Ethical approval for the HKGP and this study was granted by the central institutional review board (IRB) (HKGP-2021-001 and HKGP-2022-001) and the IRBs of the Department of Health (L/M257/2021), the Joint Chinese University of Hong Kong/New Territories East Cluster (2021.423 and 2023.120) and the University of Hong Kong/Hospital Authority Hong Kong West Cluster (UW 21–413 and UW 23–289).
Participants
For the HKGP, both asymptomatic individuals and symptomatic probands suspected of having a genetic disease were prospectively identified and recruited across a range of medical specialities at the three partnering centers of the HKGI. All participants received pretest genetic counseling and provided informed written consent following the unique three-tier consent and assent model designed by the HKGI43. As described in our pilot study, detailed phenotype information, including family history and symptom onset, was collected and recorded using HPO terms18.
HKGP participants whose samples were subjected to GS, variant calling and classification before November 2024 were included in this study18. Probands with suspected genetic disease(s), together with their family members, and who had finished genetic diagnosis were included in the diagnostic cohort. Unrelated Chinese participants, including both healthy and affected singletons, in addition to parents from duo and trio family structures, were included for the analysis as the HKGP Chinese cohort. Notably, individuals exhibiting phenotypes associated with selected dominant genes (described in the Gene selection section below), as well as participants with offspring demonstrating phenotypes related to selected recessive genes, were excluded from the respective analyses. To ensure unrelatedness, PLINK (version 2.0) was employed to assess the biological sex and the relatedness among the remaining participants in the HKGP Chinese cohort, with one participant removed from each pair (parents will be retained for non-singleton participants) exhibiting a kinship coefficient greater than 0.177 (ref. 41). Participants with conflicting self-reported sex and sequencing data imputed sex (PLINK 2 –impute-sex) were removed for this study. Chinese ethnicity was determined on the basis of self-reported data and validated through ancestry admixture analysis, where Chinese ethnicity was identified as the predominant ancestry using SNVstory44.
Enrollment criteria
-
1)
Undiagnosed disorders
-
a)
The definition for undiagnosed disorders is disorders without a specific diagnosis after thorough evaluation through clinical assessment and routine investigation.
-
b)
HKGP will recruit patients who meet the following criteria:
-
i)
The patient has a medical condition that meets the aforesaid definition.
-
ii)
Consent of the patient is obtained for providing and sharing medical information and samples.
-
iii)
The patient (or parents or legal guardian) agrees to trio testing—that is, blood sample to be taken from patient and both parents. In case trio testing is not possible, the decision will be made based on the relevant specialists’ assessment.
-
i)
-
a)
-
2)
Cancers with clinical clues linked to possible hereditary components
-
a)
The definition is as follows:
-
i)
Having more than one first-degree or second-degree relative with confirmed cancer; or
-
ii)
Developing cancer at a younger age than expected for that cancer type; or
-
iii)
Pediatric patients with cancer; or
-
iv)
Having more than one type of cancer in the same person
-
i)
-
b)
Recruitment criteria for patients with hereditary cancer and genetic predisposition to cancer would be:
-
i)
The patient is pathologically confirmed with cancer that meets the above definition; and
-
ii)
Consent of the patient is obtained for providing and sharing medical information and samples.
-
i)
-
a)
-
3)
Other patients who will benefit from GS (under the theme ‘Genomics and Precision Health’ of the main phase of HKGP)
-
a)
‘Genomics and Precision Health’ is a cohort that aims to improve the health of individuals with and without specific diseases by harnessing the power of genomics technologies. The health of individuals can be improved by genomics technologies according to clinical, personal, economic and system utilities.
-
a)
-
4)
Unaffected first-degree family members aged older than 18 years of the above three cohorts
Exclusion criteria
Exclusion criteria include patients with known genetic cause for their condition or patient/parents/legal guardian/substitute decision-maker unwilling to participate in the study.
GS and variant detection
The detailed workflows for sequencing and data analysis of short-read GS were previously described18. In brief, whole blood (or buccal/saliva when necessary) was collected, and genomic DNA was extracted for polymerase chain reaction (PCR)-free short-read GS using the KAPA HyperPlus Kit and sequenced on Illumina NovaSeq 6000 or X Plus to achieve a mean coverage of ≥29.5×. After passing quality control checks, the GATK-based standard bioinformatics pipeline was used for secondary analysis. In short, reads were aligned to the GRCh38 reference using BWA (version 0.7.17) with duplicate removal via Picard (version 2.27.4), and variant calling for autosomes, sex chromosomes and the mitochondrial genome was performed using GATK HaplotypeCaller, Mutect2 (version 4.2.6.1), CNVKit (version 0.9.9), Manta (version 1.6.0) and ExpansionHunter (version 3.1.2) to detect SNVs, indels, CNVs, SVs and STRs45,46,47,48.
