This study was approved by the Oxford Tropical Research Ethics Committee (OxTREC: no. 15-19), the National Institute of Medical Research in Tanzania (no. NIMR/HQ/R.8a/Vol.IX/3408) and the Uganda National Council of Science and Technology (UNCST: no. HS529ES), and was conducted in accordance with the Declaration of Helsinki. Written informed consent was obtained from parents or legal guardians of all participating children, with age-appropriate assent obtained where applicable. Participants did not receive financial compensation for study participation.
Study design
This study used data from the prospective, multicenter, observational, Aggressive Infection-Related East Africa Lymphoma (AI-REAL) study43, which was conducted in two phases. In phase I, we trained different liquid biopsy models and assessed their performance for the diagnosis of EBV+ BL using histopathology with a limited IHC panel and review by a minimum of 2 histopathologists as the GSP in 212 participants with clinically suspected lymphoma. In phase II, we performed a head-to-head comparison between the liquid biopsy and gold-standard tissue biopsy to assess the TAT for the two methods (n = 58 participants with clinically suspected lymphoma) and to conduct an additional independent external validation for the best-performing model (n = 56 participants with clinically suspected lymphoma).
Participants
The study enrolled all children and young adults, from the age of 3 years to 25 years, clinically suspected of having lymphoma and consenting to participate in the study from August 2019 to July 2023 from four hospitals in Tanzania and Uganda, that is, the Muhimbili National Hospital, Kilimanjaro Christian Medical Centre, Bugando Medical Centre and St. Mary’s Hospital Lacor. Participants with suspected lymphoma who had previously received chemotherapy, immunotherapy or any investigational agent for lymphoma, or participated in a clinical trial on lymphoma, were excluded. Phase I of the study was conducted from August 2019 to July 2022 and enrolled a total of 313 participants, whereas phase II was conducted from August 2022 to July 2023 and enrolled a total of 64 participants (Extended Data Fig. 1).
Tissue biopsies were obtained by a qualified surgeon through excisional biopsy and immediately fixed in 10% neutral buffered formalin. The formalin-fixed specimens were then transported to the local pathology laboratory for further processing and diagnostic evaluation. In parallel, liquid biopsy samples were collected via venipuncture into Roche Cell-Free DNA Collection Tubes, following standard phlebotomy procedures. The tubes were transported to the molecular laboratory, where plasma was separated by centrifugation and stored at −80 °C until further analysis.
Laboratory procedures
Extended Data Fig. 7 provides a summarized overview of the laboratory procedures performed for both the liquid and the tissue biopsies. All measurements were taken from distinct, independent samples.
Gold-standard pathology
Before study initiation, we performed a comprehensive training and infrastructure needs assessment at the three local study pathology laboratories. To ensure optimal histopathology assessment for the study and local capacity building, technical staff and the four clinical study pathologists underwent training, followed by competency assessments conducted by expert hematopathologists in tissue fixation, embedding, sectioning and IHC staining techniques. The training included hands-on sessions and on-site mentoring over a period of 1 year. An automated IHC staining platform (Ventana Benchmark GX) was installed and maintained at all participating laboratories for the duration of the study and 3-monthly internal quality control procedures were implemented, including the use of known positive controls. Tissue samples were fixed in 10% neutral buffered formalin and embedded in paraffin according to standard protocols. H&E staining was performed on all samples to evaluate morphology and IHC was done using the previously described limited IHC antibody panel for BL.
This diagnostic approach utilizes a three-phase scoring system. The first phase combines typical BL morphology with immunostains BCL2, CD10 and CD20 to establish a diagnosis. Cases that remain inconclusive proceed to a second phase with CD38, CD44 and Ki67, whereas unresolved cases undergo a third phase with FISH analysis for MYC–Ig translocation. In our study, we used the first-phase immunostains (BCL2, CD10 and CD20) to support the diagnosis of BL, because this phase has previously been shown to achieve 81% concordance with the World Health Organization SoC pathology and is well suited for the limited-resource settings12,44. During the study, we purchased and procured all relevant antibodies and ensured that these were available. Finally, we introduced digital whole-slide imaging to enable slide review by a minimum of two local, study histopathologists and to gauge external third opinions as required45. This approach was adopted as the gold standard for evaluating the performance of the liquid biopsy methods.
