Ethics statement
The TRUE-HF study (NCT05008692) was conducted under approved protocols from the University Health Network (UHN) Research Ethics Board (Toronto, Canada; REB no. 20-5205). Written informed consent was obtained from all participants before enrollment. Details of the study protocol are available21 and a summary is provided herein. The statistical analysis plan is included in Supplementary information.
Retrospective analyses of the All of Us Research Program used de-identified data accessed through its controlled-access platform. The program operates under a centralized institutional review board and all participants provided written informed consent for data collection and secondary research use. Analyses complied with program policies and applicable ethical and regulatory requirements.
Study participants of the TRUE-HF observational study
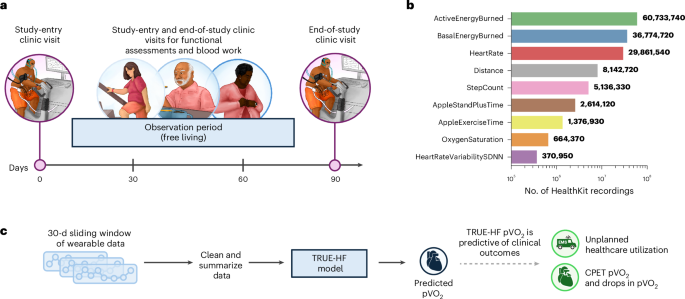
The TRUE-HF study (NCT05008692) enrolled outpatients with HF receiving care at the UHN (Fig. 1). Eligible participants were aged ≥18 years and could adequately comprehend English independently or with a caregiver’s help. Research coordinators provided study information at the time of informed consent and contacted patients for follow-up.
During the initial enrollment visit (study entry), we provided patients guidance on setting up their Apple Watch. During this session, we also educated patients on how to use Apple Watch. The appropriate electronic case report form (eCRF) and electrocardiogram (ECG) applications were downloaded and available on iPhone. Apple Watch ECG has only been validated for patients aged >22 years; therefore, only participants older than 22 years were asked to download the Apple ECG app. Participants interacted with an eCRF iOS mobile application. The data gathered in the application were not used to treat the patient. All patients underwent CPET, comprehensive bloodwork, clinical examination and a supervised 6MWT during study-entry and end-of-study clinic visits.
All demographic and clinical measurements recorded during the study-entry and end-of-study clinic visits were captured in Research Electronic Data Capture, ensuring data transcription for the study cohort. In collaboration with Apple developers, an eCRF iOS mobile application was developed using Swift to communicate study information, conduct daily surveys and gather HealthKit wearable data from patients securely and de-identified during the study duration. The wearable-derived data were not used to inform clinical decision-making.
Daily surveys captured unplanned healthcare utilization events during the free-living observation period. Patients were instructed to complete daily surveys assessing the following symptoms: increasing shortness of breath, leg swelling, palpitations, chest pain, light-headedness and fainting. In addition, the daily surveys assessed whether a patient required the following in the last 24 h: changes to medication, intravenous furosemide, unscheduled health visit, emergency room visit and/or hospital admission. Every month, we also conducted monthly fitness tests, where patients were instructed to partake in a monthly unsupervised 6MWT and Tecumseh cube test. Instructional videos could be used asynchronously to support these tests. We excluded patients with nonadherence, defined by wearing their Apple Watch <1 d throughout the 90-d free-living period.
In deriving prediction models, there is no hypothesis test to guide sample size calculation. Therefore, we aimed to calculate sample size requirements for various CI widths around clinically acceptable measures of model discrimination. The study’s sample size was determined using a classification-based approach to predict a >10% decline in pVO2, a threshold previously associated with worse outcomes in patients with HF23,24,25,26. With an assumed AUROC of 0.70, a sample size (n = 200 with completed CPETs) was estimated to provide a robust lower bound of the CI for the study objective, including model development (n = 150) and held-out test (n = 50). To prevent analytical bias, model evaluation on the held-out test set was prespecified and conducted only after the study was completed and all participants had exited follow-up.
External validation cohort: All of Us
The external validation cohort was constructed using data from the NIH All of Us Research Program v8 to validate the unplanned healthcare utilization experiments. Initially, 1,664 participants within the All of Us dataset had a documented diagnosis of HF and were available for analysis using wearable data from Fitbit devices. Among these, 400 individuals were excluded due to incomplete or missing EHR data necessary for event adjudication and accurate cohort characterization (Extended Data Fig. 2).
