPACT and external germline/somatic testing, 63 were enrolled and assigned to each cohort. Bottom, genomic breakdown by gene for each cohort. e, Biospecimen collection sites across timepoints. Anatomical distribution of metastatic biopsies collected longitudinally (T0–T3), with most samples from liver, pancreas and peritoneum. cfDNA, cell-free DNA; CR, complete response; M, months; mPC, metastatic pancreatic cancer; PR, partial response; SD, stable disease.
Participant enrollment occurred between 11 January 2021 and 7 March 2024. All participants received maintenance olaparib 300 mg orally twice a day daily and pembrolizumab intravenously every 3 weeks for 6 months and then every 6 weeks until disease progression or unacceptable toxicity. Select participants were allowed to continue beyond radiographic progression if deriving clinical benefit.
Serial tumor tissue (baseline, on-treatment and after progression when feasible) and matched peripheral blood samples were collected (Fig. 1b−d). These were used to investigate genomic instability, tumor immune microenvironment and neoantigen landscape features associated with therapeutic response and resistance. Specifically, whole-exome sequencing (WES), cell-free DNA and multiplex immunofluorescence (mIF) were employed from available biospecimens collected from different sites at different timepoints (Fig. 1c,e) to evaluate mutational signatures, neoantigens and immune infiltration across responders and non-responders (Methods). This translational framework enabled broad exploratory analyses of biological correlates of response to POLAR.
Baseline characteristics
Baseline clinical characteristics are summarized in Table 1. The median age at diagnosis was numerically similar across cohorts (median age, 62−65 years), with balanced sex distribution. Most participants presented with de novo stage IV disease, and all had an Eastern Cooperative Oncology Group (ECOG) performance status of 0 or 1. Tumor histology was predominantly adenocarcinoma, with a small number of acinar (n = 3) and adenosquamous (n = 1) carcinomas in cohort A. Baseline carbohydrate antigen 19-9 (CA19-9) and carcinoembryonic antigen (CEA) values at enrollment were similar between cohorts A and B but higher in cohort C, consistent with greater disease burden. Molecular characteristics at enrollment of the 63 participants are summarized in Fig. 1d and Table 2. Among cohort A, BRCA2 mutations were the most common (n = 18, 53%), followed by BRCA1 (n = 10) and PALB2 (n = 6). In cohort B, ATM mutations were the most common (n = 8, 53%), followed by CHEK2 mutations in n = 3 (20%) and BLM (n = 1), FANCC (n = 1), MUTYH (n = 1) and MUTYH_+ATM (n = 1) co-mutation. Cohort C had no detectable HRD or ncHRD mutations by design.
Primary outcomes
Among 63 participants enrolled in the POLAR trial, 46 (73%) had RECIST-evaluable disease at initiation of maintenance therapy, as assessed by independent radiology review. Objective responses were observed across all three cohorts, although the number of responses varied by genomic context (Fig. 2a). In cohort A, the ORR was 35% (7/20; 95% CI: 15−59%), which did not meet the prespecified co-primary endpoint. In cohorts B and C, the ORR was 8% (1/12; 95% CI: 0−38%) and 14% (2/14; 95% CI: 2−43%), respectively.
a, Waterfall plots show the best percent change in target lesion size per RECIST version 1.1 in cohorts A, B and C. In cohort A, bars are color coded by the gene harboring a core HRD mutation: BRCA1, BRCA2 and PALB2. In cohorts B and C, bars are shaded by IMPACT-HRD quartile, a genomic scar score derived from MSK-IMPACT targeted sequencing. Higher quartiles (3−4, darker blue) reflect greater levels of genomic instability associated with HRD and were used to stratify ncHRD (B) and HRD-negative (C) tumors. b, Kaplan–Meier curves for PFS by cohort. Median PFS was 8.3 months in cohort A, 4.8 months in cohort B and 3.3 months in cohort C. c, Kaplan–Meier curves for OS by cohort. Median follow-up was 37 months (95% CI: 27−47). In cohort A, the 2-year OS rate was 56% (95% CI: 41−76%), and the 3-year OS rate was 44% (95% CI: 28–69%). Dashed lines indicate 24-month and 36-month landmarks. Number at risk is shown below each plot. Y, year; NA, not available.
