Feasibility in routine cancer diagnostics
The investigated cohort comprised of 935 unique patients for whom WGS was requested between January 2021 and November 2022. The mean age at sequencing was 60.9 years, with 54% of patients being female. The most common cancer types were NSCLC (23%), CUP (16%) and soft tissue sarcoma (10%) (Supplementary Table 1a).
The suitability of tissue samples for WGS was based on cryosection analysis of fresh frozen biopsies or surgical specimens. A pathologist annotated viable tumor cells for manual microdissection to enrich the tumor cell percentage of all samples. If samples were too small for automated isolation with a robot (<6 mm), in-house DNA extraction was performed to improve sequencing feasibility for smaller samples34. As 90 samples lacked any tumor cells in the cryosection, 888 suitable tissue samples could be prepared for WGS.
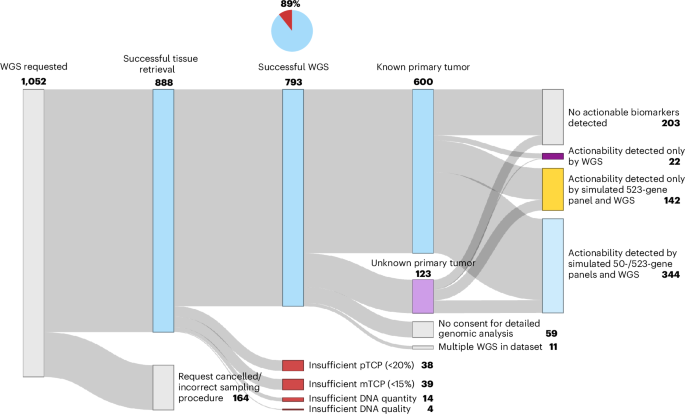
Successful diagnostic sequencing reports were generated for 89% (793 of 888) of samples (Fig. 1). Cytology samples achieved a lower success rate (57%; 20 of 35) than surgical specimens or biopsies (91%; 773 of 853) (Supplementary Table 1b,c). WGS performed on archived samples achieved comparable success rates (90%; 135 of 150). Notably, the median turnaround time was 6 working days (average, 6.7; range, 3–22) from sample reception at the sequencing facility until reporting (Extended Data Fig. 1).
Starting from 1,052 total WGS requests, reasons for discontinuation and failure as well as diagnostics outcomes are indicated. Patients who did not have consent for patient-level genomic data analysis were included only for general patient characteristics. Panel coverage of a commonly used 50-gene panel and a 523-gene panel was simulated. Potential actionability includes standard-of-care and experimental biomarker-informed treatments. pTCP, pathologist-assessed tumor cell percentage.
Source Data
Among 95 samples that failed to yield WGS reports, 40% (38 of 95) contained an insufficient pathologist-assessed tumor cell percentage (<20%) despite enhancement of tumor cellularity34. These samples were not processed for sequencing. Although WGS normally requires a minimum molecular tumor cell percentage (mTCP, based on sequencing results) of 20%, previous analyses indicated that mTCPs between 15% and 20% maintained a sensitivity of more than 95% without increased false positives. We followed institutional practice in our analyses by adopting this mTCP threshold of 15% while disclaiming in the patient report that biomarkers may have been missed when mTCP was between 15% and 20%. Forty-one percent (39 of 95) had an mTCP below 15% after sequencing and were thus considered failed. Finally, 15% (14 of 95) had insufficient DNA quantity (<50 ng), and 3% (4 of 95) had library preparation failures due to inadequate DNA quality.
Clinical actionability and simulation of panel-based testing
Next, we investigated the presence of potentially clinically actionable biomarkers identified through WGS. We focused on biomarkers indicating positive or negative selection for available reimbursed or experimental treatments, as registered in the electronic health record (EHR) following molecular tumor board (MTB) discussion.
A total of 793 WGS reports were generated, but consent for patient-level analysis of EHR data was available for only 723 patients (Table 1 and Supplementary Table 1b). Eleven patients had multiple WGS analyses during their patient journey (Supplementary Table 2b).
Overall, we found that 73% (527 of 723) of patients harbored at least one potentially actionable biomarker. An overview of cancer drivers and actionable biomarkers can be found in the supplementary data (Supplementary Fig. 1 and Supplementary Table 2a,c–e) as well as in the comprehensive ‘cancer vignette’ visualization of the genomic landscapes of all patients in this study and the NSCLC subcohort. In these visualizations, genomic characteristics of the cohort described in this study are compared with data of ~5,000 patients present in the Hartwig Medical database (Extended Data Figs. 2–4), demonstrating that patients in the current study are representative from a genomics perspective.
