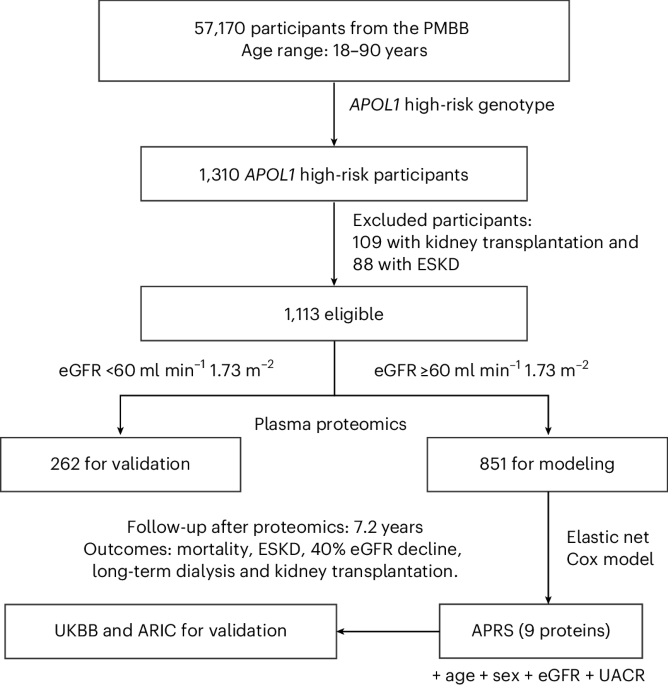
PMBB cohort
The PMBB is a large academic biobank that recruits participants from the University of Pennsylvania Health System, with recruitment beginning in 2008. To date, the PMBB has enrolled over 250,000 participants—approximately 30% of whom are from non-European ancestries—and around 57,170 individuals have undergone whole-exome sequencing41. Demographic information, medical history (via International Classification of Disease (ICD) codes), medication use and clinical assessments were extracted from the EHR. Laboratory tests, including serum creatinine, blood urea nitrogen and electrolyte levels at baseline, were also obtained. The eGFR was calculated using the creatinine-based 2021 Chronic Kidney Disease Epidemiology Collaboration (CKD-EPI) equation42.
All participants provided written informed consent for the use of their biospecimens, genetic data and EHR data for research. Genomic DNA samples were transferred to the Regeneron Genetics Center and stored at –80 °C until sample preparation. Whole-exome sequencing was performed with reads mapped to Genome Reference Consortium Build 38 (GRCh38); samples failing quality metrics (for example, low sequencing coverage) were excluded, as described previously. Ancestry was estimated by exome data using principal component analysis. A set of high-quality, common single-nucleotide polymorphisms overlapping with HapMap3 was extracted. Principal components were first calculated for HapMap3 samples and then used as the reference space, onto which all study samples were projected. To classify ancestry, a kernel density estimation approach was applied to the joint distributions of the first four principal components for each HapMap3 population. For each sample, normalized likelihoods of belonging to each reference population were obtained. Reference populations were then assigned if likelihoods exceeded prespecified thresholds. Based on these assignments, samples were grouped into one of the following ancestral classes: African, European, East Asian, South Asian or Admixed American. Subsequently, the APOL1 was analyzed in all African ancestry participants with available exome sequence data41.
Individuals were classified into high-risk APOL1 genotype groups based on the presence of the G1 (G1a and G1b) and G2 risk alleles. Participants with two risk alleles (G1/G1, G2/G2 or G1/G2) were classified as high-risk11,12, whereas those with zero or one risk allele (G0/G0, G0/G1 or G0/G2) were classified as low-risk. We included all participants aged ≥18 years with available high-risk APOL1 genotype and at least one subsequent follow-up record or documented clinical event (for example, mortality and dialysis). Exclusion criteria included known ESKD or with kidney transplantation at baseline, missing or ambiguous APOL1 data and excessive missing data in key clinical covariates. To minimize reverse causation, clinical diagnoses (for example, hypertension, diabetes and cardiovascular disease) were required to have been recorded in the EHR prior to the blood draw. Laboratory values used as baseline covariates were extracted from tests performed within a prespecified window of 2 months before to 1 month after the plasma draw; for each participant, we used the single measurement closest to the draw date to reduce temporal misalignment. For UACR, we used the most recent measurement obtained within the 2 years preceding the plasma draw. When only a urine protein–creatinine ratio or a dipstick protein result was available, we converted these to estimated UACR using the conversion equations from ref. 43. This work was conducted under University of Pennsylvania Institutional Review Board (IRB)-approved protocols (815796, 813913, 855821 and 857403; study protocol).
