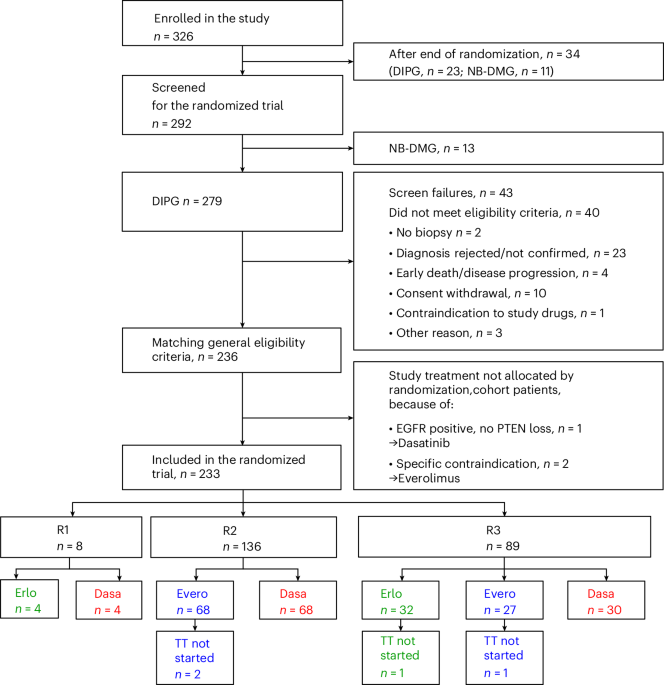
Population of the study and control cohort
A total of 326 patients were enrolled between 2 October 2014 and 6 May 2020 (Fig. 1). Thirty-four patients were treated with everolimus after the trial steering committee’s recommendation to stop randomization.
Thirty-four patients were enrolled after the end of randomization (Amendment 7) and were, therefore, not included in the analysis. Thirteen patients with NB-DMG were randomized in the trial after Amendment 7, which authorized the enrollment of DMG located outside the pons, and these patients were not considered for the main analysis. Dasa, dasatinib; Erlo, erlotinib; Evero, everolimus; TT, treatment.
Among the 292 patients enrolled before randomization closed, 43 were screen failures. Thirteen with non-brainstem DMG (NB-DMG) H3K27-altered became eligible in the trial after the change of the WHO classification. Of these 43, 33 did not meet eligibility criteria (including 23 with a diagnosis centrally rejected), and 10 withdrew consent. Three more patients were not randomized for other reasons. The most frequent differential diagnoses were ganglioglioma, pilocytic astrocytoma, pediatric high-grade glioma of the subclass MYCN (pedHGG-MYCN), MYB−QKI angiocentric glioma and NF1-related glioma (Supplementary Table 1).
In total, 233 patients were randomized in the three subtrials (Table 1) until 20 September 2019 when the independent data monitoring committee recommended stopping randomization for futility (Fig. 1). Ninety-five patients were randomized to receive everolimus, 102 to receive dasatinib and 36 to receive erlotinib. Sex was balanced. Median age was 8.1 years (range, 1.8−30.3). Seventy-two percent were treated in France. Other brainstem structures than pons were involved in 43%, and 3% had metastases at diagnosis.
The median interval between radiological diagnosis and treatment initiation was 26.5 days (interquartile range (IQR), 21–32) (Supplementary Table 2).
Sixty-six patients with biopsy-proven DIPG treated in Gustave Roussy before the start of BIOMEDE23,24 were used as historical controls, excluding those who had received any BIOMEDE drugs. Most received radiotherapy and temozolomide.
Trial design and endpoints
Patients with radiologically suspected DIPG first consented to biopsy. After diagnosis confirmation (DMG H3K27-altered with an epicenter in the pons) by central pathology reviewer, enrollment in the treatment randomization based on biomarkers was offered in case of age over 6 months, absence of active intratumoral hemorrhage, ability to undergo radiotherapy, Lansky Play Scale >50% and life expectancy longer than 3 months. Metastatic disease was allowed.