We simulated the coverage of commonly used 50-gene (small) and 523-gene (comprehensive) NGS panels within our WGS dataset and show that these test approaches would detect actionable biomarkers in only 49% (352 of 723) and 70% (505 of 723) of patients, respectively (Fig. 1 and Supplementary Table 3a–c). Furthermore, in 8% (40 of 505) of patients for which actionable biomarkers could be detected using a comprehensive panel, WGS identified additional actionable biomarkers. In 3% of patients (22 of 723), no potentially actionable biomarkers would have been identified using panel-based NGS, yet WGS revealed clinically relevant alterations; these were predominantly gene fusions (n = 11) and HRD in the absence of pathogenic mutations in known homologous recombination pathway genes (n = 7). Collectively, these findings suggest that about one in ten patients may benefit from the broader actionability detection of WGS compared with comprehensive panel-based screening.
We note that patients with CUP represent a unique subcohort. As WGS serves a critical role in establishing the initial cancer diagnosis, delivery of non-biomarker-informed systemic therapy due to demystification of the tumor type is as relevant as finding an actionable biomarker. We therefore analyzed patients with known primary tumor diagnoses (n = 600) and patients with CUP (n = 123) separately.
Actionability in patients with known primary tumors
Among 600 patients with known primary tumors, 73% (437 of 600) presented one or more clinically actionable biomarkers, averaging 1.3 actionable biomarkers per patient (median, 1; range, 0–5) (Fig. 2a,b). The median tumor mutational burden (TMB) was 3.3 mutations per megabase (Fig. 2c). Notably, 27% (162 of 600) of tumors exhibited biomarkers linked to reimbursed treatments, most commonly EGFR mutations (70 of 600). Sixty-three percent (379 of 600) of patients had biomarkers enabling experimental treatments, mostly based on a high TMB (≥10 mutations per megabase; 147 of 379) (Fig. 2d, Supplementary Fig. 1 and Supplementary Table 2c–e).
Data are shown separately for patients with known primary tumors (blue, n = 600) and CUP (purple, n = 123). a, Number of biomarkers and actionable biomarkers reported per patient. Boxes indicate the interquartile range (IQR), center lines the median and whiskers the extreme values within 1.5 × IQR. Outliers are represented by dots. b, Number of patients with zero, one, two or three or more actionable biomarkers. c, TMB distribution in mutations per megabase. The line indicates the median. d, Fractions of patients harboring actionable biomarkers per tumor type (with at least five representations in the cohort). Bars are proportioned to 100%.
Source Data
An average of 1.8 (range, 0–10) systemic treatments were given before WGS. Twenty-six percent (158 of 600) of patients had no prior systemic treatments, and 19% (114 of 600) had received a biomarker-informed treatment (Supplementary Table 2a).
Biomarker-informed treatment in patients with known primary tumors
Next, we evaluated patient allocation to biomarker-informed treatments. We excluded 12 patients who had died or were no longer in care when WGS results were reported. The average follow-up was 10.1 months (median, 8.8; range, 0.0–28.7).
Twenty-seven percent (159 of 588) of patients had a biomarker for a reimbursed treatment, and 40% (63 of 159) of them did start such treatment. Notably, 21 of these were patients with NSCLC whose EGFR-, ALK-, MET-, RET- or ROS1-targeted tyrosine kinase inhibitor treatment was changed after WGS identified biomarkers suggestive of resistance.
A common reason not to start a reimbursed biomarker-informed treatment was that detected biomarkers contained no new actionable insights over previously initiated treatments (21%; 34 of 159). Additionally, three patients were enrolled in a clinical trial based on biomarkers for which a reimbursed treatment was also available. Another major reason not to pursue biomarker-informed treatment was related to ongoing response or complete remission after previous treatments, including surgery or radiotherapy (18%; 28 of 159). This number could thus change with longer follow-up. Finally, five patients did not start a reimbursed biomarker-informed treatment due to a poor performance status (Supplementary Table 2a).
Sixty-three percent (373 of 588) of patients had actionable biomarkers enabling experimental biomarker-informed treatments, with 19% (71 of 373) starting with this treatment. Sixteen percent (58 of 297) were allocated to a clinical trial, and 3% received biomarker-informed treatment through either a compassionate use program (8 of 373) or an early access protocol (5 of 373).