To align the All of Us cohort’s clinical severity with the TRUE-HF population, we restricted the cohort to include only patients with documented prior unplanned healthcare utilization. We defined unplanned events as inpatient hospitalizations (excluding planned procedures) or intravenous furosemide administration40. For each participant, the study-entry date was set as the later of the following two dates: the discharge date of their qualifying unplanned healthcare event or the first day of available wearable sensor data collection. To ensure adequate data availability, we excluded participants with <30 d of wearable data after study entry. Participants were then filtered to ensure adequate wearable data coverage, defined as at least 40% daily measurement coverage during the observational period after study entry, resulting in the exclusion of additional participants.
Finally, we defined the observation endpoint (‘end-of-study’ visit) as either the occurrence of a second unplanned healthcare utilization event within 120 d or the completion of a follow-up period that matched the TRUE-HF cohort median duration (approximately 94.5 d), with a maximum of 120 d. Participants who did not meet either endpoint criterion were excluded, resulting in a final external validation cohort comprising 193 individuals. Patient demographics are summarized in Extended Data Table 2.
Wearable data and feature engineering
The following data were collected, with informed consent from study participants, from Apple Watch through HealthKit during the approximate 90-d free-living period (that is, excluding days of baseline and end-of-study follow-up clinic visits): step count, exercise time, distance traveled, stand time, active energy burned, basal energy burned, heart rate, heart rate variability and O2 saturation. These variables were selected because they were the most frequently and consistently recorded during the interim analysis of the monitoring period and details of each are defined in Apple HealthKit41.
A standardized summarization protocol addressed the varying temporal resolutions across data types. First, abnormal data record errors were removed using an outlier approach. Records with values >3 s.d. from the population mean for each data type were removed. Next, we constructed representation of the wearable data that could integrate large-scale data for downstream usage. Specifically, first, we normalized the disparate data streams by synthesizing 90-min aggregated metrics (mean, median, minimum, maximum and s.d.) of HealthKit variables (defined above); the sum was used instead of the mean to more accurately capture the overall exercise quality of these HealthKit data types during the 90-min time window. To maintain an estimate of variable trajectories during periods of sensor nonrecording, we employed intrapatient forward-filling imputation to address gaps in the 90-min summary data, thereby preserving the autoregressive integrity of the time series. We used intrapatient forward-filling imputation to maintain the autoregressive integrity of the time series and conservatively estimated variable trajectories during sensor nonrecording.
TRUE-HF framework details
With wearable data and incorporating patient-specific clinical information such as sex, race, age, prescription dosages, weight and height, our model predicts an individual patient’s cardiopulmonary fitness and changes in their fitness over time. All nine wearable-derived features presented in Fig. 1b were included as model inputs.
Our new method leveraged a contextualized DL model to retain and analyze temporal trends across 30 d of patient-wearable data, providing near-continuous daily monitoring through next-day predictions. To achieve this, our model incorporated three distinct components: (1) it contextualized temporal representations of the data using a bottom-up approach, extending from 90-min intervals to full-day aggregation; (2) it integrated patient-specific clinical information directly into the wearable data features, allowing for adaptive feature calculations; and (3) it explicitly considered the temporal constraints, recognizing that daily activities are influenced by preceding days, and used this to make ongoing predictions for each day.
The TRUE-HF model (Extended Data Fig. 1) processed 30 d of 90-min summaries of wearable data, starting with sequential 90-min summaries and assembling them into larger time windows, allowing the model to learn temporal relationships at different temporal resolutions while improving processing efficiency. This approach is embodied in the model architecture, a bottom-up variant of the transformer model, which optimizes the feature map and reduces the temporal resolution through pooling37,42,43.
HealthKit data are first tokenized through one-dimensional convolutions to collapse features along the temporal domain44. The input is then processed through a TRUE-HF block consisting of a transformer layer followed by a pooling layer. The transformer layer learns relationships within the temporal resolution in each TRUE-HF block. Subsequently, the pooling layer aggregates pairs of consecutive time points (that is, 90–180 min). Using four consecutive TRUE-HF blocks, we effectively analyzed time resolutions of 90 min, 180 min, 360 min, 720 min and 1,440 min (daily resolution). The final prediction layer then aggregated information across the previous 30 d, inclusive, to predict the current day’s measurement.
To enhance the model’s understanding of wearable data, TRUE-HF incorporates patient-specific clinical information (described above), enabling it to learn different operations based on input attributes rather than treating all inputs equally. We achieved this by augmenting TRUE-HF with a feature-wise linear modulation block that modulates activations in the neural networks based on clinical details45. The model used only demographic information from the baseline clinic visit to maintain temporal causality.
