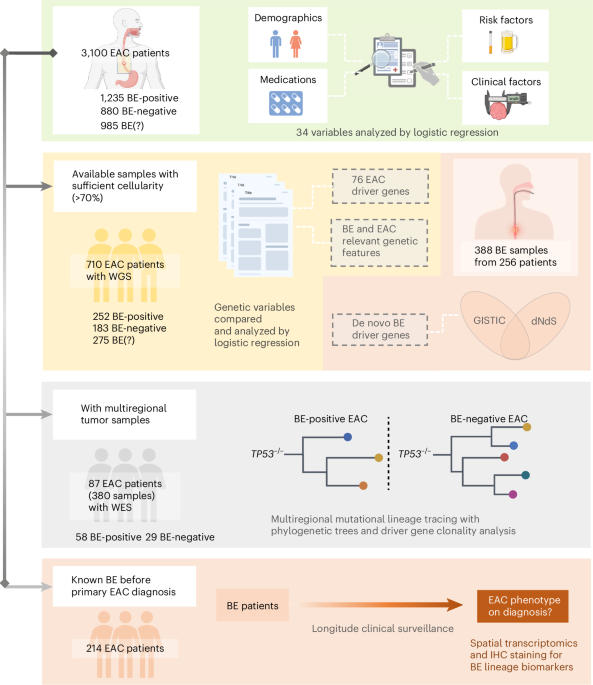
Patient cohorts and study design
A multicenter consortium, Oesophageal Cancer Clinical and Molecular Stratification (OCCAMS), prospectively recruited a cohort of 2,115 patients with EAC on a curative pathway across 25 UK centers18. This included gastroesophageal junction cancers, but gastric cardia cancers were excluded. OCCAMS was registered and approved by relevant research ethics entities (East of England Ethics Committees, UKCRNID-8880, REC 07/H0305/52 and 10/H0305/1). Study sites were trained to include details of a prior history of BE and any evidence of BE on diagnosis. In addition, the endoscopic, surgical and histopathological reports were manually reviewed to classify each case phenotypically according to the presence or absence of macroscopic or microscopic evidence of BE metaplasia adjacent to the tumor. For precancerous BE samples used as a comparison, we used cases from the BEST2 and Barrett’s Biomarker studies (East of England Ethics Committees 10/H0308/71, LREC01/149 and registered in the Integrated Research Application System (IRAS) (ID 57563, 15949)).
All participants in OCCAMS, BEST2 and Barrett’s Biomarker studies provided individual informed consent, and data were pseudonymized.
Selection of cases
The inclusion criteria for the OCCAMS study select patients with a confirmed diagnosis of adenocarcinoma of the esophagus and gastroesophageal junction who were fit enough for treatment on a curative pathway, which was generally neoadjuvant chemotherapy and surgery. For these cases, we aimed to collect pretreatment samples for sequencing, but, where this was not possible, a sample was taken from the surgical resection specimen. For patients with early disease, treatment comprised endoscopic therapy (endoscopic mucosal resection with or without radiofrequency ablation). A small number of patients with advanced-stage disease were included who were initially deemed to be curative but in whom the full staging showed more advanced disease not suitable for a curative pathway.
Comprehensive clinical research guidelines were developed in the Fitzgerald research group and followed by trained clinical and research staff at all OCCAMS study centers. At each study center, eligible patients with EAC were identified and approached regarding participation in the OCCAMS study and their desire to join as participants. Alternatively, OCCAMS research staff in the clinic asked the patient’s permission to be contacted with more information via mail and a follow-up phone call. Patients had the opportunity to ask questions and think about their involvement by talking to their general practitioner, for example, and could return the signed form later. Consent was obtained from patients to contact their general practitioner to inform them of their participation in OCCAMS and to obtain relevant medical information from the cancer registries and other National Health Service (NHS) data controllers.
