Study participants
A total of 540 participants were included in the full analysis set for clinical data. Participants’ characteristics are shown in Table 1. Participants with PD (n = 314) included both carriers (n = 128) and non-carriers (n = 186) of GBA1 variants (Appendix A). The individuals without PD included healthy controls non-carriers of GBA1 variants (n = 175, HC) and non-manifesting GBA1 variant carriers (n = 51, GBA-NMC). When age and sex of GBA-NMC were compared to the other groups, no differences were detected. Participants with PD were significantly older than HC (P= 0.006) and more frequently males (P = 0.028). More than half of the HC participants were partners of people with PD, to mitigate the effect of diet or other lifestyle-associated variables on gut microbiome composition.
Clinical profile of GBA-NMC
To identify possible clinical elements that could stratify GBA-NMC for their risk of developing PD, we compared the severity of motor and non-motor symptoms between HC and GBA-NMC using a wide range of clinical scales and questionnaires (Table 1). We found worse motor symptoms in the GBA-NMC group, either subjectively reported (Movement Disorder Society Unified Parkinson’s Disease Rating Scale (MDS-UPDRS) part II, P = 0.0008, q = 0.0076) or objectively assessed (MDS-UPDRS part III, P = 0.0153, q = 0.0627), with the former remaining significant even after adjustment for multiple testing. In terms of non-motor symptoms, no differences in constipation or global autonomic function were detected; however, GBA-NMC showed significantly higher scores in the MDS-UPDRS part I (P = 0.0036, q = 0.0228) and more severe urinary symptoms (P = 0.0198) and cognitive impairment (β = 0.92, P = 0.0197, odds ratio = 2.5, 95% confidence interval: 1.2−5.5), although these did not survive to multiple testing adjustment (both q = 0.0627). No differences in depression, anxiety or olfactory function were observed. Within the GBA-NMC, 10 individuals reached the threshold for estimated probability according to the MDS prodromal criteria calculated based on available information (listed in Methods)16.
Overall, our clinical data suggest that, in our GBA-NMC cohort, there may be a group of individuals who exhibit some prodromal symptoms (for example, subthreshold parkinsonism and some dysautonomia) and, thus, might be in their prodromal phase of PD. Additional biological markers indicating PD proximity are needed to better refine individualized risk of PD.
Significant microbiome alterations in PD
Microbiome profiles were successfully generated for 464 individuals (150 HC, 43 GBA-NMC and 271 PD). We first compared microbiomes of patients with PD, carriers (n = 109) or non-carriers (n = 162) of GBA1 variants, and found 44 Metagenomic Species Pan-genomes (MSPs) out of 627 with prevalence of at least 10% in our cohort that were different in abundance at P < 0.05 by two-sided Wilcoxon rank-sum test. However, none remained significantly different after Benjamini−Hochberg correction for multiple testing at q < 0.05 (or even q < 0.6). By contrast, the comparison of either group with HC (n = 150) revealed numerous MSPs significant at q < 0.05 (50 and 83 for carrier and non-carrier groups, respectively; the difference is likely due to the higher statistical power for the more numerous non-carrier group). β diversity analysis of Bray−Curtis distances by PERMANOVA confirmed that there was no significant microbiome difference between carrier and non-carrier groups (adonis P = 0.46), whereas the difference between HC and either group was very significant (P = 0.001). We conclude that, in overt PD, GBA1 genetic status impacts the microbiome composition much less than the disease itself.
We, therefore, pooled all patients with PD regardless of their genetic status (n = 271) to identify the PD gut microbiome signature at maximal statistical power and compared them with HC (n = 150). A total of 176 species were differentially abundant at P < 0.05 (Supplementary Table 1 and Extended Data Fig. 1a; 103 at q < 0.05). In PD, Actinobacteriota were enriched at phylum level and Bifidobacteriaceae at family level (Supplementary Table 2), as previously reported12,17,18. By contrast, species enriched in HC belonged to Lachnospiraceae C and Ruminococcaceae. These families include butyrate producers such as Roseburia and Dysosmobacter, which may be antiinflammatory. Bifidobacterium and Faecalibacterium genera were enriched in PD and HC, respectively. Enrichment of Bifidobacterium and depletion of butyrate producers such as Faecalibacterium in patients with PD were previously reported19,20. Species that showed the greatest increase in PD included Streptococcus mutans, Bifidobacterium longum, Bifidobacterium dentium and Lactobacillus paragasseri, whereas species that showed the greatest depletion in PD included Roseburia intestinalis, Roseburia inulinivorans and an unclassified Faecalibacterium. Further analyses were based on the 176 species differentially abundant at P < 0.05.
