Study populations and sample collection
Cross-sectional cohorts
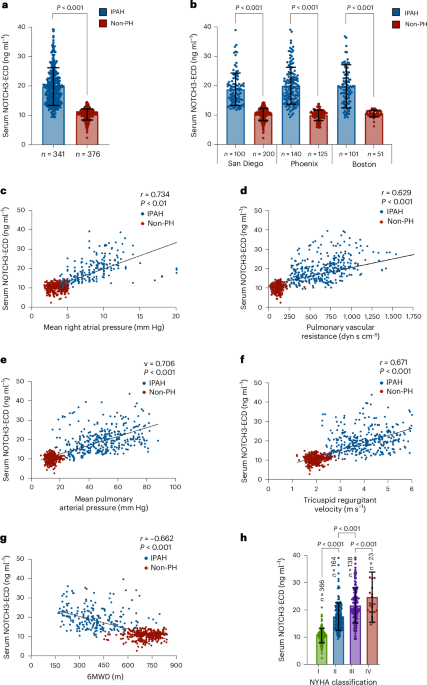
Between 2010 and 2021, serum samples were prospectively collected from 341 individuals with IPAH and 376 without PH (Table 1) being treated at the University of California, San Diego (UCSD) (San Diego cohort: 100 IPAH, 200 non-PH), the University of Arizona (Phoenix cohort: 140 IPAH, 125 non-PH) and the Massachusetts General Hospital (Boston cohort: 101 IPAH, 51 non-PH) (Table 1). A sample size of 341 individuals with IPAH and 376 individuals without PH was calculated to provide 90% power to detect a minimum effect size of 0.27 for the difference in serum NOTCH3-ECD levels between the two groups with a two-sided α = 0.05. The San Diego and Phoenix IPAH patient samples were collected on an outpatient basis, whereas the Boston patients with IPAH had serum collected during ICU hospitalization for IPAH. Informed consent was obtained for all individuals from each cohort. Patient sex was determined by assigned sex according to each patient’s hospital records. All IPAH blood samples were collected within 1 month of RHC or ECHO. Control individuals in the San Diego and Phoenix cohorts were healthy paid volunteers, whereas the Boston control cohort comprised ICU patients without PH, being treated for other non-lung-related diseases (51 nonintubated patients comprising 37 multi-trauma orthopedic patients without lung contusion, acute respiratory distress syndrome or pulmonary embolism; 10 individuals recovering from elective large abdominal, vascular or orthopedic operations; 3 patients with a closed head injury; and 1 patient with amyotrophic lateral sclerosis). All individuals, including controls, underwent RHC as part of this study. All IPAH patients tested negative for HIV and active hepatitis viral infection, as well as anti-nuclear, anti-centromere, anti-mitochondrial, anti-double-stranded DNA, anti-topoisomerase 1, anti-Ro and anti-La antibodies. No patient with IPAH was undergoing evaluation for liver transplantation or had received a previous liver transplant. Patients with IPAH did not undergo genetic testing at the time of diagnosis and none had a family history of PAH (WHO group 1) or other PH (WHO groups 2–5). The determination of IPAH was made by an integrated assessment by an experienced PH pulmonologist or cardiologist, with further adjudication as needed by a committee of PH physicians. Inclusion and exclusion criteria for the diagnosis of IPAH is given in Supplementary Table 1.
Patients with subtypes of WHO group 1 PAH associated with previously diagnosed heritable mutations, methamphetamine, scleroderma, HIV, congenital heart defects, portal hypertension or pulmonary veno-occlusive disease were not included in the primary analysis, although serum samples from these patients were collected from UCSD and PH centers in the United States and United Kingdom for secondary comparative analysis. Serum samples were also obtained from individuals with non-PH vasculitides that affect the lung as well as individuals with malignancies expressing NOTCH3 for additional secondary comparative analysis. Informed consent was obtained for all individuals at participating institutions for the above patient groups. Sample numbers for secondary comparative analyses were determined by availability of serum samples over the 9 years of collection from multiple participating institutions, UCSD and the UCSD Biorepository. Only IPAH and non-PH serum samples were included in the primary analysis.