Outcomes and follow-up
The primary outcomes were defined as a composite of kidney events (long-term dialysis, ESKD diagnosis, kidney transplantation or a ≥40% decline in eGFR from baseline) and all-cause mortality. Mortality was included given its clinical importance and as a critical competing risk for kidney failure44. For APOL1 high-risk individuals with eGFR ≥60 ml min−1 1.73 m−2, the mean follow-up duration was 7.18 ± 2.89 years, with a mean time to event of 6.51 ± 3.07 years. Outcome data were obtained through medical record review and ICD-10 codes. Participants were censored at the time of death, loss to follow-up or the end of a 10-year follow-up period, whichever occurred first.
Proteomic profiling
PMBB plasma samples were collected at baseline and subjected to proteomic profiling using the SomaScan platform (SomaLogic), which employs modified aptamers (SOMAmers) to probe the human proteome22. SomaScan v.4.1 pools 7,524 SOMAmers to probe 6,386 distinct proteins (https://menu.somalogic.com/). The SomaScan assay is a highly multiplexed, sensitive and reproducible proteomic technology that has been extensively validated and used in numerous clinical studies23,30,31,32,33,45,46,47,48. The SomaScan assay is based on the use of modified DNA aptamers, called SOMAmers, which are designed to bind specific protein targets with high affinity and specificity22. Each SOMAmer is uniquely tagged with a DNA barcode that allows for quantification using a custom DNA microarray. In brief, the diluted plasma samples were incubated with a pool of SOMAmers, and, subsequently, the SOMAmer−protein complexes were captured on streptavidin-coated beads, washing away unbound proteins and SOMAmers. The bound SOMAmers were then eluted and hybridized to a custom DNA microarray containing complementary sequences to the SOMAmer barcodes. The microarrays were scanned using a SureScan Dx Microarray Scanner (Agilent Technologies), and the fluorescence intensity of each SOMAmer was quantified as a measure of the relative abundance of its corresponding protein target.
Raw data were processed using SomaScan Data Analysis Software (SomaLogic) to generate relative fluorescence units (RFUs) for each SOMAmer. The RFUs were then normalized using a set of internal calibrator samples to adjust for any assay-specific biases and to ensure comparability across different plates and runs. Rigorous quality control measures were implemented throughout the proteomic profiling process to ensure data integrity and reliability. These included the use of internal calibrator samples, quality control metrics for sample and assay performance and the exclusion of any samples or SOMAmers that failed to meet predefined quality criteria49. The log2-transformed RFUs were then used for subsequent statistical analyses and model development.
UKBB cohort
The UKBB is a large-scale, community-based cohort comprising over 500,000 participants aged 40–69 years at recruitment (2006–2010) from 22 assessment centers across the United Kingdom. For validation purposes, we included all participants who underwent plasma proteomic profiling50. This subset is representative of the overall UKBB population. Among these, 1,171 individuals of African descent were identified—204 with APOL1 high-risk. APOL1 genotypes were imputed (using Data-Field 21007) based on the TOPMed R2 reference panel after phasing with Eagle v.2.4 and converting from GRCh37 to GRCh38 via LiftOver51. ESKD was identified using data_coding_19 codes N185 and N180 in fields 41270 and 41280 and using READV3_CODE mapped to ICD-10 codes N185 and N180 in Table 1060; hypertension was captured using data_coding_19 with the regular expression ‘I1[012345]’ in fields 41270 and 41280 and using READV3_CODE mapped to ICD-10 with the same expression in Table 1060; diabetes was defined using data_coding_19 with the regular expression ‘E1[01234]’ in fields 41270 and 41280 and using READV3_CODE mapped to ICD-10 with the same expression in Table 1060; kidney transplant history was ascertained using data_coding_19 code Z940 in fields 41270 and 41280 and using READV3_CODE mapped to ICD-10 code Z940 in Table 1060; dialysis treatment was identified through procedure_concept entries matching ‘[Dd]ialysis’ in Table 936 and TERMV3_DESC entries matching ‘[Dd]ialysis’ in Table 1060; mortality outcomes were obtained from death register data in Table 1058; serum creatinine measurements were drawn from measurement_concept_id 37392176 and 3020564 in Table 931, from repeat-measure identifiers p30700_i.* and p23478_i.* in field 30700 and from TERMV3_DESC entries matching ‘[Ss]erum [Cc]reatinine$’ in Table 1060; and cystatin C was obtained via measurement_concept_id 3030366 in Table 931. Olink measurements were log2 transformed and normalized to harmonize scales. Missing proteins in Olink were imputed by mapping overlapping proteins to the SomaScan scale and predicting absent targets with a multi-output penalized regression trained on SomaScan data (statistical analysis protocol). This study was conducted under application number 273810. The validation from the UKBB for this project was approved by the University of Pennsylvania IRB (protocol 855821).
