Study overview—ethics
The VOS phase 3 trial (ECOSPOR III) was a multicenter, randomized, double-blind, placebo-controlled study conducted at 56 US and Canadian sites from July 2017 to September 2020 (ClinicalTrials.gov: NCT03183128). The study protocol and amendments are available for download from Feuerstadt et al.12. The institutional review board for each study site reviewed and approved the protocol and applicable amendments, and all patients provided written informed consent at screening (Supplementary Table 3). The trial followed the CONSORT reporting guideline.
Study overview—trial design
The VOS phase 1 trial was an open-label study conducted at three sites to evaluate the safety, dosing and efficacy of VOS for the prevention of multiply recurrent C. difficile, as described previously18. A subsequent phase 2 randomized, placebo-controlled (2:1, VOS:placebo) study evaluated a fixed dose for the prevention of multiply recurrent C. difficile, as described previously13. For the phase 3 study, eligible adults 18 years of age or older with three or more CDI episodes within 12 months, inclusive of the qualifying acute episode, were enrolled. The qualifying episode was defined as follows: (1) three or more unformed bowel movements over two consecutive days, (2) a positive C. difficile toxin test and (3) symptom resolution after 10−21 days of standard-of-care antibiotics.
Antibiotic treatment for the qualifying episode of rCDI was at the discretion of the treating investigator. Patients were stratified by age (<65 years or ≥65 years) and antibiotic received for the qualifying episode (vancomycin or fidaxomicin). Patients were randomly assigned 1:1 to VOS (approximately 3 × 107 spore colony-forming units) or matching placebo administered as four oral capsules once daily over three consecutive days12. All patients, site staff and the sponsor were blinded to treatment assignment. Patients were instructed to take a laxative 1 day prior to treatment initiation to reduce residual antibiotic in the gastrointestinal tract, to minimize inactivation of this live biotherapeutic. Patients were monitored for up to 24 weeks for CDI recurrence, defined as follows: onset of three or more unformed bowel movements per day over two consecutive days, a positive C. difficile stool toxin assay, assessment by the investigator that antibiotic treatment was warranted and persistence of diarrhea until initiation of antibiotics.
The primary efficacy outcome measure was the rate of CDI recurrence up to 8 weeks after treatment initiation in patients who received VOS compared to patients who received placebo. Preplanned exploratory microbiome endpoints were composition of the gastrointestinal microbiome from baseline through week 24 in each treatment group and composition of bile acids in stool from baseline through week 8 after treatment in each treatment group. In this expanded post hoc analysis, we investigated engraftment of VOS spore-forming species in the context of previous VOS trials13. Then, we explored the impact of the engraftment in the VOS phase 3 trial (ECOSPOR III) on broader patient microbiome composition and stool metabolites (that is, bile acids and fatty acids) through week 24.
Sex was not considered in the clinical trial study design. Sex was determined based on self-reporting or health records, and sex-based analyses were connected and reported for the ECOSPOR III primary endpoint in Berenson et al.56. A treatment effect of lower CDI recurrence in VOS versus placebo did not differ between males and females. Therefore, sex-based microbiome analyses were not executed.
VOS manufacturing and dose characterization
Donor screening and the manufacturing program were reviewed by the FDA12,13,18. Donors underwent extensive examinations, including laboratory testing before, during and after donation periods before the material was released for manufacturing. The manufacturing process uses ethanolic inactivation and other clearance steps to kill and remove non-spore microorganisms, including potential adventitious bacteria17 and viruses16. The preparation of VOS capsules relies upon filtration and centrifugation operations, which remove approximately 99% of the non-spore fecal mass, resulting in an average of less than 20 mg of residual solids per dose13. The increased quantity of residual solids reported here versus that reported in the phase 2 study is due to higher dosing in the phase 3 trial. VOS batches were characterized using an in vitro microbiological growth assay. Spore preparations are treated with a germinant mixture and then inoculated into a panel of 11 media conditions spanning diverse carbon sources, in quadruplicate and grown out anaerobically for 5 days. Growth assay material, as well as the original dose material, underwent genomic DNA extraction and shotgun metagenomics sequencing (MGX), and the resulting spore-forming species lists were combined to generate a single VOS dose definition.