Patients with pathologically assessed tumors and diagnosed with adenocarcinoma of the esophagus between 2002 and 2022 were included. Patients with tumor histology other than EAC were excluded, and a majority (n = 233, 67%) were esophageal squamous cell carcinoma (ESCC) cases. Furthermore, patients with ‘open & shut’ surgery with more advanced disease than expected were also excluded. This was because a tumor sample was generally not collected for these patients; therefore, tumor phenotype ascertainment would have not been possible. In addition, few data were recorded on the baseline questionnaire forms for such patients. A small number of cases (n = 22, <1%) were missing age or gender, and these were excluded.
Pathology review
A strict expert pathology review was performed for all cases. At least two pathologists reviewed each EAC case: one pathologist from the referring study center and another pathologist from the OCCAMS central study center at Cambridge University Hospitals who had more than 20 years of experience in upper gastrointestinal cancer. Tumors were staged based on International Union Against Cancer/American Joint Committee on Cancer (UICC/AJCC) TNM guidelines (7th edition)31. The T, N and M stages were assigned using the available information in the patient’s medical records, including clinical chart notes, endoscopic ultrasound, positron emission tomography, endoscopic mucosal resection (EMR) and histopathological reports after surgical resection. We used the most advanced stage prior to or at the time of surgery for patients who received neoadjuvant therapy.
The presence of BE adjacent to EAC for OCCAMS cases was based on one of two criteria—endoscopic (macroscopic) visual changes observed at prestaging evaluation with pathology review showing intestinal metaplasia at the time of surgical resection, which was assessed by expert gastrointestinal pathologists of the recruiting OCCAMS sites. Intestinal metaplasia was also identified in cases without macroscopic evidence of BE upon expert review of the pathology specimen. All pathologists followed a specific synoptic report proposed by the College of American Pathologists. Additionally, pathologists followed the OCCAMS study protocol, which required thorough assessment for BE in the proximal and distal resection margins and tumor. Tumor sampling was done for all borders of the resected tumor and the tumor bed to minimize sampling error. The number of biopsy specimens varied based on tumor size.
Clinical data collection
Trained research staff collected baseline characteristics using chart review or during structured face-to-face interviews using a uniform case report form (CRF) across the 25 sites in the OCCAMS Consortium. All covariates used in the analysis originated from the study CRF. Patient baseline characteristics were collected on demographic, anthropometric and environmental exposures. Weight and height were measured objectively at the baseline visit or from the next closest record to the baseline. Gender was determined based on self-reported questionnaire data. Overall survival time (in years) was calculated from the date of diagnosis to the date of death or the date the patient was last seen in the clinic. Vital status was ascertained from all-cause mortality. All patients who consented to participate provided the minimum reporting standard, which required demographic and clinical details.
Research staff transcribed and entered data captured on the OCCAMS CRFs into the study database. These data were anonymized and stored in a secure central database hosted on Cambridge University Hospitals NHS Foundation Trust servers. Several data management issues should be noted as the data collection process may introduce errors or biases. Errors during the baseline interview (for example, failure to ask questions or record a response) or lack of information in the case notes/electronic records may contribute to missing data. As many patients are of advanced age, recall bias may also introduce discrepancies (for example, answers to history of heavy drinking or smoking).
Data preparation and variable construction
Processing of baseline clinical and epidemiological data
The raw and fully anonymized baseline data for OCCAMS (R data file format, .Rdata) and BEST2 (comma-separated values file format, .csv) were exported to a university-furnished computing device in June 2022. The files containing the data for OCCAMS and BEST2 were collated, processed and screened for completeness, accuracy and consistency. Data were cleaned, removing or correcting any inconsistencies, inaccuracies or implausible values. All pragmatic strategies to minimize missing data were implemented. The cleaned dataset was then carefully checked against the raw data to ensure quality data preprocessing. The datasets and data-cleaning code were saved as plain-text files and tracked and managed using version control software (Git/Subversion). All data processing was carried out using R version 4.2.3 on macOS Ventura 13.3.1.