Source data
Notably, 13 of 33 patients (39%) in cohort A had non-measurable ‘undefined’ disease at baseline due to profound radiographic responses to prior platinum-based chemotherapy, a prerequisite for POLAR eligibility; all had baseline measurable disease before platinum exposure. Among these participants, 10 remained progression free on POLAR maintenance for more than 4 months (two with PFS > 6 months and eight with PFS > 12 months)—a time window commonly used to confirm radiographic response on repeat imaging—and all had a minimum follow-up of 16 months. These findings are consistent with favorable underlying HRD biology. In a post hoc exploratory analysis that included these participants with durable PFS > 4 months, the exploratory response rate for cohort A increased to 52% (95% CI: 34−69%; 17/33), suggesting enrichment for HRD-associated disease and the inherent platinum sensitivity.
The PFS for each of the three cohorts is presented in Fig. 2b. At the time of data cutoff (25 June 2025), the 6-month PFS rate was highest in cohort A at 64% (95% CI: 49−82%); however, it did not meet the co-primary endpoint of 6-month PFS rate of 77% (the protocol is attached as supplementary data). This was followed by cohort B at 47% (95% CI: 27−80%) and cohort C at 13% (95% CI: 3.7−48%). Median PFS was 8.3 months (95% CI: 5.3−NR), 4.8 months (95% CI: 4−12) and 3.3 months (95% CI: 1.9−4.8) for cohorts A, B and C, respectively. Stratified multivariable Cox analysis showed that baseline CA 19-9 was significantly associated with PFS (hazard ratio = 1.02, 95% CI: 1.02−1.03) independently of other known risk factors, including age, tumor location, duration of prior platinum therapy, gender, ECOG performance status and initial stage.
Secondary outcomes
Disease control was achieved in the majority of participants, with disease control rate (DCR) of 80% in cohort A (16/20; 95% CI: 56–94%), 75% in cohort B (9/12; 95% CI: 43–95%) and 50% in cohort C (7/14; 95% CI: 23–77%). These findings mirrored the trend observed across PFS and OS, supporting the clinical relevance of genomic stratification. Among cohort A evaluable responders (complete response/partial response, n = 7), the median duration of response (DoR) was 6.8 months (95% CI: 3.4–NR). When including patients with non-measurable (labeled as ‘undefined’) disease at baseline (n = 13), the median duration of disease control was 32 months (95% CI: 6.1–NR). Median follow-up was 37 months (95% CI: 27–47), and median OS for cohort A was 28 months (95% CI: 12–NR), 18 months (95% CI: 13–NR) for cohort B, followed by 10 months (95% CI: 8.9–24) for cohort C (Fig. 2). In cohort A (Fig. 2c), the 2-year OS rate was 56% (95% CI: 41–76%), and the 3-year OS rate was 44% (95% CI: 28–69%). Stratified multivariable Cox analysis showed that baseline CA 19-9 was significantly associated with OS after adjusting for known baseline risk factors (hazard ratio = 1.02, 95% CI: 1.02–1.03) (Supplementary Table 1). Additional prespecified secondary endpoints not reported in this paper include best overall response assessed by immune RECIST (iRECIST), biomarker response endpoints (CEA and CA 19-9 kinetics) and RECIST/iRECIST-concordant PFS analyses, which will be reported separately.
Subgroup analysis by BRCA2, PALB2, BRCA1 and ncHRD gene mutations
Subgroup analysis by mutation type within cohort A (HRD) and cohort B (ncHRD) revealed heterogeneity in clinical outcomes. Participants with BRCA2 (n = 18) and PALB2 (n = 6) mutations demonstrated numerically similar PFS and OS but longer than participants with BRCA1 mutations (n = 9) (Supplementary Fig. 2). The median PFS was 9.9 months (95% CI: 3.6−NR), 12.0 months (95% CI: 6.2−NR) and 6.1 months (95% CI: 4.1−NR) for BRCA2, PALB2 and BRCA1, respectively The median OS was 28.0 months (95% CI: 9.9−NR), 27.0 months (95% CI: 11−NR) and 18.0 months (95% CI: 12−NR), respectively. At 24 months, OS rates were 59% (95% CI: 40−88%) for BRCA2, 67% (95% CI: 38−100%) for PALB2 and 42% (95% CI: 18−94%) for BRCA1. From cohort A (n = 33), 15 participants had both germline and somatic sequencing data available, and the rest (n = 18) had only circulating tumor DNA (ctDNA) available. For zygosity status: 10 biallelic loss (seven BRCA2, two PALB2 and one BRCA1), three monoallelic loss (two BRCA2 and one PALB2)and two indeterminate (one BRCA2 and one BRCA1) (Supplementary Table 1). In cohort B, ATM mutations were the most common type (n = 9, 60%). Participants with ATM and non-ATM ncHRD gene mutations (CHEK2, FANCC and MUTYH) had similar median PFS to other ncHRD mutations (4.8 months (95% CI: 2−NR) versus 6.5 months (95% CI: 4−NR)) (Supplementary Fig. 3). Participants with ATM mutation had numerically longer OS (18 months (95% CI: 15−NR) versus 14 months (95% CI: 12−NR)).