ARIC study
The ARIC study enrolled 15,792 participants (aged 45–65 years) from four US communities (Washington County, Maryland; Forsyth County, North Carolina; Jackson, Mississippi; and Minneapolis, Minnesota) between 1987 and 1989, with follow-up visits every 3 years initially and subsequently at varying intervals52. We included 314 African American participants with APOL1 high-risk genotyping (performed using TaqMan assays for G1 and G2)8,53 who attended visit 2 (approximately 3 years after baseline) with eGFR≥60 ml min−1 1.73 m−2; proteomic profiling on these visit 2 biospecimens was performed using the SomaScan 5K assay. To ensure comparability with the PMBB, we truncated follow-up at 10 years from baseline. This study was conducted under application number MP4524. The validation from the ARIC for this project was approved by the University of Pennsylvania IRB (protocol 855821). All validation analyses were independently performed by statisticians from different institutions.
Human kidney samples
Kidney tissue samples (n = 474 for RNA-seq and n = 325 for proteomics) for this study were procured from surgical nephrectomies, ensuring that only the normal parts of the tissue, specifically those at least 2 cm from any cancerous lesions, were used for analysis. An honest broker deidentified the samples and collected corresponding clinical information, such as age, race, sex and diabetes and hypertension status, in addition to creatinine values. The eGFR was subsequently determined using the latest CKD-EPI equations42. Kidney samples were formalin fixed, paraffin embedded and stained with periodic acid–Schiff. Whole-tissue imaging was conducted using the Aperio system, which is a platform that digitizes and assists in analyzing pathology slides. Samples were scored in an unbiased manner by a specialized renal pathologist23,25,26. The use of these samples and data was approved by the University of Pennsylvania IRB under the category of ‘exempt’, negating the need for informed consent due to the deidentified nature of the study samples.
Kidney tissue RNA-seq and data processing
RNA isolation, sequencing and analysis were performed as previously published27. Total RNA was isolated from kidney tissue using the RNeasy Mini Kit (Qiagen) according to the manufacturer’s instructions, including the DNase digestion step. RNA quality was assessed by Agilent Bioanalyzer 2100. The cDNA library was prepared using NEBNext Ultra II RNA Library Prep Kit for Illumina. Then, cDNA libraries were sequenced on an Illumina NovaSeq 6000 platform using the NovaSeq PE150 protocol. Adaptor and lower-quality bases were trimmed with TrimGalore (v.0.4.5). Reads were aligned to the human genome (hg19) using STAR (v.2.7.3a). Gene and isoform expression levels of transcripts per million were estimated using RSEM (v.1.3.0).
Kidney tissue proteomics and data processing
Kidney tissue samples were snap frozen, cryopulverized (CryoMill; Retsch) and lysed using T-PER extraction reagent supplemented with protease inhibitors (Roche Diagnostics). Protein concentrations were determined via bicinchoninic acid assay (Thermo Fisher Scientific). Similar to plasma proteomics, the tissue proteomic profiling was performed using the SomaScan v.4.1 platform (SomaLogic), which uses slow off-rate modified DNA aptamers (SOMAmers) to quantify protein targets. To ensure data consistency, quality control measures included hybridization controls, pooled calibrators and buffer-only replicates for monitoring background signals and batch effects. Data underwent normalization to correct for within-run hybridization variability, followed by intrastudy median normalization.