CUP tissue-of-origin analysis and actionability
The outcomes of the WGS-based tissue-of-origin prediction algorithm (Cancer of Unknown Primary Prediction Algorithm (CUPPA)) are routinely reported for all patients27. We found that for 49% (60 of 123) of CUP cases, a tissue of origin could be predicted confidently (confidence score ≥0.80). In an additional 14% (17 of 123), a conclusive diagnosis was also achieved by combining findings with lower confidence scores with prior clinicopathological findings, including from radiology, nuclear medicine or endoscopy (Fig. 3). Collectively, WGS led to a definitive tumor-type diagnosis for 63% of CUP patients (77 of 123; Supplementary Table 4a,b).
Treatment decision outcomes and frequencies and types of treatments administered following WGS are shown for all CUP patients analyzed (n = 123). For definitively solved CUP cases and those where WGS aided diagnostic resolution, biomarker-informed and non-biomarker-informed treatments represent tumor-type-specific therapies. In cases that were not fully solved, biomarker-informed and non-biomarker-informed treatments reflect treatments guided by tumor-type indications derived from WGS, even when some diagnostic uncertainty remained.
Source Data
For 73% (90 of 123) of CUP patients, at least one potentially actionable biomarker was identified (Fig. 2a,b). After demystification of the tissue of origin, 11% (14 of 123) had biomarkers linked to reimbursed treatments, and 69% (85 of 123) had biomarkers linked to an experimental biomarker-informed treatment. Although these rates are similar to those in previous literature27, we noticed that definitively solved CUPs harbor fewer standard-of-care biomarkers than cancers that already had a clear diagnosis. For NSCLC patients with a previous CUP history, only 5% of cases (1 of 23) had an actionable biomarker, whereas this was 61% (91 of 150) for the directly diagnosed NSCLC patients. This raises the question of whether the lack of standard-of-care actionable pathway is related to diagnostic uncertainty in general or defines a separate tumor-type subgroup.
The median TMB in the CUP cohort was 5.5 mutations per megabase (Fig. 2c,d and Supplementary Table 2a) and slightly higher than tumors with known primary origin.
Treatments in patients with CUP
Forty-seven percent (58 of 123) of CUP patients received a systemic treatment after WGS: 49% (38 of 77) following solved cases and 43% (20 of 46) following unsolved cases. However, more than one-third of CUP analyses (49 of 123) were done in the context of a second opinion and care after WGS diagnostics was mostly given in the referring hospital, with clinical follow-up not accessible for this study due to privacy regulations. When excluding 38 patients whose systemic treatment information was entirely unknown, we found that 64% (38 of 59) of solved cases received systemic treatment and 76% (20 of 26) of unsolved CUPs. Overall, 68% (58 of 85) of cases with known treatment data received a systemic treatment after WGS.
Most CUP patients were allocated to non-biomarker-informed standard-of-care treatments based on new WGS-derived diagnostic insights (46%; 39 of 85). A smaller proportion received biomarker-informed reimbursed treatments (7%; 6 of 85) or were enrolled in clinical trials for biomarker-informed experimental treatments (11%; 9 of 85). One patient was assigned to a non-biomarker-informed clinical trial, and three patients whose diagnoses remained unresolved received empirical CUP regimens (Fig. 3 and Supplementary Table 4a).
Although tissue-of-origin prediction was not a primary objective for the 600 patients with known primary tumors, the CUPPA analysis is part of the standardized WGS data analysis workflow for all patients. Interestingly, the algorithm provided added diagnostic value for 3% (20 of 600) of patients with established diagnoses, mainly resulting in improvements of soft tissue sarcoma subtyping. This had direct treatment consequences for 13 of 20 patients. Furthermore, for an additional six patients (1%; 6 of 600), a complete revision of an established diagnosis was made, with immediate treatment implications for one (Supplementary Table 4c).
Detection of PGVs
Finally, we analyzed the identification of clinically relevant PGVs that can be found because the WGS tests includes a normal blood sample as control for somatic variant detection. Consent for germline analysis was available for 96% of patients (692 of 723). Following updated national guidelines, PGVs were reported as present in the tumor without explicitly stating germline status43, which is why we performed this analysis in a post hoc setting. When a variant was identified as a potentially relevant PGV during MTB discussion, patients were referred for genetic counseling and germline analysis, leveraging pre-existing WGS data.