Safety
Treatment was generally well tolerated in line with the known adverse event profiles of both drugs, and no new safety signals were observed. No grade 4 or 5 treatment-related adverse events (TRAE) occurred. Grade 3 TRAEs included anemia (n = 10, 15%) and abdominal infection (n = 1, 1.6%). Grade 2 immune-related adverse events (irAEs) included colitis (n = 1), hyperglycemia (n = 1), pneumonitis (n = 2), pancreatitis (n = 1) and hyperthyroidism (n = 1). Grade 3 irAEs occurred in n = 4 participants and included pneumonitis and colitis (Table 3).
Exploratory integrated biomarker analyses
Exploratory analyses reported in this paper were prespecified in the trial protocol and were conducted to investigate molecular and immunologic correlates of clinical benefit. These include analyses of ctDNA dynamics, tumor genomic features and mutational patterns, immune infiltration and associations with clinical outcomes. Specific analytic implementations, subgroup definitions and correlation analyses were exploratory and intended to generate hypotheses rather than to formally test prespecified endpoints. In addition, selected post hoc analyses were performed to further explore emerging signals observed in the data and are clearly identified as such in the paper.
ctDNA dynamics and clinical benefit
To explore the association between early ctDNA dynamics and clinical outcomes, we analyzed 30 representative participants’ plasma pairs (n = 60) from baseline (T1) and 6-week (T2) plasma across the three cohorts (A: n = 14 pairs, B: n = 8 pairs, C: n = 8 pairs) using MSK-ACCESS. Of these, n = 56 samples (93%) passed quality control, and only n = 32 (57%) had detectable somatic mutations. Notably, most samples at T1 had low mean variant allele frequency (VAF) less than 0.004 (Extended Data Fig. 1a), indicative of molecular residual disease (mRD). Change in mean VAF between T1 and T2 were generally minimal (range, −0.0031 to 0.013), and a trend toward higher VAF increases was observed among participants with PFS ≤ 6 months compared to those with PFS > 6 months (P = 0.063; Extended Data Fig. 1b,c). At data cutoff, five participants (7.9%) achieved durable clinical benefit with PFS exceeding 36 months. Four of these had undetectable or near-undetectable mean VAF (0.00014 and 0.00009) at both T1 and T2 timepoints; the T1 sample of the fifth participant did not pass quality control, but T2 mean VAF was 0 (Extended Data Fig. 1d). Despite unavailability of direct tumor tissue genomics in many participants, using n = 56 MSK-ACCESS ctDNA samples, we were able to recreate the OncoPrint summarizing their somatic alterations (Extended Data Fig. 1e).
HRD genotype and irAEs in relation to durable benefit
To explore the relationship between clinical and molecular features associated with clinical benefit, we generated participant-level swimmer plots (Fig. 3a) integrating PFS, genomic alterations (mutations in DNA damage response (DDR), HRD and ncHRD genes and KRAS and homozygous deletion of CDKN2A), baseline biomarkers (CA19-9 and neutrophil-to-lymphocyte ratio (NLR)) and irAEs. Eleven durable responses were predominantly observed in cohort A, including exceptionally long-term responders (PFS > 18 months) with BRCA2 (n = 8), PALB2 (n = 2) and BRCA1 (n = 1) germline mutations. One participant in cohort B with a germline BLM mutation has remained on POLAR for more than 36 months with a durable partial response (−42.2% by RECIST version 1.1). All irAEs (n = 6) occurred among patients who had PFS > 6 months across three cohorts. Other variables such as KRAS mutation allele, CDKN2A loss and baseline NLR did not separate groups with different PFS outcome.