Finally, by incorporating a causal self-attention mechanism, we explicitly constrained temporal learning to move forward only (autocorrelation) in the TRUE-HF framework42,46,47. This mechanism introduces autoregressive properties into temporal learning. It safeguards predictions for any given day being influenced only by data from that day and preceding days. We leveraged casual attention to enable additional semi-supervised training, as described below47.
Model training
All iterative model training was performed exclusively within the first 154 patients, whereas the final 63 patients were used solely for held-out testing. We excluded the days of the study-entry and end-of-study clinic visits from training.
To mitigate the lack of daily clinical CPET and 6MWT outcome labels, we utilized semi-supervised learning targeting linear approximated values of each test across the study48,49,50.
In our study, explicit daily labels were absent, and only baseline and end-of-study clinic visits provided clinical status for our target outcomes (CPET pVO2 or clinical 6MWT). To establish a reasonable approximation of our target outcome (pVO2 or clinical 6MWT) for each of these 30-d windows, we used linear interpolation of clinical outcomes recorded at the initial study-entry assessment and follow-up visits, yielding daily outcomes (Supplementary Fig. 2).
The final TRUE-HF model was an ensemble of ten models, each trained using the same TRUE-HF framework but with different random seeds. The TRUE-HF model uses exclusively wearable data and clinical data to predict future states, never past states. The average prediction from these models was used to derive TRUE-HF predictions.
Extending the TRUE-HF model to All of Us
The All of Us dataset provided per-min wearable measurements exclusively for HeartRate and StepCount, whereas critical features from the original TRUE-HF model of ActiveEnergyBurned, BasalEnergyBurned, Distance, AppleStandPlusTime, AppleExerciseTime, OxygenSaturation and HeartRateVariabilitySDNN were unavailable.
To address these differences, we employed a knowledge-distillation approach, specifically a teacher–student training strategy51,52. In this approach, predictions generated by a more comprehensive teacher model (TRUE-HF model) served as training targets (pseudo-labels) for a streamlined student model that accommodated the reduced feature set. Given the substantial feature gap between the original TRUE-HF model and the All of Us-compatible variant, we introduced a ‘teacher-assistant’ model to facilitate knowledge transfer and mitigate performance degradation53.
This teacher-assistant model retained all original wearable features but used a reduced clinical feature set aligned with the All of Us cohort. For each training batch, pseudo-labels were generated from a randomly selected member of the original TRUE-HF ten-model ensemble. Subsequently, the ensemble of trained teacher-assistant models, which provided pseudo-labels to train the final All of Us-compatible TRUE-HF-RS model, relies exclusively on HeartRate, StepCount and the reduced clinical feature set.
All TRUE-HF and TRUE-HF-RS models were trained exclusively on the training set of the TRUE-HF cohort (n = 154). The All of Us Research Program was used exclusively for external validation.
Model validation and outcomes
We compared our TRUE-HF pVO2 and TRUE-HF 6MWTD against clinically measured CPET pVO2 and 6MWTD, as well as to Apple VO2Max and sixMinuteWalkTestDistance, respectively. To predict the CPET value from the end-of-study clinic visit, we used wearable data collected over the 30 d preceding it. This ensured that all model inputs reflected free-living conditions, unaffected by structured CPET or tests conducted on the visit day. Apple VO2Max and Apple sixMinuteWalkTestDistance were collected through our iOS mobile application. Note, certain conditions or medications that limit heart rate may cause an overestimation of the Apple VO2Max algorithm—as communicated in Apple’s user interface19,54.
To further assess our predictions’ accuracy in detecting pVO2 changes over time, we measured the model’s ability to detect declines at the end-of-study clinic visit, measured against the study-entry clinical measurements. For CPET pVO2, a end-of-study drop in CPET pVO2 was defined as a ≥10% reduction from study-entry visit to end-of-study clinic visit and was chosen because a ≥6% to 10% decrease in pVO2 is associated with an increased risk of medium-to-long-term hospitalization or death in patients with HF23,24,25,26. To classify patients as a ≥10% drop in pVO2, we calculated a percentage difference between the last model prediction (that is, TRUE-HF prediction the day before the clinic visit) to study-entry CPET pVO2. The same assessment method was used for the TRUE-HF 6MWTD model, where we tested correlation measures and drops in 6MWTD and classified a decline in 6MWTD (10% reduction in distance walked), respectively.