An intermediate and consistent microbiome alteration in GBA-NMC
We then investigated whether some of the PD-related gut microbiome alterations were detectable in the GBA-NMC group (n = 43) relative to HC (n = 150) by comparing species abundance in the two groups. A total of 43 species were significantly different at P < 0.05 (Supplementary Table 3 and Extended Data Fig. 1b), fewer than in the comparison of HC with PD, possibly, at least in part, because of the loss of statistical power due to the low number of GBA-NMC. Of these, 21 were common with the 176 species significantly altered in patients with PD compared to HC, and all but one were enriched or depleted in GBA-NMC individuals coherently with enrichment or depletion in patients with PD, using the direction of Cliff’s δ as a guide (positive or negative, respectively), which is unlikely to happen by chance (χ2 test, P = 5.7 × 10−3). The effect sizes observed for the common MSPs in the comparisons between GBA-NMC versus HC and PD versus HC were highly correlated (red dots, Fig. 1a).
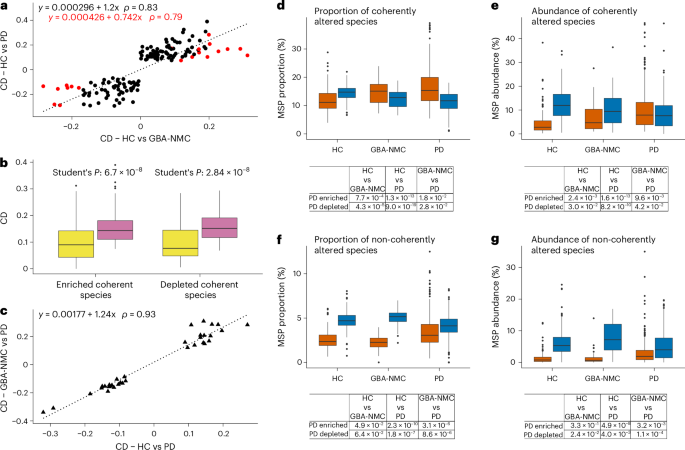
a, Correlations of Cliff’s δ (CD) of 142 coherent species found by comparing HC individuals with GBA-NMC or patients with PD. Negative CD values are species found enriched in HC, and positive CD values are species found enriched in both GBA-NMC and PD. Red dots highlight the 20 significant and coherently altered species (two-sided Wilcoxon P < 0.05) overlapping between HC−PD and HC−GBA-NMC comparisons. ρ denotes Spearmanʼs correlation coefficient when considering the 142 or 20 species. b, Average CD of coherently enriched (n = 81) or depleted (n = 61) species in GBA-NMC (yellow) and PD (mauve) relative to HC. c, Correlation of CD values of 34 species altered significantly (two-sided Wilcoxon P < 0.05) and non-coherently in the comparison of HC and patients with PD and in the comparison of PD and GBA-NMC. ρ denotes Spearmanʼs correlation coefficient. d,e, Abundance and proportion of coherent altered species in HC, GBA-NMC and PD and corresponding P values (determined by one-sided Studentʼs t-test for different comparisons); orange and blue refer to coherent enriched and depleted species, respectively. f,g, Abundance and proportion of non-coherent altered species in the three study groups and corresponding P values (determined by one-sided Studentʼs t-test); orange and blue refer to non-coherent enriched and depleted species, respectively. In the box plots, central line denotes the median, boxes the central quartiles, whiskers the extreme quartiles and dots the outliers.