a, Swimmer plot showing PFS (solid bar) and OS (shaded bar) annotated with clinical and genomic features for all POLAR participants, stratified by each cohort. Bars are annotated by enrollment DDR gene (for example, BRCA1, BRCA2, PALB2, ATM and CHEK2), zygosity, baseline CA19-9 level, NLR, irAE status, KRAS allele and CDKN2A homozygous deletion status. b, OncoPrint with WES showing genomic alterations across POLAR patients (n = 35) with annotation tracks for cohort, KRAS allele, HRD gene class, WGD, IMPACT-HRD scores and neoantigen quartile. c, Spaghetti plot: longitudinal changes of tumor response in patients with different levels of TIL in tumors (H&E) of the participants from cohort A. Lines are colored by TIL density score. TIL-high tumors were associated with longer PFS. d, mIF CD8+ T cell analysis: from tumors at baseline with mIF available (n = 33). Cohort A had higher CD3+CD8+ T cell infiltration (P < 0.05), and participants with longer PFS (>4 months) had higher TIL density (P = 0.055).
Source data
Cohort A (HRD) tumors are enriched for immunogenic mutational patterns and immune infiltration
WES of 35 baseline tumors (n = 15, n = 10 and n = 10 in cohorts A, B and C, respectively) revealed that cohort A (HRD) exhibited a distinct and significantly more immunogenic mutational landscape compared to cohort C (platinum sensitive without HRD) (Fig. 3b and Extended Data Fig. 2). Specifically, both total indel burden (median of 10 (interquartile range (IQR): 5−12)) and frameshift indel burden (median: 8 (IQR: 4−11)) were significantly higher in cohort A than in cohort C (median of 2 for both parameters) (P < 0.01), indicating active error-prone repair processes that preferentially generate mutation-derived neoantigens (Extended Data Fig. 2b). Although non-synonymous single-nucleotide variant (SNV) burden and total predicted neoantigen burden did not differ significantly across cohorts, this enrichment of frameshift indel mutations in HRD tumors likely contributes to enhanced neoantigen quality23,25,26.
In parallel, WES-derived tumor mutational burden (TMB) was significantly higher in cohort A compared to cohort C (median: 2.8 mutations per megabase (IQR: 2.2−4.0) versus 1.25 (IQR: 0.50−1.90), P = 0.035) (Fig. 3b and Extended Data Fig. 2a). IMPACT-HRD scores were also significantly higher in cohort A compared to cohort C (median: 47 (IQR: 22−61) versus 24 (IQR: 12−37)) (Extended Data Fig. 2a,c). These genomic features were accompanied by greater tumor-infiltrating lymphocyte (TIL) density on hematoxylin and eosin (H&E) (n = 38) in cohort A than in cohort C (median: 3.5 (IQR: 3−4) versus 2 (IQR: 2−3), P = 0.035) (Extended Data Fig. 2c). Tumors of participants with longer PFS demonstrated more abundance of TILs (Fig. 3c). Across all samples, IMPACT-HRD score positively correlated with predicted neoantigen burden (R = 0.48, P = 0.008; Extended Data Fig. 3a).
Neoantigen burden correlates with HRD but not with CD8+ T cell infiltration
Using mIF, we quantified more specific immune cells infiltrated across tumors with available neoantigen and IMPACT-HRD score. Among the mIF-available baseline tumors (n = 33), CD3+CD8+ TILs were more abundant in cohort A versus cohort C (P < 0.05) and in participants with PFS > 4 months versus PFS 4 months (P < 0.05) (Fig. 3d). Although inferred CD4+ T cell (CD3+CD8−) infiltration showed modest positive trends with neoantigen burden (R = 0.31, P = 0.2 and R = 0.30, P = 0.23, respectively), CD8+ T cells showed no correlation (P = 0.95). CD68+ macrophage infiltration trended inversely with IMPACT-HRD score (R = −0.41, P = 0.089). Programmed death ligand 1 (PD-L1) expression and T-cell-to-macrophage ratios did not significantly correlate with IMPACT-HRD score or neoantigen burdens (Extended Data Fig. 3).