The following common methodology was used to clean data for both OCCAMS and BEST2 studies. Due to the inclusion criteria, age at diagnosis and gender were complete. Ethnicity was recoded into ‘white’ or ‘other’ as there were too few observations in other ethnicity codes, which is not unexpected for the BE/EAC patient population.
The age at diagnosis for EAC and BE cases and age at recruitment for reflux controls, as well as BMI at baseline, were categorized into groups. This was done to create a more meaningful comparison for these measures. Age at diagnosis was categorized into four groups: younger than 50 years, 50−59 years, 60−69 years and 70 years or older. BMI was calculated using the baseline weight and height (kg m−2), and BMI categories were defined according to standard ranges of underweight (<18.5), normal weight (18.5−24.9), overweight (25−29.0) and obese (≥30). Underweight cases were included among normal weight owing to very small frequencies in the cohort (<2.5%). The continuous distribution and the grouped frequencies were used in descriptive analyses, and only the categorical variables for age and BMI were included in regression models.
Cigarette smoking was collapsed into a binary variable, with ‘former’ and ‘current’ recoded as ‘ever’ smoker and ‘never’ remaining as defined. Additionally, if the average number of cigarettes per day was recorded as zero, smoking status was set to ‘never’; if it was a non-zero value, then smoking was set to ‘ever’. The number of pack-years was calculated by multiplying the number of packs of cigarettes smoked per day by the number of years of smoking.
The self-reported responses for medication use frequency of aspirin, non-steroidal anti-inflammatory drugs (NSAIDs), proton pump inhibitors (PPIs), H2-receptor antagonists (H2RAs) and over-the-counter (OTC) acid suppressants included ‘Never’, ‘No’, ‘Past Use’, ‘Occasional Use’ and ‘Current Use’. However, responses such as ‘Past Use’ and ‘Occasional Use’ are open-ended, so, to mitigate this issue, responses were recoded to binary ‘Ever Use’ and ‘Never Use’. The duration of medication use was recorded in years, months, weeks and days. The total duration of use (in years) was calculated for each medication type by summing the individual measures. Additionally, if the frequency was set to ‘Never’ and a non-zero total duration of use was reported, the total was set to null. Conversely, if a non-zero duration was recorded, then the frequency of use was set to ‘Ever Use’. Aspirin and NSAID frequency of use were combined into a single variable measuring use of either medication. Similarly, a single variable for the use of any acid suppressant medication was derived using the frequency of use of PPIs, H2RAs and OTC acid suppressants.
Alcohol intake was recorded as the number of units of beer, wine and spirits consumed per week. These individual measures were summed into a single continuous variable for the total number of alcoholic drink units consumed per week. Heavy drinking status was self-reported by patients in both studies.
Frequency of reflux symptoms was reported as ‘Never’, ‘Sometimes’, ‘Often’, ‘Daily’ and ‘Unknown/sporadic’. Duration of reflux symptoms was harmonized into an ordinal variable with four ranges (Never, 5 years, 5−10 years, >10 years and Unknown/sporadic). A single variable was created that combined all measures related to reflux symptoms and acid suppressant medication use. This variable is referend to as the ‘derived heartburn symptoms status’ variable (Supplementary Fig. 3). In addition, a single variable for use of any acid suppressant medications was derived based only on the acid suppressant medication use variables (patient on acid suppressant and use or duration of PPIs, OTC acid suppressant medications or H2RAs).
For variables that contained responses with undefined free text or numeric ranges instead of a single value, either the response was set to missing or the mid-range was calculated. For example, a free text input of ‘undistilled only’ for total alcohol unit intake was set to missing, and a response of ‘3−5’ cigarettes per day was recalculated to ‘4’ per day. Continuous variables where a negative numeric value was recorded were recoded to missing as per CRF instructions.
The UK regions for OCCAMS and BEST2 study centers were determined based on their locations and classified using the International Territorial Level 1 (Office of National Statistics). Finally, for EAC cases only, combined TNM staging was created according to UICC/AJCC 7th edition guidelines31.