PRS
A previously published genome-wide PRS for CKD29 was applied. For each individual, the PRS was calculated as the weighted sum of risk alleles, with weights corresponding to the effect sizes reported in the original study. In brief, the PRS was computed as \(\mathrm{PRS}={\sum }_{i=1}^{M}{\beta }_{i}\times {\mathrm{dosage}}_{i}\), where M is the number of variants with non-zero weights and βi is the effect size for variant i. To enable genome-wide PRS computation, we used imputed genotype data from the PMBB (Release 2.0), which includes participants genotyped using the Illumina Global Screening Array (Freeze 2.0). Genotypes were phased with Eagle2 and imputed using Minimac4 on the Michigan TOPMed r3 reference panel (GRCh38). The imputed dataset was filtered to retain high-confidence variants (average R2 > 0.3 or directly genotyped in either batch) and minor allele frequency > 0.01, ensuring compatibility with published PRS weights. PRS values were computed in PLINK 2.0 using the overlapping variant set between the published PRS and the PMBB-imputed genotypes. The PRS analysis was included primarily as a benchmark to contextualize the incremental predictive value of the APRS.
CRIC proteomics model
For benchmarking, we applied the previously published CRIC proteomic risk score, which was developed using SomaScan profiling of 65 plasma proteins in patients with established CKD28. The original coefficients were applied to our SomaScan data after log2 transformation and harmonization of units. For each participant, a CRIC score was calculated as the weighted sum of these proteins. The performance of the CRIC score was evaluated using tAUC and C-index for the composite outcome in the PMBB.
Marker selection and model construction
To identify the optimal combination of proteins for outcome prediction, we implemented an elastic net penalized Cox proportional hazards modeling framework based on the random generation of candidate protein panels. This strategy is conceptually related to random subspace approaches54 and ensemble feature selection methods55,56 but differs in that resampling is performed on the feature space rather than on individuals, and each candidate protein panel is evaluated independently. This was implemented through the following sequential steps: (1) initial protein screening and candidate panel generation by identifying proteins showing significant univariate associations (Benjamini−Hochberg-adjusted P < 0.05) with the outcome (top 20%) and then randomly sampling 40–90% of these proteins without replacement for each candidate panel; (2) elastic net modeling with grid search, fitting an elastic net Cox model for each sampled protein panel and conducting a comprehensive grid search across the mixing parameter α (ranging from 0.1 to 0.9 in 0.1 increments) and the regularization strength λ (spanning 100 logarithmically spaced values); (3) performance evaluation via eight-fold cross-validation using the mean C-index and large-scale combinatorial exploration of approximately one million unique candidate protein panels to thoroughly explore the vast combinatorial space of protein interactions and identify the optimal combination; (4) intermediate feature selection by stability, retaining proteins with a selection frequency of at least 30% across candidate panels to avoid multiple testing problems and reduce noise, ensuring that only consistently predictive proteins were retained; and (5) final model construction and refinement using the intermediate protein set with another round of elastic net penalized Cox proportional hazards modeling to eliminate highly correlated proteins and optimize model parameters based on the most promising candidates identified in the initial screening and evaluation process. The final proteomic signature was defined as the panel of proteins and corresponding penalty parameters that achieved the highest average cross-validated C-index across all iterations.