Notably, using the direction of Cliff’s δ, examination of the coherence of variation of the 176 MSPs significantly different between HC and PD individuals revealed that 142 were coherently altered in GBA-NMC and PD relative to HC (Supplementary Table 4), which is very unlikely to happen by chance (χ2 test, P= 3.9 × 10−16). Of these, 81 MSPs were enriched and 61 were depleted in PD; we term these coherent enriched and coherent depleted species, respectively. Effect sizes of these species in GBA-NMC and PD relative to HC were highly correlated (Fig. 1a) and were, on average, significantly lower in GBA-NMC than in PD (Fig. 1b), suggesting that the GBA-NMC microbiome may be in an intermediate state between that of HC and PD. The remaining 34 species were found not to vary coherently in GBA-NMC and PD relative to HC and are hereafter termed non-coherent species; 18 were non-coherent enriched and 16 were non-coherent depleted in PD (Supplementary Table 4). Their Cliff’s δ values in GBA-NMC and HC relative to PD were highly correlated (Fig. 1c). Among the coherent species, we observed enrichment of oral residents (S. mutans and L. paragasseri) and proinflammatory Ruminococcus gnavus and the depletion of butyrate producers (Roseburia or Faecalibacterium prausnitzii). By contrast, non-coherent enriched species included Bifidobacteria, one of the characteristic alterations of the PD gut microbiome17, suggesting that their enrichment may take place at the clinical onset of the disease or develop during its course.
The species coherently altered in GBA-NMC and PD relative to HC represented a similar fraction (slightly over 25%) of the microbiome in all study groups, but the abundance and proportion of PD-enriched species significantly increased from HC over GBA-NMC to PD, whereas the abundance and proportion of PD-depleted species significantly decreased (Fig. 1d,e). The non-coherent species represented approximately 7–9% of microbiome abundance and 7% of species proportion. This part of the microbiome varied little between HC and GBA-NMC but very significantly increased and decreased for enriched and depleted species, respectively, in PD relative to HC or GBA-NMC (Fig. 1f,g), supporting the view that it evolves mostly once PD is manifest.
In sum, over a quarter of the gut microbiome significantly changes in PD relative to HC, of which we distinguished two components. A major one evolves consistently from HC across GBA-NMC to PD, the extent of changes being lower in GBA-NMC than in PD. A minor component changes abruptly in overt PD. Because similar coherent changes in gut microbiome composition are observed in at-risk individuals, such as GBA-NMC, and in patients with PD, we suggest that the coherently altered species of the PD gut microbiome may represent a prodromal feature of PD (the ‘prodromal-PD microbiome’), possibly contributing to PD development.
Associations of microbial alterations and clinical variables in PD
We estimated microbiome alterations by eight different tests, which capture related but different aspects of microbiome composition. The first four tests were based on abundance of different species types (coherent enriched or depleted; non-coherent enriched or depleted); the other four were based on proportion of the same species types. Abundance and proportion were computed as described in the Methods. Intuitively, an individual could have a higher proportion of coherent enriched species but a lower overall abundance of these species than another; the microbiome of the former would appear more altered than that of the latter by the test of proportion but less altered by the test of abundance. We ordered patients with PD by the fraction of the microbiome estimated by each of the eight tests and found that different quartiles vary greatly in all cases—Q1 and Q4 differed 10−20-fold by abundance and 2−3-fold by proportion (Extended Data Fig. 2a–d). This analysis shows considerable heterogeneity of microbiome composition among patients with PD.
We hypothesized that higher gut microbiome alterations were associated with worse clinical features in patients with PD. To test this hypothesis, we compared clinical variables of individuals from quartiles with highest (n = 68) and lowest (n = 68) microbiome alterations, estimated by each of the eight tests. Extended Data Table 1 shows the results from the comparison using the abundance of coherent depleted species as test of microbiome alterations; all comparisons are displayed in Supplementary Table 5. Some variables were significantly different in less than 25% of comparisons (≤2/8 comparisons; for example, age or body mass index (BMI)); we considered them to be either weakly correlated or even not correlated with microbiome alterations. By contrast, certain variables were significantly different in more than 75% of comparisons (≥6/8 comparisons), and we suggest that they are highly correlated with microbiome alterations (Fig. 2 and in bold in Extended Data Table 1). They fell in two classes.