Variable selection
After baseline data cleaning and screening in the OCCAMS cohort and as informed by the results of the literature review, a total of 32 variables across five domains were deemed relevant and included (Supplementary Table 2). To select variables for inferential analysis, a purposeful selection process was followed:
-
1.
Unconditional logistic regression was used to obtain univariable odds ratios and 95% confidence intervals for the association of each variable with BE-negative EAC compared to BE-positive EAC cases.
-
2.
Variables with P < 0.25 and missing data <60% overall were preselected and included in a multivariable logistic regression model with BE-negative EAC as the outcome compared to BE-positive EAC.
Only variables with P < 0.05 or those deemed to have epidemiological or clinical importance were selected in the final stage. Directed acyclic graphs were also used to determine which variables should be included as potential confounders.
WGS
WGS with a matched germline reference was available for one EAC sample from each of 710 patients and for 388 samples across 256 patients with BE. WGS was performed on as many cases as possible, but, prior to inclusion, each sample had to pass a strict pathology consensus review on a fresh-frozen section and show cellularity greater than 70%. Methods for sample quality control, DNA extraction and WGS were as previously described12,19, and cohort details, single-nucleotide variant (SNV), copy number and structural variation analysis are detailed as follows.
BE cohort
The BE cohort includes 388 samples with 205 sequenced from single BE samples and 79 pooled sequencing from 183 multilevel samples. There are sequencing data from 284 endoscopies of 256 patients in total. For those pooled for sequencing, we took the highest pathology as the grade annotation. This cohort was selected to capture early genomic events preceding invasive EAC. Specifically, there are 51% (145/284) non-dysplastic Barrettʼs esophagus (NDBE) and 49% (139/284) dysplasia, including 24% (68/284) low-grade dysplasia (LGD) and 25% (71/284) HGD, including very few T1a samples. Patients with NDBE as their highest pathology were followed for a median of 3.1 years (IQR: 2.2−4.4) until their last surveillance endoscopy. Most dysplastic lesions were treated at diagnosis, whereas a few historical untreated LGD cases were followed for a median of 1.3 years (IQR: 0.8–2.7). These precancerous patients were from the BEST2 and Barrett’s Biomarker studies (East of England Ethics Committees 10/H0308/71, LREC01/149 and registered in the IRAS (ID 57563, 15949)).
BE and EAC sample processing
Strict pathology consensus review was observed for these BE and EAC samples, with a minimum 70% cellularity requirement before inclusion. All tissue samples were snap frozen. For the OCCAMS study, peripheral blood was used as the germline reference, and, in cases where this was not possible, a sample of normal squamous epithelium located at least 5 cm away from the lesion or normal duodenal tissue was used instead, according to standard practice.
Methods for sample quality control, DNA extraction and WGS were as previously described12,14,19. In brief, the 710 EAC and 205 BE samples sequenced by Illumina, the Cancer Research UK Cambridge Institute and the Wellcome Sanger Institute underwent WGS to a target depth of 50×. Matched germline samples were sequenced to a target depth of 30×. Reads were then aligned with BWA-MEM to the 1000 Genomes Project version of the GRCh37 human reference genome32. Each of the 79 BE sample pools and matched germline samples sequenced by Genomics England were processed in two aliquots to combined target depths of 150× and 75×, respectively, and reads were aligned to GRCh38. Sequencing quality checks were conducted using the FastQC package (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and polymerase chain reaction (PCR), and optical duplicates were flagged using Picard MarkDuplicates (https://broadinstitute.github.io/picard/) after alignment.
Single-nucleotide and copy number variant calling
For the 710 EAC and 205 BE samples, somatic variants were called using Strelka version 2.0.15 (ref. 33) and annotated using Variant Effect Predictor (VEP) version 78 (ref. 34). Mutation burden was derived from each sample’s VEP files by summing the number of SNVs and indels across the genome. For the additional 79 BE samples sequenced by Genomics England, Strelka version 2.9.4 and VEP version 91 were used, and mutation burden was calculated by taking the average across the aliquot VEP files for each sample. LiftOver was used to convert mutation loci between versions of the human reference genome. Mutations per megabase was calculated using the length of the reference genome (3,137,454,505 bp). GISTIC2.0 was used to detect recurrently deleted or amplified regions of the genome using raw copy number values obtained from ASCAT version 2.1 (refs. 35,36) for the 710 EAC and 205 BE samples and from Canvas version 1.38.0.1554 for the 79 BE samples.