The model risk score is: \({\mathrm{score}}=\exp \left({\sum }_{j}{\beta }_{j}\times {x}_{\!j}\left(t\right)\right)\), where \(({{\beta }}_{j})\) are the coefficients and \(({x}_{\!j}\left(t\right))\) are the protein markers. The APRS formula is:
$$\begin{array}{l}\hat{y}({t|}{\mathbf{x}})=\exp \left(1.72\times {\mathrm{SPON1}}+1.18\times {\mathrm{SUMO2}}+1.08\times {\mathrm{EPHA}}10 \right.\\ \,\,\,\,\,\,\,\,\,\,\,\,+0.89\times {\mathrm{REG3A}}+1.37\times {\mathrm{WFDC2}} + 1.56 \times {\mathrm{LYZ}} + 0.68 \times {\mathrm{MMP7}} \\ \,\,\,\,\,\,\,\,\,\,\,\,+0.55\times {\mathrm{NPPB}}-0.95\times {\mathrm{CILP2}}+0.22\times {\mathrm{UACR}}_{{\mathrm{per}}\,{\rm{doubling}}}\\ \,\,\,\,\,\,\,\,\,\,\,\, \left. +0.19\times {\mathrm{Age}}_{{\mathrm{per}}\,10\,{\mathrm{years}}}-0.27\times {\mathrm{eGFR}}_{{\mathrm{per}}\,5\,{\mathrm{mL}}\,{\mathrm{min}}^{-1}\,1.73\,{\mathrm{m}}^{-2}}+0.49\times {\mathrm{Male}}\right)\end{array}$$
Model performance evaluation
We evaluated the performance of the predictive model using a comprehensive set of metrics designed for survival analysis. The evaluation function takes as input the true survival outcomes from the training and test sets, along with the predicted risk scores and the timepoints at which the predictions are made. The function first prepares the data by aligning the predicted risk scores with the true survival outcomes and applying inverse probability of censoring weights (IPCW) to account for censoring in the test set. The IPCW for each sample i at time \(t\) is calculated as: \({\mathrm{IPCW}}_{i,t} = \frac{1}{\hat{S}\left(t \mid {\mathbf{X}}_i\right)}\) where \(\hat{S}\bigl(t \mid {\mathbf{X}}_i\bigr)\) is the estimated probability of being uncensored at time \(t\) given the covariates \({\mathbf{X}}_{i}\). The samples are then sorted by descending risk score at each timepoint. Let \(n\) be the number of samples and m be the number of timepoints. For each timepoint \(t\), we define: \({\rm{TP}}_{t}\): true positives, \({\rm {FP}}_{t}\): false positives, \({\rm {TN}}_{t}\): true negatives, \({\rm {FN}}_{t}\): false negatives. Next, the function calculates various performance metrics at each timepoint, including: AUC: \({\mathrm{AUC}}_{t}={\int }_{0}^{1}{\rm {TPR}}_{t}\left({\rm {FPR}}_{t}\right),{\rm {dFPR}}_{t}\) where \({\rm {TPR}}_{t}\) is the true-positive rate (sensitivity) and \({\rm {FPR}}_{t}\) is the false-positive rate (1 − specificity) at time \(t\). Model sensitivity, specificity F1 score, accuracy, Matthews correlation coefficient (MCC), positive predictive value (PPV) and negative predictive value (NPV) were calculated as: \({\rm {Sensitivity}}_{t}=\frac{{\rm {TP}}_{t}}{{\rm {TP}}_{t}+{\rm {FN}}_{t}}\), \({\rm {Specificity}}_{t}=\frac{{\rm {TN}}_{t}}{{\rm {TN}}_{t}+{\rm {FP}}_{t}}\), F1t\(=\frac{2\times {\rm {Precision}}_{t}\times {\rm {Recall}}_{t}}{{\rm {Precision}}_{t}+{\rm {Recall}}_{t}}\,\) where Precisiont \(=\frac{{\rm {TP}}_{t}}{{\rm {TP}}_{t}+{\rm {FP}}_{t}}\), and Recallt \(=\frac{{\rm {TP}}_{t}}{{\rm {TP}}_{t}+{\rm {FN}}_{t}}\), Accuracyt \(=\frac{{\rm {TP}}_{t}+{\rm {TN}}_{t}}{{\rm {TP}}_{t}+{\rm {FP}}_{t}+{\rm {TN}}_{t}+{\rm {FN}}_{t}}\), MCCt \(=\frac{{\rm {TP}}_{t}\times {\rm {TN}}_{t}-{\rm {FP}}_{t}\times {\rm {FN}}_{t}}{\sqrt{\left({\rm {TP}}_{t}+{\rm {FP}}_{t}\right)\left({\rm {TP}}_{t}+{\rm {FN}}_{t}\right)\left({\rm {TN}}_{t}+{\rm {FP}}_{t}\right)\left({\rm {TN}}_{t}+{\rm {FN}}_{t}\right)}}\), \({\rm {PPV}}_{t}=\frac{{\rm {TP}}_{t}}{{\rm {TP}}_{t}+{\rm {FP}}_{t}}\) and \({\rm{NPV}}_{t}=\frac{{\rm {TN}}_{t}}{{\rm {TN}}_{t}+{\rm {FN}}_{t}}\). These metrics are computed by considering the predicted risk scores as a binary classifier at each timepoint, with the threshold determined by the point that maximizes the sum of sensitivity and specificity. Finally, if multiple timepoints are evaluated, the function computes the mean of each performance metric weighted by the probability of survival at each timepoint:
$$\bar{M}=\frac{{\sum }_{t=1}^{m}{M}_{t}\times \hat{S}\left(t\right)}{{\sum }_{t=1}^{m}\hat{S}\left(t\right)}$$
where \({M}_{t}\) is the performance metric at time \(t\) and \(\hat{S}\left(t\right)\) is the estimated probability of survival at time \(t\). The evaluation function returns a dictionary containing the performance metrics at each timepoint and, if applicable, the weighted mean of each metric across all timepoints. All performance metrics are reported on a 0–100% scale for simplicity.