Overall, PGVs could be identified in 6.5% (45 of 692) of patients. About half of these variants had not been previously detected through routine diagnostics (23 of 45). Most showed a second somatic hit in the tumor (33 of 45). However, 12 did not show a second hit, and in one case, a confirmed pathogenic BRCA2 germline variant was identified where the somatic hit had been lost in the tumor. No patient had more than one relevant PGV (Extended Data Fig. 5 and Supplementary Table 5).
Overall clinical utility
To evaluate the overall utility of WGS in providing clinically valuable insights, we synthesized relevant results at a patient level: having a biomarker indicating reimbursed treatment, a WGS-based solution or improvement of a diagnosis, a clinically relevant PGV or a combination of these (Supplementary Table 6).
WGS provided clinically relevant results for 35% (211 of 600) with known primary tumor diagnoses (Fig. 4a). For patients with CUP, this amounted to 67% (83 of 123) (Fig. 4b). Overall, clinically valuable insights were obtained for 41% of all patients (294 of 723) who received WGS-based molecular testing.
Utility is synthesized (green) per investigated parameter (blue, reimbursed care biomarkers; orange, PGVs; purple, solved or clarified diagnosis). a,b, Six hundred patients with known primary tumor diagnoses (a) and 123 patients with CUP (b) are presented separately. Missing cells in the rows presenting PGV detection indicate patients who had indicated they did not want to be informed about potentially inherited variants.
Source Data
Improvement of OS
Finally, we investigated overall survival (OS) after WGS in patients with survival data (n = 719) (Supplementary Table 7). OS was improved in patients with at least one potentially actionable biomarker who received biomarker-informed therapy after WGS compared with those harboring an actionable alteration who did not receive such treatment (hazard ratio (HR) = 0.78; 95% confidence interval (CI) 0.63–0.96). The median increased from 309 to 405 days (+31%), favoring the treated group. Median OS of patients without actionable biomarkers did not differ significantly from either actionable-biomarker subgroup (Fig. 5a).
a, Kaplan–Meier curves comparing patients without potentially actionable biomarkers (Actionable−), patients with ≥1 potentially actionable biomarker who did not receive a biomarker-informed treatment after WGS (BIT−) and patients with ≥1 potentially actionable biomarker who did receive biomarker-informed therapy (BIT+). b, Within the actionable cohort, patients not receiving biomarker-informed therapy are split into those receiving only non-biomarker-informed systemic therapy (BIT−Rx+) and those without any post-WGS systemic therapy (BIT−Rx−), shown alongside patients who received biomarker-informed treatment (BIT+). c, Among patients with ≥1 potentially actionable biomarker, OS is stratified by the number of prior systemic therapy lines at the time of WGS: 0 (left), 1 (center) or ≥2 (right). Dotted lines indicate the median survival and are annotated with the corresponding number of days. HRs are reported with 95% CIs in parentheses. BIT, biomarker-informed treatment; Rx, non-biomarker-informed systemic treatment.
Source Data
In a subanalysis that split the group not receiving biomarker-informed therapy into patients receiving no treatment at all or receiving systemic therapies (Fig. 5b), biomarker-informed therapy remained superior to no therapy (HR = 0.60; 95% CI 0.47–0.78), with an almost threefold longer median OS (146 versus 405 days), whereas the difference versus non-biomarker-informed systemic therapy was smaller (+11%) and no longer significant.
When stratifying by the number of prior systemic treatment lines at the time of WGS (Fig. 5c), the OS benefit of biomarker-informed therapy was concentrated in patients without prior systemic therapy. These patients (median OS not reached after four years) outperformed those receiving only non-biomarker-informed systemic therapy (median OS 427 days; HR = 0.52; 95% CI 0.29–0.92) as well as those receiving no systemic therapy (median OS 214 days; HR = 0.37; 95% CI 0.21–0.64) despite having actionable biomarkers. By contrast, among patients with ≥1 prior systemic line, biomarker-informed therapy did not confer an OS advantage over non-biomarker-informed therapy.
These patterns were consistent irrespective of whether pre-WGS treatments had already included biomarker-informed therapy (Supplementary Fig. 2a–c). We also observed no significant OS differences between patients with CUP and known primary tumors (Supplementary Fig. 2d).
As these analyses represent real-world, non-randomized data, the prognostic and predictive effects of biomarkers for survival and treatment effects cannot be discriminated, and groups were not adjusted for baseline clinical condition (for example, performance status).