Selection and calling of driver genes
Previously reported driver genes in EAC were derived from genes listed in Frankell et al.19, and genomic regions were identified using Ensembl BioMart37. These gene regions were then used to extract alterations from the outputs of VEP and GISTIC2.0. Driver mutation status was determined based on the alteration type (for example, missense, nonsense or frameshift) using Strelka and VEP. One or more affected copies were deemed as a mutation.
To identify driver genes associated with BE, we predicted SNV mutations using observed/expected mutation ratios calculated by dNdScv. Copy number driver genes were identified by overlapping genes located within peak regions detected by GISTIC2.0 with those in the COSMIC consensus and previously identified EAC driver genes.
Mutational signatures
Mutational signatures discovery within the cohort was carried out using SigProfilerExtractor38 on 997 samples as previously described12. The optimal signature configuration was determined by selecting from a range of signature combinations (from five to 17) based on the highest stability and lowest Frobenius reconstruction error for a signature combination. The optimal configuration was composed of 14 signatures, and its validity was confirmed by independent analysis using Bayesian methodology from sigminer39. Subsequently, deconstructSigs40 was employed to deduce the mutational contributions of these processes to each sample across the entire cohort presented here.
Copy number, whole-genome duplication and aneuploidy
An amplification is defined as a ploidy-adjusted copy number greater than 2, and a deletion is defined when the copy number value is 0. The percentage of aberrant genome is calculated as the proportion of the genome, excluding sex chromosomes, where the rounded copy number does not equal the rounded ploidy. The fraction of loss of heterozygosity is defined as the percentage of the genome where the minor allele frequency is less than 0.5, relative to the entire genome excluding sex chromosomes. Raw copy number values from ASCAT and the PCAWG-11 consensus purity pipeline (https://github.com/PCAWG-11) were used to determine samples with whole-genome duplication based on tumor ploidy and the extent of loss of heterozygosity41. Per-sample ploidy and purity were also inferred using this method.
Identification and classification of amplicon and chromothripsis events
Copy number segments were called using CNVkit version 0.9.8, and regions of amplifications of size 50 kb and copy number >4.5 were used as input for the identification of amplified regions and reconstructed using AmpliconArchitect42,43.
The classification of amplicons into ecDNA and BFB events was done using AmpliconClassifier (https://github.com/AmpliconSuite/AmpliconClassifier). Chromothripsis events were performed using ShatterSeek (https://github.com/parklab/ShatterSeek). High-confidence chromothripsis events were defined as ≥6 interleaved intrachromosomal structural variants, ≥7 contiguous segments oscillating between two copy number states and fragment joins testing positive, plus either chromosomal enrichment or exponential breakpoint distribution. Low confidences were defined as the same structural variant criteria but 4–6 segments oscillating44.
WES
WES was conducted and analyzed in 380 samples from 87 patients with multiregional chemotherapy-naive tumor samples (Extended Data Fig. 4a). Samples were obtained from the macrodissection of FFPE esophagectomy or esophogastrectomy specimens (approximately 0.5 × 0.5 cm). The tumor samples were required to be spatially distinct, so the number of tumor samples depended on the size, quality and spatial arrangement on the specimen. Only specimens that yielded at least two tumor samples were included, and the number of tumor samples ranged from two to six, with a mean of 4.3. The specimens covered all clinical stages, with a skew toward later stage, with 61% being stages III and IV, in keeping with a generally late stage of presentation. FASTQ files were aligned to GRCh37 using BWA-MEM, with duplicates marked by Picard version 2.9.5. Variant calling was performed using GATK Mutect2 version 4.1.7.0, using multisampling and FFPE settings.