Statistical analysis
Continuous variables are presented as median ± s.d. or IQR, and categorical variables are presented as frequencies and percentages. The independent t-test was used for normally distributed continuous variables, the Mann−Whitney U-test for non-normally distributed continuous variables and Pearson’s χ2 test for categorical variables. Missing data were handled across the entire PMBB dataset. Variables with more than 15% missingness were excluded. Remaining missing values were imputed using multiple imputation by chained equations; imputation was performed separately within the training and test sets to avoid information leakage. Survival analyses were conducted using the Kaplan−Meier method, and differences in survival probabilities among the study groups were assessed using the log-rank test. The proportional hazards assumption was assessed using covariate-specific and global tests based on Schoenfeld residuals (Grambsch–Therneau), together with graphical evaluation of log–minus–log survival plots and plots of scaled Schoenfeld residuals versus time. Where evidence of non-proportionality was observed, time-dependent effects were modeled by introducing interactions between the covariate and functions of time (for example, log(time)) or by fitting stratified Cox models. The predictive performance of the developed models was evaluated using a comprehensive set of metrics, including AUC, C-index, specificity, sensitivity, F1 score, precision, recall, accuracy, FPR, FNR, MCC, PPV and NPV. To systematically identify proteins correlated with the nine APRS proteins, we computed multiple similarity metrics between each APRS protein vector and all other proteins measured by SomaScan. In the radar plot, proteins from APRS were ordered using a greedy traversal of the core–core correlation matrix, iteratively selecting the most strongly correlated unvisited protein. Non-APRS protein angles were determined by a weighted circular mean of APRS protein angles, with weights derived from their absolute correlations to APRS proteins.
The APOL1 high-risk with eGFR ≥60 ml min−1 1.73 m−2 group was divided into an 80% training set and a 20% test set using stratified sampling. In the training set, an elastic net Cox model was applied to select protein markers. Model performance was evaluated using eight-fold cross-validation to minimize overfitting, and predictive accuracy was assessed with AUC and tAUC at various timepoints57. The final model included nine proteins plus age, sex, baseline eGFR and log2(UACR) and was fitted using an elastic net Cox model. Sample size was estimated using the Schoenfeld method for the Cox proportional hazards model and an events per variable (EPV) approach (with EPV set at 15). The EPV method yielded the required sample size of 640 participants. By contrast, based on a two-sided significance level of 0.05, an expected 30% marker-positive rate, an anticipated event rate of 25% and a target hazard ratio of 1.648—assuming median dichotomization of the model score—a total of 685 participants was determined to provide an effective statistical power of approximately 85%. Net benefits from decision curve analysis were calculated by weighting false positives and false negatives under the assumption of equal misclassification costs58. Feature stability was checked by bootstrap enumeration. NNT was calculated as the reciprocal of the absolute risk reduction, where absolute risk reduction = control event rate × relative risk reduction (RRR = 0.27, reported for inaxaplin)13. Two-tailed P values less than 0.05 were considered statistically significant. All analyses were performed using Python 3.9.13 and R 4.3.2.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.