The drugs tested were administered during and after normofractionated radiotherapy at 54 Gy. Three drugs were compared: everolimus 5 mg m−2 per dose once daily, erlotinib 125 mg m−2 per dose once daily or dasatinib 85 mg m−2 per dose twice daily. The trial was designed to randomize these three drugs in patients with biopsy-proven DIPG according to specific biomarkers (Extended Data Fig. 1). According to the biomarker constellation present, three randomizations were possible: R1 (erlotinib versus dasatinib) in patients where tumor was EGFR positive without PTEN loss; R2 (everolimus versus dasatinib) in patients where tumor showed only PTEN loss without EGFR overexpression; and R3 (erlotinib versus everolimus versus dasatinib) in patients with tumor showing both EGFR overexpression and PTEN loss (Extended Data Fig. 1). In total, eight patients were randomized in R1, 136 in R2 and 89 in R3 (Fig. 1 and Extended Data Table 1). Patients with biomarker-negative tumors received dasatinib without randomization and were excluded from the final analysis. Patients without information on biomarkers (biopsy not diagnostic but centrally reviewed radiologic features of DIPG, necrotic tumor not amenable to immunohistochemical analysis or lack of material for biomarkers) were assigned in R3. This design enabled pairwise drug efficacy comparisons by distributing R3 patients according to a given pairwise comparison; these OS comparisons were the primary objective. Considering the smallest pairwise comparison (erlotinib versus everolimus), 79 events and 90 patients were required to achieve 80% power to compare OS curves using a two-sided log-rank test (α = 20%), if the hazard ratio of death was 0.62, equivalent to a 10.6% increase in the 2-year OS (5% versus 15.6%) assuming an exponential survival distribution, proportional hazards and an expected accrual of 22.5 patients per year (East software; Cytel). Considering the expected biomarker distribution, we planned to recruit a total of 250 patients in the entire randomized trial to ensure 90 patients in the smallest pairwise comparison (erlotinib versus everolimus). Comparison with historical controls was also part of this primary objective.
Secondary endpoints were 2-year OS, progression-free survival (PFS) and safety profile. The biomarkers associated with disease outcome, the association between biomarkers and drug efficacy and the radiographic changes observed after radiotherapy were the planned exploratory objectives.
Survival analysis
The trial did not show any significant OS difference (Fig. 2a–d and Extended Data Table 2). With a median follow-up of 5.3 years, median OS since the biopsy was 11.1 months (95% CI: 9.7−11.7) in the trial compared to 10.8 months in the control cohort (95% CI: 9.5−13.0). No difference was observed for any of the treatment arms compared to the historical control, with median OS of 9.7 months (95% CI: 7.8−14.6), 9.9 months (95% CI: 8.8−11.2) and 11.9 months (95% CI: 10.7−14.2) for patients treated with erlotinib, dasatinib and everolimus, respectively (Fig. 2e and Extended Data Table 2).
The main outcome measures of the trial were based on pairwise comparisons of OS between treatment groups allocated by randomization. For these comparisons, all randomized patients were included, including those who did not start the treatment allocated by randomization (intention-to-treat analysis). OS was estimated using the Kaplan−Meier method, from the date of randomization to the date of death from any cause. a, OS curves: erlotinib versus dasatinib. This comparison includes 70 patients from the R1 subtrial and a subset of patients from the R3 subtrial. Median OS was 9.0 months (95% CI: 7.4−14.4) for erlotinib and 8.5 months (95% CI: 5.7−10.7) for dasatinib. The hazard ratio (HR) stratified by PTEN status was HR(erlotinib/dasatinib) = 0.87 (95% CI: 0.52−1.46), P = 0.59. b, OS curves: everolimus versus erlotinib. This comparison includes 59 patients from the R3 subtrial. Median OS was 10.2 months (95% CI: 7.3−14.8) for erlotinib and 10.5 months (95% CI: 7.6−12.3) for everolimus. The HR stratified by EGFR status and PTEN status was HR(erlotinib/everolimus) = 0.94 (95% CI: 0.54−1.65), P = 0.84. c, OS curves: everolimus versus dasatinib. This comparison includes 193 patients from the R2 subtrial and a subset of patients from the R3 subtrial. Median OS was 11.3 months (95% CI: 10.3−13.4) for everolimus and 9.4 months (95% CI: 8.2−10.8) for dasatinib. The HR stratified by EGFR status was HR(everolimus/dasatinib) = 0.89 (95% CI: 0.66−1.19), P = 0.42. d, PFS in patients treated with everolimus or dasatinib. This comparison includes 193 patients from the R2 subtrial and a subset of patients from the R3 subtrial. Median PFS was 6.4 and 7.1 months, respectively. No significant difference was observed. e, OS since the biopsy in the three treatment arms compared to the historical control. With a median follow-up of 5.3 years, the median OS from the time of biopsy was 11.1 months (95% CI: 9.7−11.7) in the trial cohort compared to 10.8 months (95% CI: 9.5−13.0) in the historical control cohort. No significant difference was observed between any of the treatment arms and the historical control, with median OS of 9.7 months (95% CI: 7.8−14.6), 9.9 months (95% CI: 8.8−11.2) and 11.9 months (95% CI: 10.7−14.2) for patients treated with erlotinib, dasatinib and everolimus, respectively.