CfDNA extraction, library preparation and targeted sequencing
CfDNA extraction, library preparation and targeted sequencing were performed at two dedicated study laboratories; the Haematology clinical Research Laboratory at the Muhimbili University of Health and Allied Sciences in Tanzania and the Central Public Health Laboratory.
Sample collection and processing
Approximately 8 ml of whole blood was collected in Roche circulating cfDNA (ccfDNA) tubes or PAXgene blood ccfDNA tubes. Plasma was separated by centrifugation at 1,600g for a duration of 15 min continuously, followed by a second centrifugation of the separated plasma at 4,500g for 15 min. Separated plasma was stored in 1.5-ml Eppendorf tubes at −80 °C.
Extraction and quantification of cfDNA
CfDNA was extracted from plasma using the QIAamp Circulating Nucleic Acid Kit (QIAGEN, cat. nos. 51304 and 51306) according to the manufacturer’s instructions. Briefly, it involved four main steps; first, samples are lyzed to inactivate DNases and RNases and allow complete release of nucleic acids from bound proteins, lipids and vesicles. Second, the lysates are transferred onto a QIAamp Mini column and circulating nucleic acids are adsorbed from a large volume on to the small silica membrane as the lysate is drawn through by vacuum pressure. Third, while the nucleic acids remain bound to the membrane, contaminants are washed away in a three-step wash process. The final step involves elution of highly purified nucleic acid. The quantification of cfDNA was done with a Qubit fluorimeter 3.0 using the qubit high-sensitivity assay (Thermo Fisher Scientific).
Library preparation and targeted sequencing
Libraries were constructed using 50 ng (in 30 μl) of extracted cfDNA with the ThruPLEX Tag-Seq HV kit according to the manufacturer’s protocol. The process involved addition of unique ThruPLEX HV Unique Dual indexed PCR primers to aid with sample tracking. Seven cycles were performed to ensure a yield of >500 ng depending on the concentration of the input DNA. This was followed by a purification step through magnetic separation using AMPure XP beads (Beckman Coulter). The final library was quantified and validated using the Qubit HS kit and the Bio-analyser High Sensitivity DNA kit, according to the manufacturer’s instructions. Library hybridization and capture were done using the xGen hybridization capture kit according to the manufacturer’s instructions.
Panel design and verification
The procedure used a custom-made NGS panel targeting mutational hotspots in 17 EBL-related and HL-related genes (MYC, IGH, IGK, IGL, ID3, TP53, TNFAIP3, NFKBIE, SOCS1, EP300, BTK, STAT6, CSF2RB, ITPBK, XPO1 and B2M), selected based on results from previous studies and EBV genes expressed in latently infected cells: EBER1, EBER2 and EBNA2. The final panel manufactured by Integrated DNA Technologies (IDT) consisted of 731 probes at a size of 148 kb to permit compatibility with either the iSeq100 or the MiSeq sequencing platforms. The final panel design with the genomic coordinates of our genes of interest is listed in Supplementary Table 4. Target enrichment was performed using a hybrid capture approach and sequencing was conducted twice weekly on the MiSeq platform using the MiSeq reagent kit v2 (300 cycles) at a loading concentration of 10 pM, with 6 samples per run.
Variant calling
Structural variants were detected by split-read analysis using GRIDSS and SNVs were called using VarScan 246. The following filtering strategies were used to call somatic variants:
-
(1)
Panel of normal plasmas: we excluded variants that occurred in ≥2 samples in a separate cohort of 12 healthy controls.
-
(2)
Population databases: all variants were annotated against gnomAD and those with a population allele frequency (minimum allele frequency (MAF)) > 1% were excluded.
-
(3)
VAF: to minimize the possibility of including germline variants, we excluded all variants with VAF ≥ 40%, in addition to the initial exclusion of low-level artefacts with VAF < 1%.
-
(4)
Sequencing depth and read support: only variants with a minimum sequencing depth of 500× and at least 5 mutant reads were retained for downstream analysis.