Sampling
Stool samples were collected before treatment (baseline within 3 days after completion of antibiotic treatment for the qualifying CDI episode) and after treatment at study weeks 1, 2, 8 and 24. Samples were collected in tubs and shipped on frozen gel packs to a central laboratory where they were homogenized, aliquoted and then immediately frozen at −80 °C. Unamended (neat) aliquots were used for bile acid and fatty acid measurements, and 12% weight/weight stool suspensions in 95% ethanol were generated for MGX.
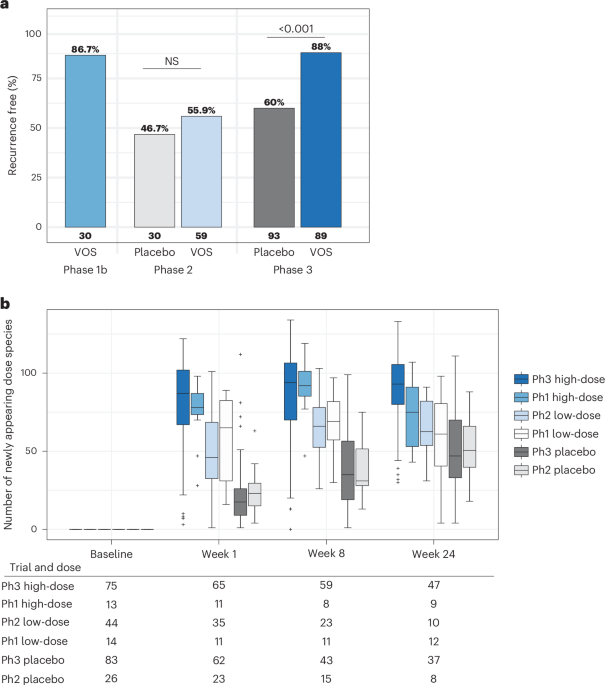
In a previously published post hoc analysis of combined VOS phase 1 and phase 2 results, dosing levels of a phase 1 dose-ranging study were divided into ‘low-dose’ and ‘high-dose’ categories, with the phase 2 ‘fixed dose’ assigned to the low-dose category13. Based on these results, the phase 3 study was designed such that patients would receive the high dose. Here, metagenomics samples from the phase 1 and phase 2 studies, collected prior to on-study CDI recurrences and at shared timepoints with the phase 3 trial, were included for cross-study engraftment comparisons. For additional details and demographic information, see Khanna et al.18 and McGovern et al.13. The institutional review board for each study site reviewed and approved the protocol, and all patients provided written informed consent.
Additionally, to generate a non-CDI comparator dataset, a single stool sample was collected from each member of a cohort of self-identified healthy individuals who reported no oral, intravenous or intramuscular antibacterial exposure within the last 12 weeks and no antifungal, antiviral or antiparasitic exposure within the last 8 weeks. The cohort had an average age of 30 years and an average body mass index of 23 kg m−2 and was 44% female. Samples were collected and processed with the same procedure as samples from the clinical study.
Microbiome and metabolomics profiling
For the phase 3 trial, DNA was extracted in-house from patient stool and drug materials using the Omega Mag-Bind Universal Pathogen Kit (Norcross). Libraries were prepared using Illumina DNA Flex kits and sequenced on the Illumina NovaSeq platform to a target sequence depth of 10 gigabases for patient samples at Clinical Laboratory Improvement Amendments (CLIA)/College of American Pathologists (CAP)-certified and Good Laboratory Practice (GLP)-compliant microbiome profiling facilities at Diversigen, Inc. Microbe taxonomic profiling for all MGX data used MetaPhlAn2 software57,58, which aligns MGX sequenced reads to species-specific markers to produce relative abundance matrices. A proprietary database of species markers was used that included taxonomic markers sourced from genomes of species in the Seres strain library as well as publicly available genomes. To monitor for contamination, each 96-well DNA extraction plate included a control sample composed of liquid media from a pure bacterial culture. After DNA extraction, a second control composed of bacterial DNA was added to a well in each sequencing plate. Both controls were sequenced alongside study samples, and no unexpected organisms were detected.