Source data
Because there was no recommendation for treatment at progression, some patients switched to the drug used in other arms, and some patients received reirradiation at the time of progression, especially if PFS was superior or equal to 6 months. We considered reirradiation as a possible confounding factor for OS analysis. The reirradiation rate was similar in the three treatment arms: 25% of patients underwent reirradiation—30% in the everolimus arm, 22% in the dasatinib arm and 22% in the erlotinib arm (Fisherʼs exact test, P = 0.32).
To allow comparison of survival with studies not mandating the biopsy as a diagnostic procedure, median OS after radiological diagnosis was measured: 10.3 months (95% CI: 8–14.8), 10.5 months (95% CI: 9.5–11.5) and 12.1 months (95% CI: 10.9–14.6) for patients randomized to erlotinib, dasatinib and everolimus, respectively.
PFS was not different in the three treatment arms (log-rank test, P = 0.89; Extended Data Fig. 2a).
The 2-year OS was similar in the three arms: 144 patients (62%) survived less than 1 year; 70 (30%) survived between 1 year and 2 years; and 19 (8%) survived more than 2 years (Extended Data Table 2).
Safety
Safety and outcome of the biopsy procedure
Neurosurgeons performing the biopsies were either authorized based on their previous track records for this procedure or trained in two dedicated sessions (S.P. and K.A.). No biopsy-related deaths were reported. Median hospital stay for the biopsy was 2 days (range, 1–5). In the 157 biopsy procedures that were randomly centrally reviewed, 38 patients experienced neurological worsening, of which 27 occurred within 48 hours of the biopsy and were, therefore, considered related to the procedure. Fourteen severe (grade 3−4) complications were reported, regardless of center biopsy volume.
Whatever the depth of the tumor biopsy judged centrally on postoperative computed tomography or magnetic resonance imaging (MRI), the median variant allele frequency (VAF) of the H3K27M mutation was approximately 40% (Extended Data Fig. 3a–d). Tumor mutation burden was not associated with the VAF of the histone H3K27M mutation (Extended Data Fig. 3e). Nucleic acid extraction yielded a median of 9.8 µg of DNA and 2.9 µg of RNA of sufficient quality (that is, RNA integrity number (RIN) > 5), enabling subsequent genomic analyses in 176 cases.
Treatment-related adverse events
Treatments were well tolerated. No patient died from study treatment-related complications. Seventy-eight percent of patients experienced grade 3 or grade 4 adverse events during treatment. When considering adverse events of any grade, eye (P < 0.0001), skin (P = 0.004) and infectious (P = 0.042) adverse events were more frequent with erlotinib, whereas metabolic adverse events were more frequent with everolimus (P = 0.0003) (Extended Data Fig. 3f,g, Extended Data Table 3 and Supplementary Table 3). Severe (grade 3 or higher) skin adverse events were significantly more frequent with erlotinib (P < 0.0001), and severe renal (P = 0.0054) and gastrointestinal (P = 0.038) adverse events were significantly more frequent with dasatinib. Irrespective of the imputability to the treatment or the disease, neurological severe adverse events reported did not differ in their frequency among treatment arms (Fisherʼs exact test, P = 0.65) (Supplementary Table 3); the three most frequent were headaches, hydrocephalus and dizziness. Treatment was stopped because of toxicity in 20%, 3% and 14% for erlotinib, everolimus and dasatinib, respectively (Fisherʼs exact test, P = 0.004). The tolerance of everolimus was better than reported in breast cancer trials25 and similar to a recent report in children with low-grade gliomas26.
Exploratory outcomes
Response to therapy
Clinical improvement any time during first-line treatment was reported in 75% of the patients, whereas the clinical status was stable in 19% and deteriorated in 6%. Clinical response did not differ among treatment arms (Supplementary Table 4). Mean duration of improvement was 3.4 months, with no difference between arms. The OS of the patients with clinical improvement (11.2 months (95% CI: 10.3−12.3)) did not differ from the one of the 44 patients remaining clinically stable (9.2 months (95% CI: 6.4−10.9)) (log-rank test).
Radiologic improvement as reported by the treating physician during first-line treatment was observed in 121 patients (54%), whereas the disease remained stable as best response in 70 patients (31%) or progressed in 32 patients (14%) with no difference among treatment arms (χ2 test, P = 0.402).
Pseudoprogression was reported in 110 of 233 patients (49%) with no significant difference among arms (χ2 test, P = 0.870).