Comparison of individuals with least (L) and more (M) altered microbiome across the study groups: for PD individuals, top and bottom quartiles of the distribution according to the abundance of coherent depleted species (n = 68 each); for GBA-NMC individuals, those below and above the median according to the proportion of coherent enriched species (n = 21 and n = 22, respectively); for HC individuals, top and bottom quartiles of the distribution according to the proportion of coherent enriched species (HC_L and HC_M, n = 38 each). HC individuals having PDMS-16 ≤ −5 (HC16_L, n = 21) and PDMS16 ≥ 3 (HC16_M, n = 27) are also shown. a, Disease-associated variables of patients with PD (disease duration and medication dosage). b, Clinical variables for all groups (automatic dysfunction, constipation, depression and non-motor symptoms). c, Health-related variables for all groups (DQS; appendectomy colored in gray if individuals did not undergo appendectomy and colored in black if individuals underwent appendectomy). The values of clinical variables are the means listed in Extended Data Table 1. Standard deviations and statistical significance are indicated by the thin bars, and stars indicate the significance level of the one-sided Studentʼs t-test for each comparison, except for appendectomy where comparisons are tested using two-sided χ2 test. Significance of P values: NS (not significant): P ≥ 0.05; *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. The exact P values are listed in Extended Data Table 1. BDI, Beck Depression Inventory; SCOPA-AUT, Scales for Outcomes in Parkinsonʼs Disease-Autonomic Dysfunction.
The first class corresponded to variables associated with disease severity, found to be higher in individuals in the top quartile of microbiome alterations, in agreement with our hypothesis. Among these, we detected depression, autonomic dysfunction, constipation and motor dysfunction. A higher percentage of individuals who underwent appendectomy prior to PD development was found in the top quartile; association of appendectomy and PD has been reported but remains controversial21,22,23,24.
The second class included variables associated with health, found to be worse in individuals with more altered microbiome, also in agreement with our hypothesis. These variables included cognitive and olfactory functions. Interestingly, variables related to nutrition, such as the Dietary Quality Score (DQS) or fruit and vegetable consumption, were also worse, suggesting a possible relation between food quality and milder disease profile.
We noted that the two most significantly differing variables between top and bottom quartiles were PD duration and levodopa equivalent daily dose (LEDD) (that is, the amount of antiparkinsonian medications), both associated with disease severity (Extended Data Table 1). We thus compared the microbiome of patients with PD medicated or drug naive, adjusting for disease duration, and the microbiome of patients with PD with different disease duration, adjusting for medication dose (Extended Data Fig. 3a,b). No significant differences were observed in the level of PD-enriched or PD-depleted species in medicated and non-medicated patients with the same disease duration, whereas significant differences were observed in patients with the same medication dose and different disease duration. We suggest that the microbiome evolves with the progression of the disease rather than in response to treatment.
Associations of microbial alterations and clinical variables in disease-free GBA-NMC and HC
To investigate whether the PD microbiome signature was associated with any prodromal clinical profile in GBA-NMC, we compared clinical variables of GBA-NMC individuals with lower and higher microbiome alterations, below or above the median of the abundance and proportion of coherent PD-enriched/depleted species; expectedly, non-coherent species, which change significantly in abundance only in overt disease, were not informative (Extended Data Fig. 4; split around median rather than comparison of quartiles was used to conserve statistical power). We used the threshold of 75% comparisons, also used in patients with PD, to define variables highly associated with microbiome alterations. We identified numerous significantly different parameters, mainly representative of non-motor symptoms, which can be present in the prodromal phase (Fig. 2, Extended Data Table 1 and Supplementary Table 6). Among these, scores representing motor and non-motor disability and autonomic dysfunction were all significantly higher in individuals showing microbiome alterations above the median. Notably, all GBA-NMC individuals who were identified as prodromal PD, based on the MDS research criteria16, had the abundance of coherent PD-enriched species above the median, a highly significant bias (χ2 test, P = 4.7 × 10−3), and four were at the very top of the distribution (Fig. 3; P = 5.7 × 10−4). We conclude that GBA-NMC individuals presenting with the microbiome signature closer to PD are the ones showing the strongest prodromal profile, especially in terms of autonomic and peripheral nervous system dysfunction (body-first PD). Paralleling the observation with patients with PD, where the duration of overt disease appears to be strongly associated with microbiome alterations, we suggest that, in the GBA-NMC group, the progression toward overt PD is associated with progressive microbiome alterations and that a combination of clinical and microbiome assessments may help identify those individuals who will convert to disease.
Individuals assigned to the prodromal group by the MDS research criteria (n = 8) are represented by filled dots.