Our secondary objective was to evaluate the association between TRUE-HF-predicted declines in daily pVO2 and unplanned healthcare utilization during the 90-d follow-up period. This association was compared to associations of traditional static risk factors measured at baseline and clinical models (MAGGIC, SHFM and PREDICT-HF). We defined unplanned healthcare utilization as hospitalization, unscheduled clinical visits or urgent intravenous furosemide treatment taking place between the study-entry and the end-of-study visits. The first prediction made by our TRUE-HF pVO2 model required 30 d of data. Hence, this objective was evaluated only among patients free from unplanned healthcare utilization in the first 31 d of our study.
Explainability and feature analysis
We examined the impact of removing structured monthly exercise sessions. All wearable data collected during these sessions were masked by excluding measurements within a 90-min window around the start and end times. A 30-d window was chosen before final analyses (described in the statistical analysis plan). We also evaluated how input window length affected accuracy by comparing shorter windows (10 d or 20 d) with the 30-d window using retrained models and zero-shot inference. Saliency analyses on the combined TRUE-HF model quantified feature importance, averaging saliency values daily for visualization. To assess feature contributions, we trained and compared (1) a fully connected neural network with clinical baseline variables and (2) a model using only wearable data.
Statistical analysis
The analyses and methods were prespecified before data evaluation and performed only on the held-out test set (n = 63), consisting of the last 50 patients who successfully completed an end-of-study clinic visit. We followed the STROBE and MI-CLAIM reporting guidelines55,56.
The primary analysis compared our model’s observed and predicted CPET pVO2 values. We used Spearman’s coefficient to measure the rank-based association between observed and predicted values. Pearson’s r was employed to quantify linear correlation. Together, these two correlation measures capture complementary aspects of the model’s predictive fidelity. The m.a.e. was chosen for its interpretability in the same units as the outcome. AUROC was used to evaluate the diagnostic accuracy of correctly predicting a meaningful drop in outcome measurements.
To estimate AUROC CIs, we employed 1,000 stratified bootstrap resamples and stratified data resampling techniques. Based on Delong’s test, a two-sided P < 0.05 was considered significant between models.
The secondary analysis used time-varying, extended, Simon and Makuch’s Kaplan–Meier cumulative risk curves to assess the association between an observed drop of ≥10% in their TRUE-HF daily pVO2 and unplanned healthcare utilization57. In this analysis, the cohort was stratified into two groups: (1) patients with a ≥10% reduction in daily pVO2; and (2) patients with <10% reduction in daily pVO2. We constructed cumulative incidence curves for unplanned healthcare utilization and accounted for the time-varying nature of our independent variable. To assess the probability of event-free survival between these groups, we constructed cumulative risk curves to account for the continuous change in our daily pVO2 covariate, facilitating a longitudinal comparison of clinical outcomes related to unplanned healthcare utilization. An extended Cox’s proportional hazards model quantified the HR and the strength of association between time-varying occurrence of pVO2 drops (scaled in 10% drops) in TRUE-HF daily pVO2 and subsequent clinical events58. Sensitivity analyses involving minimal covariate adjustment were performed. Due to the limited sample size, these analyses used single covariate adjustments rather than comprehensive multivariate adjustments.
Secondary analysis AUROC was computed as the maximum percentage drop in model prediction from the first model prediction to the prediction on the day for each patient, preceding any event or end-of-study clinic visit (censor). DeLong’s one-tailed test was performed to evaluate the a priori hypothesis that continuous monitoring methods would surpass static measures in discriminative performance. Multiple testing correction was applied using the Benjamini–Hochberg method (target false discovery rate = 0.05). Landmark-based, time-dependent AUROC analysis was performed during external validation to assess the model’s ability to predict outcomes based on the largest percentage drop identified up to each landmark time t59. This analysis was performed in 10-d intervals, with 30-d outcome windows repeated.
Qualitative results for the trends observed with TRUE-HF predictions were created using LOWESS-smoothed trajectories, generated by forward-filling censored or missing data across the entire study window using the last observed value before censoring or event.
Data preprocessing and model development were performed in Python using Pandas (v1.5.3), NumPy (v1.21.3) and PyTorch (v2.0.0). Correlation analyses, m.a.e. calculations and Simon and Makuch’s Kaplan–Meier estimations were conducted with SciPy (v1.7.1) and lifelines (v0.28.0). AUROC, DeLong’s test, the Benjamini–Hochberg correction and Cox’s proportional hazards models were implemented in R using pROC (v1.18.5) and survival (v3.6-4) packages and the timeROC package (v0.4). TRUE-HF model architecture definitions, wearable data preprocessing and inference workflows have been made publicly available at https://github.com/mcintoshML/TRUEHF.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.