Planned prognostic factor analysis
TP53 mutation was the main prognostic factor for OS and remained significant in multivariable analysis (hazard ratio = 2.84 (95% CI: 1.92−4.20), P < 0.0001) when adjusting for immunohistochemical biomarker status, treatment allocation, type of histone H3 alteration and age (Supplementary Tables 5 and 6 and Extended Data Fig. 4). We observed a borderline significant association between histone mutation and OS in univariate analysis (P = 0.064); compared to H3 wild-type (H3-WT), the outcome was worse in patients with H3.3K27M (hazard ratio = 1.28 (95% CI: 0.81−2.03)) and better in patients with H3.1K27M (hazard ratio = 0.81 (95% CI: 0.46−1.42)). TP53 mutations were significantly more frequent in H3.3K27M than in H3.1K27M or H3-WT (67.9%, 24.1% and 28.6%, respectively; P < 0.0001; Supplementary Table 6), and the prognostic value of histone mutation disappeared when adjusting on covariables, in particular TP53 mutation (P = 0.49 in multivariable analysis) (Extended Data Fig. 2c). Of note, as previously published27,28,29,30,31, age was significantly associated in univariate analysis with OS (P = 0.009), with, compared to patients 10 years or older, a longer OS for the youngest (hazard ratio = 0.79 (95% CI: 0.53−1.16)) and a shorter OS for patients between 5 years and 10 years of age (hazard ratio = 1.34 (95% CI: 0.99−1.80)) (Extended Data Fig. 2d). However, H3.3K27M and TP53 mutations were much less frequent in younger patients (Supplementary Table 7), and, when controlling for biological features of the tumor, the effect of young age appeared significantly reversed (hazard ratio = 1.72 (95% CI: 1.04−2.86)).
A planned exploratory analysis assessed clinical and known prognostic biomarkers associated with response to everolimus versus dasatinib, the only comparison with sufficient patient numbers; treatment effect appears relatively homogeneous across main subgroups (Extended Data Fig. 4).
Overall portrait of recurrent single-nucleotide variants
Mutation rates varied among patients, independently of H3 mutational status or OS (Fig. 3a). The number and type of non-synonymous coding variants were consistent with previous reports, with a median of 17.5 mutations per exome29,30. Missense variants accounted for most single-nucleotide variants (SNVs) (Fig. 3a).
a, Oncoplot depicting the most frequent somatically mutated genes in DIPG (n = 172), sorted in decreasing order of mutation frequency and stratified by OS—that is, less than 1 year (short), between 1 year and 2 years (intermediate) and more than 2 years (long). Twenty-two genes mutated in at least 3% of patients are shown, with mutation frequencies displayed as a bar plot on the right. Only somatic SNVs supported by more than five reads associated and with a VAF higher than 5% are shown. A stacked bar plot representing the number and types of variants per sample is displayed on the right. Germline variants supported by more than five reads and with a VAF exceeding 40% are indicated by small squares. Clinicopathological (EGFR overexpression by immunohistochemistry) and molecular annotations are displayed as bars at the top, according to the color code provided below the Oncoplot. Indels were overall rare although enriched in some genes compared to other tumor types, such as FBXO41 and PIK3R1. Some genes showed frequent stop-gain mutations (ASXL1and PPM1D) or frameshift variants (KAZN). At the bottom, a bar plot shows, for each patient, the proportion of the 15 most prevalent COSMIC SNV signatures in the cohort after removal of ‘artifactual’ signatures. Patients in the ‘short survivor’ group exhibited mutational signature profiles associated with SBS14 (concurrent polymerase epsilon mutation and defective DNA mismatch repair), SBS18 (reactive oxygen species), SBS30 (defective base excision repair) and SBS44 (defective DNA mismatch repair) and with signatures of unknown etiology (SBS33, SBS34 and SBS39). Only SBS19 (unknown etiology) was enriched in the ‘long survivor’ group. b, Distribution of VAFs for the more frequently mutated genes. The number of patients harboring an alteration is indicated on the left; each dot represents one SNV. Whiskers indicate the minimum and maximum values within 1.5× IQR. Whiskers represent the minimum and maximum values within 1.5× IQR. Q1, Q2 and Q3 correspond to the 25th, 50th (median) and 75th percentiles, respectively. c, Distribution of H3-K27M, TP53 and PIK3CA alterations, MYC gain and EGFR overexpression across survival subgroups is shown. Two-sided Fisher’s exact test, ****P < 0.0001, ***P < 0.001, **P < 0.01, *P < 0.1; that is, P = 0.01 for H3-K27M, P = 1.6 × 10−7 for TP53, P = 0.08 for PIK3CA and P = 0.001 for MYC gain. d,e, Box plots showing the distribution of the percentage of altered genome number of segments and detected by CNV analysis of WES data according to TP53 mutational status. (TP53WT, n = 73, TP53MUT, n = 99). The value (number of segments +1) is plotted on a log10 scale. Wilcoxon test, *P < 0.05, **P < 0.01, ***P < 0.001 and ****P < 0.00001, and P = 1.013 × 10−5 (d) and P = 1.9 × 10−10 (e). Whiskers represent the minimum and maximum values within 1.5× IQR. Q1, Q2 and Q3 correspond to the 25th, 50th (median) and 75th percentiles, respectively. f, Distribution of clinicopathological and molecular features shown in a in TP53-mutated and TP53-wild-type subgroups. Two-sided Fisher’s exact test, ****P < 0.0001, ***P < 0.001, **P < 0.01 and *P < 0.1; that is, P = 2.6 × 10−5 for H3 status, P = 9.3 × 10−5 for PPM1D mutation, P = 1.7 × 10−7 for ACVR1 mutation, P = 6.9 × 10−4 for MYC gain, P = 0.01 for PIK3CA mutation, P = 1.6 × 10−7 for survival, P = 2.8 × 10−12 for chr1q gain, P = 0.07 for chr2 gain and P = 2.7 × 10−11 for chr1q and chr2 gain. g, OS from the date of randomization in patients with TP53MUT and TP53WT DIPG (P = 1.25 × 10−8, two-sided log-rank test).