-
(5)
Copy number alterations (CNAs): we excluded variants that showed evidence of being affected by CNAs. This was done through an internal normalization approach in which the expected relationship between VAF and log₁₀(transformed ctDNA) was modeled using ID3 mutations (rarely affected by CNAs), and the resulting tolerance limits (±2× median absolute deviation of residuals) were applied to MYC and TP53 variants to flag likely CNA-affected outliers (Extended Data Fig. 4a,b and Supplementary Table 6).
Samples that had GSP tissue biopsy results were prioritized for sequencing to allow for clinical validation. For phase II samples, batching was not applied; instead, samples were transported to the sequencing laboratory immediately after collection and sequenced in a head-to-head comparison against GSP tissue biopsy, allowing us to measure TAT prospectively. The same twice-weekly sequencing schedule was maintained, with each MiSeq run limited to a maximum of six samples. Details on the target coordinates (Supplementary Table 4) and sequencing parameters (Supplementary Table 5 and Supplementary Fig. 2) are described in Supplementary Information.
Bioinformatics analysis
The sequence data were analyzed and stored by a bespoke pipeline (courtesy of the Oxford Molecular Diagnostic Centre) in a Health Insurance Portability and Accountability Act of 1996-compliant, Amazon Web Services cloud. Paired reads were combined and statistics generated on the total number of reads and invalid reads, using a customized tool (udini). The raw sequence data in FASTQ format were aligned to GRCh37 (hs37d5) with BWA-MEM2. Sorting was done using samtools. De-duplication of reads and generation of error rates and family sizes were done by a custom-made tool (elduderino). The de-duplicated FASQ was re-aligned to GRCh37 with BWA-MEM2 and fragment sizes calculated using read pairs via a customized script (available on request). Another customized script (available on request) was used to map the reads on to the genomic regions (targeted genes) of interest and to calculate copies per cell for the EBV genes (mean depth of EBV gene per mean depth for all other targeted genes). The last base at the end of each read was trimmed using a customized tool (trim) and the trimmed SAM file converted into a BAM file and indexed. A customized script (available on request) was used to generate summary statistics for coverage of all targets at different coverage depths and for different genes.
Customized scripts (available on request) were developed for variant calling using VarScan and annotation of the variants using Ensembl Variant Effect Predictor. IgCaller (v1.2 software utilizing the hg19 reference genome) was used to comprehensively analyze the rearrangements of the Ig genes and identify oncogenic translocations and the Genomic Rearrangement Identification Software Suite, with additional Picards options and Samtools as aligner (GRIDSS, v2.13.2), was also included as a structural variation caller. To visualize and distinguish true variants from false variants, Integrated Genome Viewer (IGV, v2.13.0) was used. EBV DNA size ratio was calculated as the proportion of EBV DNA fragments with size between 180 bp and 200 bp ((Total EBV DNA with size 180–200 bp)/(Total autosomal DNA with size between 180 bp and 200 bp)). Calculation of the size ratio and selection of the fragment size window were determined based on the methodology of a previous study looking at EBV DNA in nasopharyngeal carcinoma26. The lower the EBV size ratio, the lower the proportion of EBV DNA molecules of size 180–200 bp. The distribution of the fragment sizes for reads that map to regions of EBV and for reads that map to the autosomes was calculated and recorded as the EBV entropy and autosome entropy, respectively. This gives a measure of how wide or clustered the distribution of fragment sizes is. The EBV DNA size ratio, EBV entropy and autosome entropy were calculated using customized scripts (Fig. 3 and Supplementary Information).
Liquid biopsy predictor description
The following covariates were tested as predictors in the liquid biopsy diagnostic model:
EBER1, EBER2 and EBNA2 (EBV genes) quantity (copies per cell): calculated as the mean depth of the respective EBV gene divided by the mean depth for all other targeted genes and expressed as copies per cell.
EBV fragment size ratio (EBVSR): EBV DNA size ratio was calculated as the proportion of EBV DNA fragments with size between 180 bp and 200 bp ((Total EBV DNA with size 180–200 bp)/(Total autosomal DNA with size between 180 bp and 200 bp)). Calculation of EBVSR and selection of the fragment size window was determined based on the methodology of a previous study looking at EBV DNA in nasopharyngeal carcinoma26.