We then investigated microbiome alterations across quartiles of distribution in healthy individuals, using the same four tests based on coherent species (depleted and enriched; abundance and proportion), and found very significant differences, resembling those observed for patients with PD, albeit with lower values for the PD-enriched species and higher for the PD-depleted species in the extreme quartiles (Extended Data Fig. 5). Interestingly and notably, the comparison of clinical variables of HC individuals assigned to the extreme quartiles, denoted as lower (n = 38) and higher (n = 38) microbiome alterations for analogy with the PD and GBA-NMC groups (Fig. 2, Extended Data Table 1 and Supplementary Table 7), revealed significant differences of most variables differing in the GBA-NMC group, including depression and global motor and non-motor disability. Furthermore, these HC differed in eating habits (DQS and fruit and vegetable consumption) and appendectomy frequency, as observed for patients with PD. We conclude that microbiome alterations resembling those of patients with PD take place in a healthy population free of GBA1 genetic risk, raising the possibility that, as in the individuals with GBA1 variants, such alterations might be associated with progression toward overt disease. Of note is that, for both GBA-NMC and HC, one of the tests, based on the abundance of coherent species, revealed fewer differences than the other three (Supplementary Tables 6 and 7). We do not have an explanation for this observation, which indicates the value of using multiple tests to assess microbiome alterations, and we suggest that clinical variables significantly different by at least two of the other three tests correlate well with microbiome alterations in disease-free individuals (as reported in Extended Data Table 1).
Correlation of microbial species and clinical variables
We next investigated whether the species that compose the altered microbiome might individually be correlated with clinical variables, focusing on the 176 species differentially abundant in HC and PD. In the PD group, abundance variations of 94% of species were significantly (P < 0.05) correlated with at least one clinical variable (Supplementary Fig. 1a and Supplementary Table 8); the number of species correlated with a variable is shown in Fig. 4a. PD-enriched species and PD-depleted species were associated positively and negatively, respectively, with disease severity (for example, non-motor symptoms such as constipation, autonomic dysfunction, depression and RBD) and, conversely, negatively and positively with health (cognition and olfactory function and DQS score).
a−c, Number of microbial species significantly (Spearmanʼs correlation coefficient P < 0.05) correlated with different clinical parameters is indicated on the ordinate for patients with PD (a), GBA-NMC participants (b) and HC individuals (c). Orange and blue colors refer to species enriched and depleted in PD; positive and negative integers refer to positive and negative correlations, respectively. d−f, Number of functional modules correlated significantly (Spearmanʼs correlation coefficient P < 0.05) with different clinical parameters is indicated on the ordinate for patients with PD (d), GBA-NMC participants (e) and HC individuals (f). Orange and blue colors refer to modules enriched and depleted in PD; positive and negative integers refer to positive and negative correlations, respectively. Stars indicate the significance of the correlation bias of enriched and depleted species (two-sided χ2 goodness-of-fit test) on the positive and negative half of the graph, respectively. Significance of P values: *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. BDI, Beck Depression Inventory; HADS, Hospital Anxiety and Depression Scale; SCOPA-AUT, Scales for Outcomes in Parkinsonʼs Disease-Autonomic Dysfunction.
Notably, we found a similar profile of correlations between species and clinical variables in GBA-NMC (Fig. 4b), with variables associated with severity being mostly positively correlated with species enriched in PD and negatively with species depleted in PD (Supplementary Fig. 1b); the converse was observed for variables associated with health. However, the correlation bias was less clear than in the PD group (for example, cognition or constipation—Montreal Cognitive Assessment (MOCA) or Wexner Constipation Scoring System (WCSS), respectively), possibly because of lower statistical power due to the smaller group size and/or lower variance of the variables in the GBA-NMC group. The correlations involved 64% of the 176 species (Supplementary Table 9). Some 15% of correlations were between the same species and clinical variables as in patients with PD (Supplementary Table 10), and an overwhelming majority of those (39/42) had the same direction in both groups (Extended Data Fig. 6a). Other correlations were different, possibly due to relatively small variations of species abundances within a study group, resulting in a statistical significance in one group but not another.
Similar correlations were found in healthy individuals (Fig. 4c and Supplementary Table 11). The correlation bias was less clear than in the PD group, as observed in the GBA-NMC group, possibly for the same reasons. The correlations involved 77% of the 176 species, with 24% of them being between the same species and clinical variables as in the PD group (Supplementary Table 12) and showing the same direction in both groups (75/77; Extended Data Fig. 6b).
A notable conclusion from our analyses is that the microbial species correlated with disease severity in patients with PD appear correlated with clinical variables that estimate disease severity both in healthy individuals at higher risk of PD development due to their GBA1 genetic background and in those without known GBA1 genetic predisposition.