For species-level taxonomic profiling of patient stool, MGX reads in the phase 3 trial were first mapped to species-specific markers, to determine the number of mappable reads in each sample. To ensure robust comparisons of species numbers across samples and trial datasets, the number of reads mapping to species markers was downsampled to 163,000 reads (randomly selected) for each sample. This number of reads was chosen to optimize sequencing depth in individual samples while maintaining the total number of samples included in the phase 3 dataset and was applied to data from all three trials. Samples with fewer than 163,000 mappable reads were excluded from analyses (Extended Data Fig. 1).
To determine if the newly appearing dose species yield similar or different engraftment trends to strain-level engraftment, StrainPhlAn (version 4 with default parameters) was applied to combined patient and dose material MGX datasets. A cutoff of 0.02 or smaller normalized phylogenetic distance, calculated from StrainPhlan phylogenetic trees, was used to define same strains. This threshold represents the 10% quantile of the distribution of normalized phylogenetic distances between the independent healthy cohort and dose material, where strain overlap is anticipated to be minimal. Quantitative assessment of changes in bile acid and fatty acid concentrations was conducted on homogenized, lyophilized stool samples, spiked with internal standards and then subjected to liquid−liquid extraction, dilution and liquid chromatography with tandem mass spectrometry (LC−MS/MS) analysis using an Agilent 1290/SCIEX 5500 QTRAP system equipped with an Agilent SB-C18 reversed-phase column (Metabolon, Inc.). LC−MS/MS profiling of bile acids was conducted according to Good Clinical Practice, and assay validation included assessment for selectivity, specificity, limit of quantification accuracy and precision and stability. Fatty acid profiling was conducted according to Research Use Only specifications.
Untargeted stool metabolomics for extended bile acid analysis was carried out by Metabolon, Inc. according to Research Use Only specifications. For each sample, recovery standards were added prior to the first step in the extraction process for quality control purposes. To remove protein, to dissociate small molecules bound to protein or trapped in the precipitated protein matrix and to recover chemically diverse metabolites, proteins were precipitated with methanol under vigorous shaking for 2 minutes (Glen Mills, GenoGrinder 2000) followed by centrifugation. The resulting extract was divided into five fractions: two for analysis by two separate reversed-phase/ultra-performance liquid chromatography with tandem mass spectrometry (RP/UPLC−MS/MS) methods with positive ion mode electrospray ionization (ESI), one for analysis by RP/UPLC−MS/MS with negative ion mode ESI and one for analysis by hydrophilic interaction (HILIC)/UPLC−MS/MS with negative ion mode ESI.
We directly measured the inhibitory effect of short-chain and medium-chain fatty acids on C. difficile in an in vitro batch culture assay measuring C. difficile growth across a range of concentrations of fatty acids and three pH values both typical of a healthy colonic environment. Short-chain and medium-chain fatty acids are found in concentrations of up to 120 mM in human colonic contents, with corresponding pH ranging from 5.6 to 7.0 depending on the colonic region59,60. C. difficile was cultured in the presence of mannitol or methionine/glycine, which are two different growth substrates that C. difficile uses and metabolizes by different pathways in vivo50,51,52. All experiments were carried out with three different C. difficile ribotypes (RT-001, Clostridioides difficile American Type Culture Collection (ATCC) 9689; RT-060, Clostridioides difficile ATCC 43593; RT-087, Clostridioides difficile ATCC 43255) to assess strain or ribotype-specific impacts. Half-maximal inhibitory concentration (IC50) values were calculated from curves based on growth, quantified by measuring optical density at 600 nm (OD600) across eight concentrations of fatty acids. Each estimate of IC50 is based on the average optical density value from three replicate cultures.