Evolutionary trajectory and clonality analysis of driver genes
Somatic mutation clustering was performed using PyClone45 (version 0.13.1) to identify subclonal populations based on SNV data. Clonal phylogenies were reconstructed using ClonEvol46 (version 0.99.11) on the PyClone-derived clusters, excluding clusters with fewer than five mutations. Indels and copy number driver events were added to the trees post hoc. REVOLVER47 (version 1.0.0) was used to determine trajectories of repeated evolution from multiple regions. The number of clonal and subclonal driver and passenger mutations was assessed using dNdScv48 based on the output of PyClone.
To evaluate intratumour heterogeneity (ITH) and clonal architecture across EAC phenotypes, we analyzed and compared the following metrics using the Wilcoxon rank-sum test.
Mutational ITH: ratio of subclonal SNVs to total SNVs
Total driver mutations: non-silent SNVs/indels in previously reported EAC driver genes
Total mutations: non-silent SNVs/indels
Recent subclonal expansion score: the largest terminal node cancer cell fraction (CCF) across all samples in a case—calculated for every potentially valid tree, with the minimum score used in analysis
Clones with driver mutations: number of clones harboring ≥1 driver gene alterations
Truncal mutations: silent and non-silent SNVs present in the founding (truncal) clone
Truncal driver mutations: non-silent SNVs/indels in driver genes located in the truncal clone
Spatial transcriptomics
Spatial transcriptomics landscape of BE and EAC was measured in seven samples (five EAC and two BE; Extended Data Fig. 5a) from six patients with EAC. Samples were fresh-frozen chemotherapy-naive tumor biopsies with well-preserved tissue morphology. Spatial transcriptomics profiling was performed using the 10x Genomics Visium HD WT platform measuring approximately 18,000 genes, which achieves 2-µm resolution, enabling near-subcellular spatial granularity. Frozen tissue was selected over FFPE for Visium analysis because well-preserved fresh-frozen samples generally yield higher-quality RNA and improved transcriptomic signal49,50. We also confirmed that tissue morphology was well preserved in the frozen sections used for this study.
Cohort design and spatial transcriptomics experiment
To investigate the spatial transcriptomics landscape of BE and EAC, treatment-naive patient samples were selected from the OCCAMS database in which fresh-frozen biopsies had been collected at endoscopy and stored at −80 °C. The primary selection criterion was well-preserved tissue morphology, essential for accurately correlating transcriptomic profiles with histological features.
Given that fresh-frozen tissue may exhibit variable morphological preservation compared to FFPE specimens, all biopsies underwent blinded, standardized pathological evaluation. Hematoxylin and eosin (H&E)-stained sections were reviewed by a gastrointestinal pathologist, and overall morphology was classified as good, medium or poor based on cellularity, the presence of well-differentiated tumor epithelium in EAC cases or clearly identifiable epithelial structures in BE cases and the absence of extensive necrosis, hemorrhage or other forms of tissue degradation.
After this review, seven samples were analyzed for spatial transcriptomics (Extended Data Fig. 4a). Profiling was performed using the 10x Genomics Visium HD platform, which achieves 2-µm resolution for near-subcellular spatial granularity. All histological processing was performed by the Histology Core at the Cancer Research UK Cambridge Institute. Fresh-frozen esophageal biopsy samples were cryosectioned to 10-µm thickness. Libraries were prepared using the Visium HD Gene Expression Slide and Reagent Kit (4 rxns, PN-1000675) (10x Genomics) and sequenced. Sequencing was performed on an Illumina NovaSeq X system at a depth of approximately 300 million reads per sample, sufficient for 2-µm resolution. Raw sequencing data were processed using Space Ranger (version 3.1.3) (10x Genomics) with alignment to the GRCh38 human reference genome. Spatial gene expression matrices were generated and visualized using Loupe Browser (version 8.1.2). Each Visium HD spatial transcriptomics map was co-registered with the corresponding H&E-stained histology image using ImageJ software.