Longitudinal cohort
A separate cohort of 100 newly diagnosed, treatment-naive patients with IPAH (43 patients from San Diego, 57 patients from Phoenix), were followed for 6 years with serial blood sampling, ECHO and RHC (Extended Data Table 2). Informed consent was obtained from all 100 individuals. Sex was determined by assigned sex according to each patient’s hospital records. No living patients were lost to follow-up during the time course of the study. Blood samples were taken on the day of RHC at diagnosis, at 3 years and at 6 years. Sample numbers for longitudinal analysis were determined by availability of serum samples collected at three timepoints for each patient over a 6-year period. These patients were analyzed separately from cross-sectional cohorts.
Sample collection
Blood samples were collected by venipuncture in heparin-coated tubes (Becton Dickinson) within 1 month of the most recent RHC or ECHO. Serum was separated by Ficoll (Amersham) gradient centrifugation of whole blood and was stored at −80 °C. Serum samples were de-identified using a barcode at the point of care before being transported to the lab for analysis. For ten patients, serum was derived from blood sampled from the pulmonary artery and transseptally from the left atrium during cardiac catheterization to investigate the origin of NOTCH3-ECD as part of institutional review board (IRB)-approved research protocols. All studies were approved by the UCSD Human Subjects Program and relevant IRB committees of all participating institutions, with the following protocols: UCSD: IRB protocol no. 809539; Mass General Brigham: IRB protocol no. 2010P000982; University of Arizona: IRB protocol no. 1100000621A013, University of Alabama: IRB protocol no. 1639383-7; University of New Mexico: IRB protocol no. 00003255; University of California, Los Angeles (UCLA): IRB protocol no. 12-0738; Americas Hospital, Guadalajara, Mexico: IRB protocol no. AH-IRB-B2; Mount Sinai, New York: IRB protocol no. RC-4590; and University of Cambridge (UK): IRB and informed consent as part of the UK National Cohort Study of Idiopathic and Heritable Pulmonary Arterial Hypertension (ClinicalTrials.gov ID: NCT01907295; UK Research Ethics Committee reference no. 13/EE/0203).
ELISA
Levels of NOTCH3-ECD in human serum were quantified in nanograms per milliliter (ng ml−1) using the human NOTCH3-ECD ELISA kit (Cloud Clone, cat. no. SEL147Hu) according to the manufacturer’s protocol. Operators who performed the ELISAs were blinded to case and control status of the samples. NOTCH3-ECD standards were reconstituted with 1.0 ml of standard diluent to achieve a high standard of 100 ng ml−1. Serial dilution was performed to establish a concentration curve (ng ml−1) of 100, 50, 25, 12.5, 6.25, 3.13 1.57 and 0.78 as well as a final blank of 0 ng ml−1 Serum samples were diluted 1:5 by mixing 20 μl of serum with 80 μl of phosphate-buffered saline. Each sample of the dilution series, as well as each diluted serum sample (all 100 μl), was placed in triplicate into individual wells of a 96-well anti-NOTCH3-ECD-coated plate and incubated for 1 h at 37 °C. The liquid was removed by decanting each plate. Diluted biotinylated antibody (100 μl) was added to each well. Plates were incubated for 1 h at 37 °C. Wells were covered with 350 μl of the manufacturer’s wash solution and decanted after 1 min. Three wash cycles using 350 μl of the manufacturer’s wash solution were performed. Streptavidin-conjugated horseradish peroxidase, 100 µl, was added to each well. Plates were incubated for 30 min at 37 °C. Five wash cycles with the manufacturer’s wash solution were performed as above. Tetramethylbenzidine substrate solution, 90 µl, was added to each well. Plates were covered with aluminum foil and incubated for 20 min at 37 °C. The reaction was terminated by adding 50 μl of stop solution to each well. Absorbance was measured immediately at 450 nm using a SpectraMax M2e plate reader (Molecular Devices) and the results were collated using SoftMax Pro v5.4 (Molecular Devices). Serum samples were run on different locations on the plate each time that they were assayed, with plates randomly containing IPAH and control samples from each geographical cohort. All experiments were performed in triplicate on 3 days separately using different ELISA lots.
Cross-reactivity testing
ELISA specificity for the anti-NOTCH3-ECD antibody was tested by adding recombinant human NOTCH1-ECD (Beta Life Sciences, cat. no. BLPSN-3544), NOTCH2-ECD (Beta Life Sciences, cat. no. BLPSN-3547) and NOTCH4-ECD (Beta Life Sciences, cat. no. BLPSN-3549) peptides diluted in phosphate-buffered saline to concentrations of 0.78–100 ng ml−1. Cross-reactivity was determined quantitatively by optical density compared to blank controls. Experiments were performed in triplicate on 3 days separately, using different assay lots.