Source data
We found predicted pathogenic germline variants in 103 patients. As recently published31, we found recurrent alterations in PTCH1 (n = 3) and in homologous recombination genes—that is, PALPB2 (n = 1), BRCA2 (n = 2), BRCA1 (n = 1) and CHEK2 (n = 1). Recurrent pathogenic germline mutations not previously reported were detected in calcium channel genes for 15.7% of patients: RYR2 (n = 8), RYR1 (n = 5), RYR3 (n = 4), CACNA1G (n = 4), CFTR (n = 4) and ITPR3 (n = 5) (Extended Data Fig. 5a).
Analysis of this large cohort refined the frequency of SNVs in DIPG (Fig. 3a). In total, 133 tumors (77%) harbored H3.3-K27M, including 89 with TP53 mutations. Twenty-nine patients (17%) had H3.1-K27M (27 H3C2-mutated and two H3C3-mutated), of whom 19 carried ACVR1 mutations and seven carried TP53 mutations. Ten tumors (6%) were H3-WT after sequencing (Supplementary Table 8).
The five most frequently mutated genes, in addition to H3 genes—TP53 (57%), ACVR1 (17%), ATRX (13%), PPM1D (11%) and ASXL1 (9%)—showed an average VAF higher than 35%, without being associated with frequent chromosomal gain, suggesting early events during oncogenesis (Fig. 3b and Extended Data Fig. 5b). A similar observation was made for CDKN2C, ZEB2, PTPN11, PTEN, F8 and ATM although in few patients (≤6) only. By contrast, high PDGFRA mutation VAF was associated with gain/amplification in three of six patients, as previously reported14.
SNVs were evenly distributed across OS groups, except for a significant enrichment of H3.3-K27M and TP53 mutations and, to a lesser extent, MYC gain in patients with an OS less than 1 year (n = 107, ‘short survivor’ subgroup) (Fig. 3c). NF1 alterations were restricted to this group.
PIK3CA (18%) and PIK3R1 (9%) were the most frequent secondary events, with lower VAFs (Fig. 3b). PIK3R1 alterations were found in both H3.3-K27M and H3.1-K27M cases, with significant co-occurrence with ACVR1 alteration (Extended Data Fig. 5c). PIK3CA alterations occurred across all OS groups, as previously described32, with mutual exclusivity (P < 0.01) with H3.3-K27M. H3.1-K27M was more frequent in younger patients, whereas TP53 mutation and MYC gain were enriched in older patients (Extended Data Fig. 5d,e).
First, using unbiased pathway analysis for genes mutated in more than three patients (n = 92), we found significant enrichment of processes related to tube morphogenesis (adjusted P = 8.158 × 10−10) and cell adhesion (adjusted P = 9.06 × 10−7). Next, analysis of frequently altered DMG pathways and functions (Extended Data Fig. 5f) showed that 170 of 172 tumors (98.8%) harbored mutations or copy number variations (CNVs) in at least one of the following seven of them: RTK/MAPK, PI3K/MTOR, TGFB signaling, genome integrity, chromatin regulators, protein homeostasis and transcription factors.
Single-base substitution (SBS) signatures analysis revealed SBS1 (deamination of 5-methylcytosine), SBS6 and SBS15 (defective DNA mismatch repair) and SBS87 (thiopurine chemotherapy exposure) as the most prevalent (Extended Data Fig. 5g,h). This profile differed from previous reports based on smaller selected subsets of tumors33,34.