EBVP: represents the number of EBV DNA reads divided by the total number of sequenced reads (after removal of PCR duplicates), expressed as a proportion26.
EBV DNA and autosomal entropy: calculated as a measure of the diversity of fragment length distributions, using Shannon entropy computed across the full distribution of paired-end sequencing fragment sizes mapping to the EBV genome (for EBV entropy) and the autosomal regions (for autosomal entropy), respectively. Fragment sizes were binned into 20-bp intervals and the proportional frequency of fragments in each bin was used to derive entropy, reflecting the overall variability and dispersion of the fragment size distribution.
MDT meeting decision tree
To further evaluate the potential clinical utility of the liquid biopsy diagnostic test, we established a weekly virtual MDT meeting consisting of pediatric hematologists and oncologists, hematologists, pathologists and laboratory scientists from all study sites, as well as a senior study bioinformatician.
When tissue biopsy was reported based on H&E stain only (minimum pathology report) and the liquid biopsy result indicated BL, treatment was initiated as BL (Fig. 3). When liquid biopsy indicated non-BL, treatment was deferred until GSP tissue biopsy results were available. If the liquid biopsy report was unavailable at the first MDT meeting, treatment decisions were deferred until GSP tissue biopsy results were available.
In cases where BL diagnosis was definitive by liquid biopsy, treatment was initiated (Fig. 3). When liquid biopsy indicated non-BL, treatment was deferred until the GSP tissue biopsy result was available. If neither tissue biopsy nor liquid biopsy results were available at the first MDT meeting, treatment decisions were deferred until the second MDT meeting in the following week. Our analysis focused on the initial MDT review to capture the immediate outcome of the two tests in the early decision-making process for cases of BL, although enhanced follow-up protocols in phase II ensured that results from both liquid biopsy and tissue biopsy were obtained for all cases and integrated into patient management plans.
Turnaround time
For the tissue biopsy TAT analysis, we assessed the time from patient presentation at the hospital to the issuance of the GSP diagnostic report, divided into key intervals: time from presentation to sample collection; collection to laboratory receipt; receipt to H&E slide processing; time to issuing the first diagnostic report (based on H&E review); and time to completing the final GSP diagnostic report. For the liquid biopsy TAT, we measured the time from sample receipt in the laboratory to issuance of the diagnostic report, broken down into the intervals of cfDNA extraction and library preparation, targeted sequencing, bioinformatic analysis and final report generation. For the direct TAT comparison between GSP tissue biopsy and liquid biopsy, we specifically analyzed the time from laboratory sample receipt at the respective laboratories to the issuance of the diagnostic report for each method.
Diagnostic yield
We assessed the diagnostic yield for BL at the first MDT meeting, expressed as a percentage and calculated from the number of BL cases diagnosed at the first MDT meeting by either the GSP tissue biopsy or the liquid biopsy method divided by the total number of confirmed cases of BL.
Statistical analysis
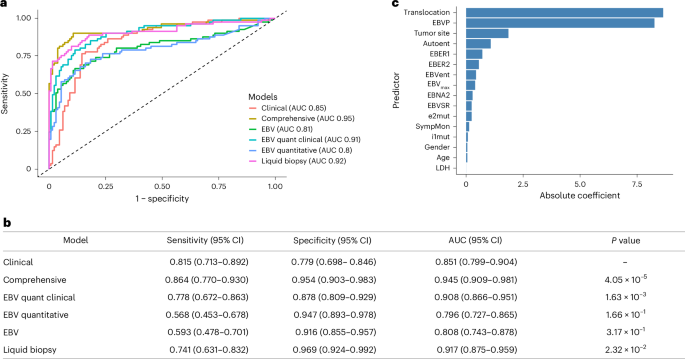
All statistical analyses were conducted in R v4.2.3. Normality of continuous variables was assessed using the Shapiro–Wilk test and visual inspection of histograms and Q–Q plots. As the data were not normally distributed, nonparametric summaries and tests were used where appropriate. Descriptive statistics were summarized as proportions for categorical variables or medians with IQRs for continuous variables. The Pearson’s χ2 test and Wilcoxon’s rank-sum test were used to assess associations between variables. We applied the Benjamini–Hochberg procedure to adjust for multiple comparisons across 19 variables. False recovery rate-adjusted P values are presented and values <0.05 were considered statistically significant. All variables showing a significant association with the diagnosis of BL were considered for inclusion in a penalized logistic regression model that we previously described and validated using a smaller cohort20. Before model fitting, candidate predictors were examined for multicollinearity using pairwise correlation and variance inflation factor analysis and highly collinear variables were excluded to improve model stability and interpretability. Priority was given for those that were most biologically or clinically relevant. Six different models were created based on a combination of different variables, as shown in Table 2.