VOS dose material from five batches (generated from four donors, with one donor contributing material to two separate batches) was assayed in vitro for production of DCA, LCA, butyrate, hexanoate and valerate by growing each in a single complex liquid medium across a 10-fold dilution series with eight dilutions in triplicate. Cultures were grown for 5 days, and then supernatant was collected for metabolite concentration quantification by LC−MS/MS as described above (Metabolon, Inc.). Metabolite concentrations at time 0, obtained from supernatants collected at the beginning of the experiment right after inoculation, were subtracted from final concentrations to account for the composition of the media. Dilutions with the highest concentration for each metabolite are reported.
Statistical analyses
The phase 3 preplanned microbiome-related analyses included evaluation of VOS dose species engraftment, changes in species composition and bile acid concentrations in VOS as compared to placebo groups from baseline to weeks 1, 2, 8 and 24 (ref. 12). Preplanned analyses of microbiome compositional and targeted metabolomic for primary and secondary bile acids were performed on the safety population, and the distribution of patient engraftment and secondary bile acid measurements within treatment group were assessed with non-parametric median and interquartile range statistics. The evaluation of engraftment, changes in species composition and bile acid concentrations presented herein are post hoc analyses consistent with the preplanned statistical analysis with the following deviations. (1) The population for the preplanned microbiome analyses comprised all patients who were randomized, received any amount of study drug and who provided at least one evaluable stool sample before treatment and at least one evaluable stool sample after treatment. In these post hoc analyses, samples collected after treatment of an on-study CDI recurrence were excluded, as treatment of C. difficile recurrence requires antibiotics, which impact microbiome signals (Extended Data Fig. 1). These post-recurrence, post-antibiotic samples were more common in patients receiving placebo, where the recurrence rate was significantly higher. Removal of these samples enables examination of the impact of treatment without the confounding impact of antibiotics. (2) For analyses not requiring both pretreatment and posttreatment samples to derive values, both single and paired pretreatment/posttreatment stool samples were used. (3) Bile acid concentrations were compared directly as absolute concentrations instead of evaluated as change from baseline to enable clear communication of results. Comparisons to VOS phase 1, VOS phase 2 and the healthy comparator cohort, as well as fatty acid concentration measurements, were not preplanned.
Figures and statistical analyses were carried out using R version 3.6.3. P values reported for comparisons between VOS and placebo arms at single timepoints were generated using two-sided MWU in R using the rstatix package version 0.7.2. This included comparisons of engraftment, engraftment within prespecified subgroups (that is, age <65 years versus ≥65 years) and prior antibiotic regimen (vancomycin versus fidaxomicin), phylum-level relative abundances and concentrations of bile acids and fatty acids. Box plots used to visualize metrics show the median (central horizontal line) and interquartile range (shaded box) in each group at each timepoint; vertical bars indicate the most extreme non-outlier values (within 1.5 times the interquartile range); and crosses (+) indicate outlier values (outside 1.5 times the interquartile range).
NMDS plots were used to visualize the community composition of patient microbiome samples in the context of healthy individuals. Differences in overall community composition between treatment arms at each timepoint were evaluated with PERMANOVA tests, and sample dispersion was assessed using the PERDISP2 procedure. MetaMDS, adonis and betadispers functions in vegan version 2.5-6, respectively, were used. Differences in genus prevalence across arms were assessed using Fisherʼs exact tests adjusted for multiple hypothesis testing by controlling the false discovery rate (FDR) with the Benjamini−Hochberg procedure. Plots were generated using the following packages: dplyr version 1.1.2, ggplot2 version 3.7, ggpubr version 0.6.0 and reshape2 version 1.1.4.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.