DIPG somatic copy number alteration landscape is associated with TP53 mutation
When analyzing genome-wide copy number alterations (CNAs), we observed marked and previously unreported heterogeneity among patients with DIPG. This included frequent whole chromosome arm losses (chr14 and chr16) associated with complex genomic rearrangements across the genome or whole chromosome arm gains (chr1q and chr2) usually associated with fewer structural alterations (Supplementary Fig. 1). Ranking samples by the number of CNV segments emphasizes that widespread complex rearrangements are more common in tumors harboring TP53 mutations (Fig. 3d, Extended Data Fig. 6b and Supplementary Fig. 1).
Copy number signature analysis highlighted a strong enrichment of CN9 in TP53-mutated tumors (Extended Data Fig. 6b,c). CN9 has been linked in pan-cancer analyses to poor survival and to TP53 mutations35. CN12 and CN13 were also enriched in TP53-mutant tumors but occurred in few patients (n = 21 and n = 15, respectively) and were positively associated with MYC gain (Extended Data Fig. 6d). CN9 and CN12 show prominent focal loss-of-heterozygosity (LOH) patterns suggestive of structural chromosome instability (CIN)35, whereas CN13 arises from cancers with LOH affecting over 70% of the genome, with whole-arm-scale or whole-chromosome-scale LOH events. CN9 correlates positively with chromothripsis state score, whereas the ‘diploid’ signatures CN1 and CN19 were negatively correlated with TP53 mutations and positively with chr1q gain (Extended Data Fig. 6c,d).
TP53 pathway alteration is not limited to dominant-negative missense mutations
Most somatic TP53 mutations affected the DNA-binding domain, mainly hotspot missense mutations, usually associated with a dominant-negative effect36 (Extended Data Fig. 6e). Overall, 37% of patients harbored LOH associated with a point mutation or indel and 5.8% with compound heterozygous mutations, and 2.3% carried subclonal compound heterozygous mutations. Eleven patients had a truncating exon-6 mutation in PPM1D, known to mimic TP53 deficiency in the DNA damage response37 (Supplementary Fig. 1 and Extended Data Fig. 6f,g). Twenty-five patients exhibited copy number loss of TP53BP1, a DNA double-strand break sensor involved in non-homologous end joining (NHEJ) DNA repair38,39. This loss, affecting a large portion of the chr15 arm (median 60%), was associated with significantly reduced expression of this gene (P < 0.05) and may impair TP53 function. The number of segments detected increases gradually with TP53 dysfunction. Tumors with TP53 alterations had a significantly higher proportion of genome altered than TP53WT tumors. TP53MUT and TP53WT tumors had a median proportion of genome altered of 26% and 13%, respectively (Fig. 3e). Finally, H3.3-K27M/TP53-mutant tumors displayed increased segment numbers and oscillations, consistent with chromothripsis (Fig. 3d and Extended Data Fig. 6f,g).
TP53 mutations were early events in DIPG with high VAF (median 0.75 (75% CI: 0.56−0.87); Fig. 3b) and were associated with MYC gain (31.3% versus 9.5%, χ2 test, P = 6.88 × 10−4) but were rare in tumors with chr1q gain or concomitant chr1q and chr2 gains (χ2 test, P = 2.83 × 10−12 and P = 2.79 × 10−11, respectively) (Fig. 3f).
TP53-mutant tumors more frequently harbored PTEN, SOX10 and F8 mutations, whereas ATM mutations predominated in TP53WT (Extended Data Fig. 6j).
TP53 mutations were associated with poor OS and characteristics of aggressiveness at transcriptional level
Median OS of patients with TP53-mutated tumors was 8 months compared to 15 months in patients with TP53-wild-type tumors (Fig. 3g).
RNA-seq revealed marked upregulation of genes involved in cell cycle regulation (Benjamini−Hochberg false discovery rate (FDR) P = 2.43 × 10−11), chromosome segregation (Benjamini−Hochberg FDR P = 4.47 × 10−16) and DNA repair (Benjamini−Hochberg FDR P = 3.14 × 10−13) in TP53MUT tumors, alongside downregulation of genes involved in neuron ensheathment (Benjamini−Hochberg FDR P = 6.06 × 10−4) and glial cell development (Benjamini−Hochberg FDR P = 1.1 × 10−3) (Extended Data Fig. 6k). Additionally, oligodendrocyte markers and ectoderm development genes were repressed in TP53-mutant tumors. Transcriptional profiling revealed a shift toward a more immunosuppressive phenotype in TP53-mutant tumors. Notably, KLRC1, the most upregulated gene, encodes the natural killer cell inhibitory receptor NKG2A, a key natural killer cell immune checkpoint, inhibiting the killing ability of CD8+ T cells. NKG2A specifically binds the non-classical human leukocyte antigen (HLA) class I molecule HLA-E, thereby transmitting inhibitory signals that impair natural killer cell function40.