For each diagnostic model, the optimal penalty parameter (λ) was selected via tenfold crossvalidation to minimize classification error and ROC curves were generated. The best-performing model was identified based on the highest AUC and pairwise comparisons of AUCs were conducted using DeLong’s test to assess the statistical significance of performance differences between models. The relative importance of variables contributing to the performance of the comprehensive model was derived from the absolute values of the coefficients obtained through LASSO regression. In this penalized logistic regression framework, variables with larger absolute coefficients exert a greater influence on the model’s predictions, reflecting their relative contribution to distinguishing cases of BL from non-BL cases. External validation was done by predicting the diagnosis on a different cohort (phase II) based on the best-performing model. In this analysis, missing data for LDH in 12 samples was imputed using multiple imputation by chained equations, implemented in the mice R package (v3.16.0). Predictive mean matching (method = ‘pmm’) was applied. Comparison of the median TAT between the liquid biopsy and the tissue biopsy was done using Wilcoxon’s signed-rank test (for paired samples).
Statistics and reproducibility
A formal sample size calculation was conducted to ensure sufficient power for assessing diagnostic accuracy. Using a binomial proportion approach (z = 1.96, expected sensitivity = 0.8, margin of error = 0.10), a minimum of 62 cases with BL was estimated, corresponding to approximately 124 participants, assuming a 1:1 case–control ratio. For multivariable model development, an events-per-variable threshold of 20, with up to 18 candidate predictors, indicated a target sample size of 720 participants. Owing to the prospective design and disruptions related to the COVID-19 pandemic, this target was not reached. The final cohort comprised 212 participants, including 81 cases of BL. To mitigate overfitting in the context of a constrained sample size and multiple candidate predictors, LASSO regression was used for variable selection and regularization. Model performance was internally validated using tenfold crossvalidation to assess robustness and generalizability, in accordance with recommended best practices for predictive model development.
Participants were enrolled consecutively as part of routine clinical care. No randomization was performed and investigators were not blinded to allocation during experiments or outcome assessment. Participants with missing outcome data were excluded from the analysis; this exclusion was predefined to preserve the integrity of outcome-based comparisons and was limited to individuals lacking key clinical endpoints or follow-up information required for the primary analyses. For other missing data, including covariates, appropriate imputation methods were applied to minimize bias and retain statistical power.
Reproducibility was supported through the use of standardized laboratory protocols, predefined diagnostic algorithms established a priori and automated bioinformatics pipelines applied consistently across all samples. Source data and analysis code are made available as described in ‘Data availability’.
Inclusion and ethics
This research was conducted through equitable collaboration between institutions in Tanzania, Uganda and the UK, with a deliberate focus on capacity strengthening, technology transfer and shared scientific leadership. NGS platforms were transferred and implemented locally, enabling all cfDNA sequencing to be performed in-country. Laboratory scientists, clinicians and bioinformaticians participated in structured, crosscountry training and reciprocal site visits, supporting hands-on skills development in library preparation, sequencing, bioinformatics analysis and clinical interpretation. Bioinformatics pipelines and reporting frameworks were deployed with local oversight, ensuring full access to data and analytical workflows for all collaborating investigators. The program supported formal academic training, including PhD, DPhil and masters training for local scientists, contributing to sustainable research capacity beyond the duration of the study. Study design, implementation, data interpretation and authorship reflected shared leadership and local ownership.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.