Stratification of HC individuals using microbiome features
In view of similar correlations between clinical variables and microbial species in all study groups, we explored the possibility of stratifying HC individuals who may be closer to disease development by the microbial species they harbor. To do so, we first selected the species involved in the same significant correlations with clinical variables in PD and the disease-free GBA-NMC (33 species). Among those, we then selected species with the highest prevalence difference in PD relative to HC (over 10%), assuming that they should be the best indicators of disease-related microbiome alterations, and the species correlated coherently with PD duration in patients, hypothesizing that they indicate alterations of the microbiome due to progression of disease in patients and, thus, possibly approaching the disease in HC. As a result, we identified 16 species, of which 10 were enriched and six were depleted in PD (Supplementary Table 13). By summing the number of PD-enriched species and deducing the number of PD-depleted species that each individual harbored, we computed a score, termed Parkinsonʼs Disease Microbiome Score-16 (PDMS-16), referring to the number of species used. PDMS-16 distribution among HC and PD individuals is shown in Extended Data Fig. 7.
We then compared clinical variables between the subgroup of HC presenting with the highest PDMS-16 (3−7; n = 27, 18% of the group) with (1) 21 individuals with the lowest PDMS-16 score (−5 and −6) and (2) the remainder of the HC group (n = 123). The individuals with the highest PDMS-16 score had more severe depression and anxiety, autonomic dysfunction and constipation, consumed less healthy food and had a more frequent history of appendectomy (Fig. 2 and Extended Data Table 1). We conclude that the PDMS-16 score can identify individuals with a clinical profile closer to that of patients with PD than the remainder of the HC group in our study population, and we suggest that, as proof of concept, it could be a lead for developing a broadly applicable, species-based score of microbiome alterations indicative of PD risk in disease-free individuals, when much larger, adequately phenotyped cohorts become available.
Alterations of microbiome functions in PD and GBA-NMC
Comparison of HC with patients with PD revealed that 146 of the 357 complete functional microbial modules from three sources (Kyoto Encyclopedia of Genes and Genomes (KEGG), Gut Metabolic Modules (GMM) and Gut Brain Modules (GBM)) were differentially abundant at q < 0.05: 121 were enriched in PD and 25 were depleted (Supplementary Table 14). The PD microbiome was enriched for modules related to neurotransmitters metabolism (dopamine, acetylcholine and GABA), nucleic acid degradation and amino acid degradation (threonine, phenylalanine and putrescine). Enrichment of the modules for dopamine degradation and DOPAC synthesis (MGB023 and MGB024, respectively) could possibly be due to levodopa treatment of patients with PD, as Actinobacteria reportedly can degrade and use dopamine18. The degradation pathway of the bacterial byproduct of protein degradation, p-cresol, was also found to be enriched in our PD cohort. The modules depleted in PD were involved in riboflavin biosynthesis, dietary carbohydrate degradation (lactose and mannose) pathways and the conversion of acetyl-coenzyme A into acetate pathway. These findings, combined with the enrichment in pathways implicated in the conversion of acetate into acetyl-coenzyme A, or of propionate into succinate, and the tricarboxylic acid cycle, support a metabolic shift from carbohydrate fermentation toward proteolysis as an energy source, in line with previous findings17,20.
The comparison of HC and GBA-NMC individuals identified only six significantly different modules at P < 0.05. This low number may reflect, in part, fewer alterations of functions in the GBA-NMC group and, in part, insufficient statistical power. We, therefore, examined the coherence of Cliffʼs δ variation of the modules significantly different in HC and PD in the GBA-NMC group. Of the 180 modules significant at P < 0.05, 143 varied coherently and 37 varied non-coherently (Supplementary Table 14), a very significant bias (χ2 test, P = 2.8 × 10−15). Cliffʼs δ values of the coherently varying modules were significantly correlated (Extended Data Fig. 8a) and significantly higher in the HC versus PD comparison than in the HC versus GBA-NMC comparison (Extended Data Fig. 8b).
We conclude that functional potential of the microbiome displays variations like those observed for the taxonomic differences.
Correlations of microbiome functions and clinical variables
Abundance variations of 92% of modules were significantly correlated with at least one of 25 clinical variables in the PD group (Supplementary Table 15). Correlations were biased in a similar way observed for individual species: enriched modules largely correlated positively with variables related to the severity of the disease and negatively to those related to health, whereas the depleted modules had an opposite bias (Fig. 4d).