We observed marked downregulation of the long non-coding RNA GATA2-AS1 in TP53MUT tumors (fold change = 0.46, adjusted P = 2.08 × 10−5), an MYC target, that represses GATA2 expression and limits tumor aggressiveness41,42,43. By contrast, TP53-mutant tumors showed upregulation of the MET protooncogene, which promotes the proliferation, invasion and poor prognosis44 in glioblastoma (GBM) and is amplified in a subset of DIPG14.
Description of the four very-long-term survivors still alive 6 years from diagnosis
Four patients were alive at last follow-up, 6 years or more after diagnosis, without meaningful sequelae. Three showed residual MRI abnormalities at the last evaluation, whereas one achieved complete remission with clearance of all TP53-mutant fluid-attenuated inversion recovery (FLAIR) hypersignals (Extended Data Fig. 7a). All patients had been off treatment for at least 1 year and were treated with an mTOR inhibitor: two received everolimus and two received sirolimus due to toxicity or initial progression. No shared alterations except H3-K27M were found. One patient harbored a clonal IDH1 R132C mutation. Two additional patients in the trial cohort (1.16%) with both IDH1 R132C and H3.3 K27M mutations survived more than 2 years: one not randomized and treated with everolimus was alive at the last follow-up, and one treated with dasatinib died after 2.19 years.
Gene expression profile of 13 long-term survivors
We compared transcriptional profiles of 13 long-term survivors with those of 82 short-term survivors. RNA-seq revealed significant downregulation of genes involved in cell cycle regulation and chromosome segregation in long-term survivors, along with upregulation of genes involved in mitochondrial electron transport and oxidative phosphorylation (Supplementary Fig. 2 and Extended Data Fig. 7b,c).
The top 10 most downregulated genes in long-term survivors were CENPE, E2F7, GPR139, IL5RA, NTS, NLRP11, POLQ, SLC7A3, CLS2A3P1 and UPS32P2. E2F7 promotes tumorigenesis via EZH2 activating the PTEN/AKT/mTOR pathway45,46. In DMG, E2F7 and EZH2 expression were positively correlated and elevated in short-survival patients (Extended Data Fig. 7d), and several E2F7 targets (n = 24, adjusted P < 0.01) were reduced in long-term survivors (Supplementary Fig. 2a). By contrast, the most upregulated genes in long-term survivors were BCYRN1, CARD11, CHRM5, HYAL1, IFI27L2, ITPK1, MRO, PDXK and TNFSF12.
We next analyzed signaling pathways by computing individual gene set variation analysis (GSVA) scores for Hallmark gene sets from the Molecular Signatures Database (MSigDB) and identified those significantly modulated between patients with long versus short OS, selecting one representative pathway per biological process (Fig. 4a). Long-term survivors showed inhibition of pathways related to DNA repair, G2/M checkpoints, MYC targets, Notch signaling, oxidative stress-induced senescence and TP53 signaling. Genes involved in PI3K/AKT/MTOR signaling were also significantly downregulated in patients with long OS. SNVs in this pathway were present across all survival subgroups. However, long-term survivors harbored no alteration in PI3KCA or NF1, showing only in-frame deletion of PIK3R1 (Extended Data Fig. 4a). None of the long-term survivors with RNA-seq data showed activation of the MAPK pathway nor a BRAF V600E or FGFR1 mutation.
a, Pathways showing significant modulation of GSVA scores between long and short survivor groups. b, Cell type composition of DIPG tumors inferred from deconvolution of bulk RNA-seq data (upper panel). The proportion of each cell type is indicated by a color code. OS (in months) is shown as a dot plot in the lower panel. c, Violin plot showing tumor cell type composition in short and long OS subgroups. Two-sided t-test, Holm−Bonferroni-adjusted P values: *P < 0.05, **P < 0.01; P = 0.024 for OPC; P = 0.29 for TAM2 and P = 0.008 for Prol. DC, dendritic cell; NK, natural killer; OPC, oligodendrocyte progenitor cell; Prol, proliferative TAM; TAM1, monocyte-derived TAM; TAM2, microglia-derived TAM.