Unsupervised clustering
To identify spatial transcriptional domains within the tissue, unsupervised clustering was performed using the Leiden algorithm51 and overlaid on the tissue image. To assess whether the transcriptionally defined clusters corresponded to histologically distinct regions, H&E-stained slides annotated with cluster assignments were reviewed by a board-certified pathologist. Cluster composition and spatial distribution were compared between BE and EAC samples to examine conserved and stage-specific transcriptional patterns.
Differential gene expression analysis was conducted using the Scanpy package (version 1.11.1) and applied to Leiden clusters52. The Wilcoxon rank-sum test, as implemented in Scanpy, was applied for pairwise comparison. Genes were ranked according to test statistics, and those with an adjusted P value lower than 0.05 were considered significant markers. Multiple testing correction was performed using the Benjamini–Hochberg FDR method53.
Profiling with BE and EAC marker genes
Cell type annotation was performed using established marker genes. Literature-derived gene panels were assembled to identify the major cell populations present in EAC, columnar epithelium, squamous epithelium, stromal and immune populations (Extended Data Table 5). Tissue type annotations were based on the read counts of marker genes, with squamous or stromal tissues assigned according to the dominant expression. For EAC and BE classification, regions expressing EAC-specific genes were designated as ‘EAC regions’, whereas regions expressing only BE-specific genes were labeled as ‘BE regions’. Bins were colored based on tissue type, with more dense regions reflecting a greater number of bins expressing relevant genes and lower density indicating less-abundant expression.
IHC analysis of BE protein markers
TFF3 and REG4 were assessed in FFPE samples (biopsies, EMRs and surgical resections) from 18 patients with EAC (eight BE-negative and 10 BE-positive). Sections (4 µm) were stained using a BOND RX automated stainer (Leica) and scanned at ×40 magnification (Leica, Aperio AT2).
TFF3 and REG4 biomarkers for the histopathological hallmark of intestinal metaplasia20,54 were immunohistochemically assessed. FFPE samples from endoscopic biopsies, EMRs and surgery were obtained from a subset of patients from the Cambridge cohort who were enrolled onto the OCCAMS study. A total of 31 sections from eight patients with BE-negative EAC (one moderate, three moderate to poorly differentiated and four poorly differentiated) and 72 sections from 10 patients with BE-positive EAC (one well differentiated, four moderate and five moderate to poorly differentiated) were stained, with matched age and gender. Pathologist evaluation was used to determine specimens of interest in surgical samples and adjacent and/or longitudinal preneoplastic lesions including BE using H&E staining. All samples were sectioned at 4-µm thickness, and IHC staining was carried out using BOND RX (Leica automated stainer). For TFF3 (Invitrogen), the heat-induced epitope retrieval (HIER) used was 1, with 10 minutes in HIER, and the antibody was used at a dilution of 0.26 µg ml−1. For REG4 (Abcam, ab255820), the HIER used was 2, with 10 minutes in HIER, and the antibody was used at a dilution of 0.26 µg ml−1. All images were acquired using an Aperio AT2 slide scanner (Leica) at ×40 magnification. Pathologist review was performed in all staining interpretations.
Statistical analysis
Logistic regression
Unconditional logistic regression was used to obtain univariable odds ratios and 95% confidence intervals for the association of each variable with BE-negative EAC compared to BE-positive EAC cases. Variables with P < 0.25 and missing data <60% overall were preselected and included in a multivariable logistic regression model with BE-negative EAC as the outcome compared to BE-positive EAC. For each comparison set, crude and adjusted odds ratio and 95% confidence interval were obtained. We performed four separate adjusted analyses per comparison: (1) minimally adjusted for age and gender only, (2) fully adjusted for all covariates, (3) fully adjusted model eliminating heartburn as a covariate and (4) tumor stage adjusted model. We then developed a logistic regression model for 435 patients, integrating the most relevant genomic and clinical data to understand whether the two phenotypes could be differentiated by integrating both datasets.