Immunoprecipitation and western blotting
Immunoprecipitation and western blotting were performed as previously described59. The antibodies used for western blotting were: human anti-NOTCH3-ECD antibody (Sigma-Aldrich, clone 2G8, cat. no. MABF937, 1:1,000), goat polyclonal anti-rat secondary antibody (Thermo Fisher Scientific, cat. no. 31470, 1:5,000), rabbit polyclonal anti-transferrin antibody (Thermo Fisher Scientific, cat. no. PA527306, 1:1,000) and horseradish peroxidase-conjugated goat polyclonal anti-rabbit immunoglobulin G antibody (Thermo Fisher Scientific, cat. no. 31460, 1:5,000). For immunoprecipitation experiments, primary antibodies were used at a concentration of 6 μg of antibody per 1 mg of total protein. Experiments were performed in triplicate on 3 days separately.
Statistical analysis
Diagnostic analyses
The primary objective was to assess the ability of serum NOTCH3-ECD to differentiate between individuals with IPAH and individuals without PH. Logistic regression was used to generate ROCs to assess the ability of serum NOTCH3-ECD levels to predict the presence of IPAH in three geographically separate cohorts (San Diego, Phoenix and Boston) and one combined cohort.
Specifically, the optimal cutoff of serum NOTCH3-ECD to maximize diagnostic sensitivity and specificity to differentiate between individuals with IPAH and individuals without PH was determined by calculating the optimal Youden’s Index and F1 score. The cutoff was established first in the San Diego cohort independently and then externally applied to the Phoenix and Boston cohorts. Finally, the cutoff value was applied to the combined cohort to evaluate potential generalizability and overall performance. The AUC, F1 score, precision and recall at the defined cutoff were calculated for each cohort independently and then as a combined group.
Prognostic analyses
The secondary objective was to assess whether levels of serum NOTCH3-ECD predict mortality in 341 individuals with IPAH (combined cohort from San Diego, Phoenix and Boston) within 3 years of sample collection. Kaplan–Meier plots were constructed with patients censored at the date last known to be alive or the date of lung or heart–lung transplantation. A log-rank test was used to compare individuals with serum NOTCH3-ECD levels above and below the predetermined cutoff values and HRs were estimated using Cox’s proportional hazards models. Separate analysis was also performed for which lung or heart–lung transplantation and mortality were both considered events and the transplant-free survival was calculated. Individual patient mortality was verified from the medical records.
Multivariate time-dependent Cox’s regression was also employed to assess the impact of serum NOTCH3-ECD levels on the 3-year mortality of 341 individuals with IPAH (combined cohort from San Diego, Phoenix and Boston) while adjusting for major prognostic factors for IPAH. The variables included in the final multivariate Cox’s regression model were those with P < 0.20 after backward, stepwise, logistic regression predicting 3-year mortality (Supplementary Table 2). The final Cox’s regression model included patient age, sex, NYHA class, 6MWD, PVR, NT-proBNP and serum NOTCH3-ECD.
To compare the ability of the REVEAL 2.0, REVEAL 2.0 Lite and COMPERA 2.0 scores to predict 3-year mortality in IPAH individuals (combined cohort from San Diego, Phoenix and Boston), both with and without the addition of serum NOTCH3-ECD, separate machine learning models were generated (machine learning code in Supplementary Note 1). Specifically, the extreme gradient boost (XGBoost) algorithm was employed due to its effectiveness in handling high-dimension data, robustness and ability to capture complex patterns. In addition, XGBoost is particularly skilled at handling missing data effectively, which typically limits the predictive ability of nomograms and other machine learning tools such as neural networks. As such, no imputation of data was performed or utilized.