Source data
Patients with DIPG surviving longer have a distinct tumor immune microenvironment
Several differentially expressed genes according to survival supported differences in the tumor microenvironment (TME). TNFSF12, secreted by microglia-derived tumor-associated macrophages (TAMs), was significantly upregulated in long-term survivors (Supplementary Fig. 2a and Extended Data Fig. 7c)47; the TNFSF12−TNFRSF12A axis has been implicated in glioma progression and microglial activation in adult GBM. As previously reported, DMGs show myeloid infiltration48. Deconvolution of bulk RNA-seq data revealed mixed TAM ontogeny in DIPG with enrichment of microglial TAMs (Mg-TAMs) in long-term survivors, whereas monocyte-derived TAMs (Mo-TAMs), proliferative TAMs and oligodendrocyte progenitor cells were enriched in short-term survivors; overall myeloid infiltration was not associated with survival (Fig. 4b,c). Expression of the immunosuppressive cytokines KLF2 and TGFB1 tended to be reduced in long-term survivors (fold change adjusted P = 0.04 and fold change adjusted P = 0.01, respectively). Patients with a prolonged survival also showed higher proportions of cells with a ‘myeloid cell proliferation’ signature from Andrade et al.48 as well as elevated resting CD4/CD8 T cell and CD4 naive central memory resting T cell signatures (Fig. 4c). Finally, HOXA7 was strongly downregulated in long-term survivors; its overexpression correlates with TAM infiltration in The Cancer Genome Atlas (TCGA) and in esophageal squamous cell carcinoma (ESCC) samples49.
Correlation between response to the treatments and clinical characteristics or biomarkers
The protocol included an exploratory analysis for efficacy biomarkers for each of the three drugs. This analysis was restricted to patients randomized to everolimus or dasatinib, as too few patients received erlotinib. The prespecified biomarkers did not influence the response to everolimus and dasatinib (Extended Data Fig. 4b,c). In addition to GSEA Hallmark PI3K/AKT/MTOR signaling, we considered alterations in a curated list of PI3K/AKT/MTOR pathway genes (Supplementary Table 9) and chr1q gain. Forest plots (Fig. 5a,b) showed association between biomarkers and treatment response (significance threshold P < 0.1). Chr1q gain was associated with improved PFS (P = 0.05) and OS (P = 0.035) with everolimus (Extended Data Fig. 4d–g). Similarly, mutations in PI3K/AKT/MTOR pathway correlated with better PFS (P = 0.02) and OS (P = 0.08) in everolimus-treated patients (Fig. 5c–f). Overexpression of the PI3K/AKT/MTOR signature showed a trend toward improved response to everolimus (interaction P = 0.1085). PIK3CA alterations were enriched in patients with high GSVA score for mTOR pathway expression (Fig. 5g and Extended Data Fig. 4g).
a, Forest plot comparing the differential effects of everolimus and dasatinib on OS according to the presence or absence of relevant biomarkers among the 147 patients with biological evaluation allocated to the everolimus versus dasatinib comparison (n = 67 versus n = 80, respectively). Chrom_1q_2gain indicates concomitant gains of chromosome 1q and 2 together. HPI3K refers to the level of expression of the PI3K/AKT/MTOR pathway. AKTmTOR denotes the presence or absence of mutations in the PI3K/AKT/MTOR pathway. Interaction P values were considered significant when lower than 0.1. The diamond represents the overall treatment effect and its width the 95% CI. Each horizontal line represents the hazard ratio (square) and its 95% CI of treatment effect for each modality of a covariate. An arrow indicates that the upper limit of the 95% CI is outside the margins. The vertical line represents the null hypothesis of no treatment effect (hazard ratio = 1). NS, no significant modulation of the level of expression of genes belonging to PI3K/AKT/MTOR. b, Forest plot comparing the differential effects of everolimus and dasatinib on PFS according to the presence or absence of relevant biomarkers. To confirm the impact of theranostic biomarkers on the response to everolimus, we compiled an independent cohort of 31 patients with DIPG with available sequencing data who were treated at Gustave Roussy with sirolimus or everolimus either before trial initiation or after randomization was stopped. The diamond represents the overall treatment effect and its width the 95% CI. Each horizontal line represents the hazard ratio (square) and its 95% CI of treatment effect for each modality of a covariate. An arrow indicates that the upper limit of the 95% CI is outside the margins. The vertical line represents the null hypothesis of no treatment effect (hazard ratio = 1). c, OS in patients without PI3K/AKT/MTOR alterations, stratified by randomized treatment. d, OS in patients with PI3K/AKT/MTOR alterations, stratified by randomized treatment (two-sided interaction test). e, PFS in patients without PI3K/AKT/MTOR alterations, stratified by randomized treatment. f, PFS in patients with PI3K/AKT/MTOR alterations, stratified by randomized treatment (two-sided interaction test). g, Oncoplot showing SNV alterations in genes involved in PI3K/AKT/MTOR pathway in DMG, ranked from left to right according to decreasing GSVA pathway scores. SNV alterations from patients without available RNA-seq data are shown on the right. Evts, events; HR, hazard ratio; Pts, patients.
Source data
For external validation, we analyzed an independent cohort of 19 patients with biologically documented DIPG treated with an MTOR inhibitor either before BIOMEDE initiation or after the completion of randomization. Patients with overexpression of the PI3K/AKT/MTOR signature had a better OS and PFS (Extended Data Fig. 4h,i).