The statistical independence of the outcomes was assumed based on the absence of repeated events and the binomial distribution of the residual variation. It is rare for this assumption of logistic regression to be violated. To ensure that the assumption of multiplicativity was satisfied, effect measure modification was assessed between BMI and heartburn and between aspirin/NSAID use and heartburn. A priori, it was known that BMI may modulate heartburn. The latter interaction was tested because the heartburn variable was partly derived from PPI use, and NSAIDs may modify the effects of PPIs in relation to EAC55.
As heartburn may be on the causal pathway, if its elimination as a covariate changed the log odds ratio by more than 10%, then it could be considered a confounder. Missing data were coded as indicator variable.
Three sensitivity analyses were performed to assess the robustness of the estimates obtained using the fully adjusted model for each comparison set. The first sensitivity analysis involved excluding any observations with missing data for the variables in the fully adjusted model, adopting a complete case approach. The second sensitivity analysis used estimates derived from multiple imputation data (detailed below). Lastly, a sensitivity analysis was conducted by excluding EAC cases with a history of undergoing BE surveillance.
No statistical method was used to predetermine sample size. However, this study represents, to our knowledge, the largest clinical and genomic cohort to date, with the next largest cohorts having 54% fewer patients with clinical11 and 31% and 49% fewer patients with EAC and BE genomically sequenced, respectively14,19. We did not exclude any data from the analyses. The experiments were not randomized. The pathologist investigators were blinded to the risk factor data (heartburn, smoking and BMI). Other investigators were not blinded to the independent variables and phenotype outcome.
Missing data
Missing data for baseline characteristics were calculated as a percentage of the total number of cases. The percentage of the recorded values is reported as a fraction of complete cases. For variables dependent on the response to other variables, the missing percentage was calculated as a fraction of cases where the first response variable was available. For example, the proportion of missing data for the duration of cigarette smoking was based on the total number of patients who self-reported current or former cigarette smoking.
Multiple imputation was performed on the datasets corresponding to each comparison group to assess how missing data might bias the observed associations. Age and gender were complete and, therefore, not included in multiple imputation. BMI group, cigarette smoking, aspirin/NSAID use, heartburn symptoms and TNM (EAC only) were imputed using multiple imputation by chained equations with the appropriate method selected based on the variable type56.
The missing data were assumed to be missing completely at random, meaning that the probability of a value being missing is not related to other data. This assumption was based on the similar distribution observed for recorded and imputed data. Furthermore, baseline data were collected by numerous research staff, and, based on our experience, we assumed that variations in the order of the CRF questions, completeness of each section and other factors may have impacted the quality and accuracy of the data collected. Therefore, we assumed that systematic exclusion of data was unlikely. The number of imputations (m) was set to the percent value of the variable with the highest amount of missing data in each dataset, which was aspirin/NSAID use, with approximately 50−60% missing data. The number of iterations (n) was set to 20, as typically recommended57.
Multiple correspondence analysis (MCA)
To delineate the genomic differences between BE-positive and BE-negative EAC, we employed MCA focusing on mutations in recognized EAC driver genes. Each mutation type was categorized distinctly to enable comprehensive analysis. This analysis was performed using the FactoMineR package in R, a robust tool for multivariate exploratory data analysis. Visualization of the MCA results was accomplished using the fviz_mca_ind function from the factoextra package.
Non-parametric data, transformations and multiple hypothesis testing
Statistical comparisons between groups were performed using either the Kruskal−Wallis test or the Mann−Whitney U-test, as indicated by the normality of the data distribution. When applicable, data were log transformed to ensure normality. The percentage of driver genes between groups was compared with χ2 testing. In cases where multiple comparisons were made, adjustment for FDR using the Benjamini−Hochberg procedure or Bonferroni correction were applied.
Computing environment
All analyses were performed using R version 4.2.3 on macOS Ventura 13.3.1 with packages ‘rstatix’, ‘mice’, ‘survival’, ‘survminer’ and ‘coxme’.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.