The binary classification problem processed using machine learning (survival versus death within 3 years) was implemented using the R caret package. The clinical characteristics of the training set were used as independent variables to develop machine learning models to predict all-cause mortality within 3 years. All variables included in the REVEAL 2.035, REVEAL 2.0 Lite35 and COMPERA 2.036 calculators were included in each individual machine learning model, with and without NOTCH3-ECD levels. There were a maximum of 13 categorical variables (demographics: men, age >60 years, eGFR < 60 ml min−1 1.73 m−2 or renal insufficiency, NYHA class, systolic blood pressure ≥110 mm Hg or <110 mm Hg, heart rate ≤96 beats per min or >96 beats per min, all-cause hospitalizations ≤6 months, 6MWD categories in respect of each calculator, NT-proBNP or BNP values respective to each calculator, pericardial effusion on echocardiogram, percentage predicted DLCO ≤ 40, mRAP > 20 mm Hg within 1 year and PVR < 5 Wood units on RHC) and one continuous variable (serum NOTCH3-ECD) included for each model with respect to the corresponding mortality calculator. Data were formatted into a binary framework for each of the categorical variables by converting them into dummy variables. To avoid perfect multicollinearity, the first of the dummy variables was dropped from the model. The final dataset was divided into training (80%) and testing (20%) data subsets.
The models were constructed using the randomForest package in R and hyperparameters, including the number of trees, mtry (number of predictors to sample at each split) and min_n (number of observations needed to split nodes). Then, the models were optimized as described previously60,61. Briefly, tuning parameters, which are modifiable variables such as the rate of learning, depth and complexity of the model, were tested for each classifier to obtain the best prediction in the training dataset. Bayesian optimization was employed for hyperparameter tuning to iteratively search for the ideal parameters. For XGBoost, the hyperparameters optimized were eta, max_depth, min_child_weight, subsample and nfold. Optimization was performed on the training sub-dataset with the ‘scoring_function’ that evaluated the model based on the resulting AUC.
The risk of mortality was predicted using the test dataset and the predictive performance was evaluated by examining the AUC. The AUC, F1 score, precision, recall, accuracy and balanced accuracy after k-fold crossvalidation were calculated and reported as the mean ± s.d. for all models after tenfold validation. K-fold crossvalidation was employed to maximize the utility of available data while providing a robust assessment of model performance. By partitioning the dataset into k subsets and iteratively using each subset for validation while training on the remainder, this technique reduces evaluation bias and delivers more reliable performance metrics than traditional single train-test splits.
Longitudinal cohort analyses
In a separate, longitudinal cohort of 100 treatment-naive IPAH individuals, the survival, change in serum NOTCH3-ECD levels, mRAP, PVR, mPAP, TRV, 6MWD and NHYA class over 6 years was analyzed. Kaplan–Meier curves were constructed for overall survival as well as transplant-free survival within 6 years of diagnosis. Patients were censored at the date last known to be alive or the date of a lung or heart–lung transplantation.
Trends of serum NOTCH3-ECD levels, mRAP, PVR, mPAP, 6MWD and TRV were compared between patients who developed progressive disease and underwent a transplantation or died within 6 years of follow-up and patients who survived until the end of the study period. To visualize the longitudinal trends of key clinical variables (NOTCH3-ECD, mRAP, PVR, mPAP, TRV and 6MWD) over the study period (years 0, 3 and 6), locally estimated scatterplot smoothing plots were generated using the ggplot2 package in R. Locally estimated scatterplot smoothing is a nonparametric method that fits local polynomial regressions to the data, allowing visualization of the central tendency without assuming a specific global functional form. In these plots, time (in years) was plotted on the x axis against the clinical variable’s value on the y axis. Each variable was displayed in a separate facet with independent y-axis scaling to accommodate differing value ranges. A mixed ANOVA was employed to analyze the role of time and prognostic status (death or transplantation versus survival) on the trends of serum NOTCH3-ECD levels, mRAP, PVR, mPAP, TRV and 6MWD.
Data are presented as mean ± s.d., unless otherwise indicated. Comparison between independent groups for continuous variables was performed using independent two-sample Student’s t-test and ANOVA with a post-hoc Tukey test, as indicated. Correlation between continuous variables was assessed by Spearman’s rank correlation coefficient. A two-sided P < 0.05 was considered to indicate statistical significance. Data were analyzed using GraphPad Prism, v9.1.2 (GraphPad Software) and R software, v4.21 (R Foundation for Statistical Computing).
Missing data
There were 3.5–5% missing data in the cross-sectional cohort and 2–3% in the longitudinal cohort. The amount of missing data was distributed without focality with respect to individual data entries or time periods. In cases where data were missing, these entries were left blank and no forms of imputation were performed.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.