Similar correlation profiles were observed for the other two study groups, GBA-NMC and HC (Fig. 4e,f and Supplementary Tables 16 and 17), albeit with a lower number of modules, likely due to lower statistical power (58.3% and 72.5% for GBA-NMC and HC, respectively).
We conclude that, expectedly, functional potential of the microbiome correlates with clinical variables similarly to species that encode it.
Validation of microbiome alterations in independent cohorts
We examined three different publicly available cohorts composed of patients with PD and healthy individuals where stool microbiome was analyzed by shotgun metagenomics. One was from the United States (PD, n = 491; HC, n = 234)12, one from Turkey (PD, n = 69; HC, n = 17)25 and one from Korea (PD, n = 78; HC, n = 68)26. There were n = 498, n = 606 and n = 572 metagenomic species, respectively, present in at least 10% of the cohort individuals (Supplementary Tables 18−20), of which the large majority was also present in our cohort (n = 408, 81.9%; n = 474, 78.2%; and n = 448. 78.3%, respectively). By abundance, these common species represented 94.2%, 78.7% and 87.8% of the microbiome, respectively.
Cliffʼs δ values computed for the HC/PD comparisons in each of the three cohorts were well correlated with the Cliffʼs δ values observed in our cohort, for all species common between a given cohort and our cohort (Fig. 5a–c). The correlations were higher for species significantly different in our cohort only (P < 0.05; Fig. 5d−f) and raised even further for species significantly different in both our cohort and another given cohort (Fig. 5g–i). Furthermore, the direction of Cliffʼ δ values for species significant in both cohorts was fully conserved in two cohorts (United States and Turkey) and 93.3% conserved in one cohort (Korea). This indicates that similar microbiome alterations take place in PD worldwide, given the geographic diversity of the cohorts studied, and that they extend to species that do not appear significantly different (P > 0.05).
a,d,g, United States cohort. b,e,h, Korean cohort. c,f,i, Turkish cohort. Top panels: all species. Middle panels: 176 species significant in our study (two-sided Wilcoxon P < 0.05). Bottom panels: species significant in both cohorts (two-sided Wilcoxon P< 0.05). Wilcoxon values are displayed in Supplementary Tables 1 and 18–20. ρ denotes Spearmanʼs correlation coefficient. CD, Cliffʼs δ; Sign., significant.
We next examined the intragroup heterogeneity of the microbiome in the three cohorts, using as lead the coherent PD-enriched and PD-depleted species defined in our cohort. Heterogeneity like the one we observed in our cohort was found in all other cohorts (Extended Data Fig. 9), indicating that, among both patients and healthy individuals, some have more altered microbiomes than others. These observations largely validate conclusions from our cohort analysis.
Clinical phenotypes were available only for the Turkish cohort (Supplementary Table 21), where microbiome profiles of sufficient quality were generated for 69 PD individuals and for 17 HC individuals with our pipeline, using the published sequencing reads. We first analyzed the patients with PD, comparing the clinical variables of the top and bottom terciles with least and most altered microbiome (n = 23 each; Supplementary Table 22). Significant differences were observed for most variables (17/24; 71%) in at least some tests; the highest number of variables (12/24, 50%) was found using abundance of depleted non-coherent species. The disease duration was significantly different (P < 0.05) in seven of eight tests, chiming with the observations in our cohort. Two other variables, the Neuropsychiatric Inventory (NPI-severity) and Wechsler Memory Scale-Visual Reproduction (WMS-VR_Short-Term_Memory), were also significantly different in the large majority (six or more) of tests. Other variables were significantly different in fewer tests, possibly because of the lower statistical power than in our cohort, but their tendency was in the expected direction—variables associated with disease severity were higher for individuals with more altered microbiome, whereas variables associated with health were lower. We conclude that the association of clinical parameters and microbiome alterations in PD that we describe for our cohort is largely validated in an independent cohort.
Analysis of the HC group (n = 17), split around the median, yielded few clear results (Supplementary Table 23), likely because of a small group size and the limited nature of clinical data available (cognitive and neuropsychiatric scales and not other recognized prodromal features of PD). We conclude that fuller validation of our observations for disease-free individuals may have to await larger, appropriately phenotyped cohorts.