Gene selection
Genes with strong or definitive gene‒disease associations, as classified by Clinical Genome Resource (ClinGen) (‘definitive’ or ‘strong’), Genomics England PanelApp or PanelApp Australia (‘green’), were prioritized. Genes with moderate evidence of association (‘moderate’ in ClinGen or ‘amber’ in PanelApp) were selectively included on the basis of consensus with referring clinicians.
For the dominant disorder-related genes used for the HKGP Chinese cohort analysis, we adopted a reference gene list of 73 dominant genes from the ACMG secondary findings gene list version 3.2 (ref. 16).
For recessive disorder-related genes, we consolidated a comprehensive list of 1,459 genes from multiple well-recognized sources to ensure broad coverage and clinical relevance. These sources included (1) 105 genes from the ACMG-recommended CS pan-ethnic gene list, including HBA1 and HBA2 for Asian individuals15; (2) 1,283 genes from ‘Mackenzie’s Mission’ version 2.2 gene list, derived from a large-scale Australian CS initiative27; (3) 101 autosomal recessive genes associated with treatable inherited disorders49; and (4) 140 additional genes from other commercially available CS panels and relevant published resources. This integrative approach was intended to maximize the clinical utility of our CS protocol by capturing both established and emerging gene‒disease associations. Detailed lists of the dominant and recessive genes are provided in Supplementary Tables 6 and 7.
Variant classification
SNVs and indels
Diagnostic cohort
Following a phenotype-driven diagnostic workflow similar to that used in the HKGP pilot study18, SNVs and indels (<50 base pairs) with allele frequencies <0.005 in gnomAD versions 2.1.1 and 3.1.2 were prioritized via inheritance-based filtering and phenotypic matching with HPO terms through Exomiser50, supplemented by virtual gene panels from Genomics England PanelApp and PanelApp Australia as described above. The pathogenicity of the variants was determined according to ACMG guidelines and up-to-date recommendations from the ClinGen Sequence Variant Interpretation (SVI) Working Group through manual curation. Specifically, mitochondrial variants were analyzed according to the ClinGen Mitochondrial Disease Nuclear and Mitochondrial Expert Panel Specifications to the ACMG/AMP Variant Interpretation Guidelines. Following the HKGP principles of reporting, we reported variants that were classified as P/LP only when their biological effects matched the patient phenotype. Orthogonal validation was performed for all P/LP variants using independent DNA extracted from the original sample. Variants of uncertain significance (VUSs) in dominant genes that meet the following criteria, agreed upon by all parties in the multidisciplinary team, including clinicians, were reported: highly compatible with the clinical phenotypes and when additional secondary assay/analysis—such as RNA sequencing, enzyme activity testing, immunohistochemical staining, imaging studies and segregation analysis—can be performed to confirm the diagnosis. Variants were visualized using Integrated Genomics Viewer (IGV) version 2.17.4 (ref. 51).
The HKGP Chinese cohort (recessive and dominant genes)
In addition to diagnostic findings, SNVs and indels in our consolidated gene lists for other clinical findings were retained for curation if their allele frequencies were <0.05 in gnomAD version 3.1.2 unless they were included on the BA1 (‘standalone benign’) criterion exception list. Through a combination of automated and manual curation (Supplementary Fig. 4), these variants were classified into three categories: reported P/LP, ACMG P/LP and ACMG VUS or benign (ACMG VUS-B).
-
a.
Reported P/LP
P/LP variants from ClinVar with three-star or four-star review status were classified by expert panels such as ClinGen or authoritative consortia such as the Evidence-based Network for the Interpretation of Germline Mutant Alleles (ENIGMA). In addition, to reduce the total number of variants for manual review, one-star or two-star review status variants were also classified as reported P/LP for recessive genes.
-
b.
ACMG P/LP and VUS or benign (VUS-B)
Other identified variants were processed through two analytic pipelines: (1) both ClinVar-reported and novel variants in the dominant gene list were classified using ACMG/ClinGen guidelines and a Bayesian classification framework; (2) ClinVar-unreported null variants in the LoF genes were classified using the PVS1 criterion. All ClinVar data were accessed and extracted on 30 June 2024.
For the variants detected in the HKGP Chinese cohort, the classification process was further refined using our previously established semiautomated brief cohort analysis workflow (S-BCAW)52. Both automated scoring and manual curation were applied throughout the curation process. For recessive genes, null variants absent from ClinVar were assigned PVS1 criterion using AutoPVS1 (version 1.1) and classified similarly21.
SVs and CNVs
Diagnostic cohort
A phenotype-driven diagnostic workflow similar to that used in the HKGP pilot study was followed. The pathogenicity of deletions and duplications was interpreted in accordance with the joint consensus standards of CNV interpretation by the ACMG and ClinGen53. Currently, there is no established expert consensus for the interpretation of other SV types. For these variant types, PVS1 was applied at an appropriate strength on the basis of the predicted impact on gene function54.
The HKGP Chinese cohort (recessive and dominant genes)
The analysis of SVs focused specifically on genes identified in the predefined gene list, where the disease mechanism is LoF. Insertions, deletions and duplications within these gene lists were curated according to the ACMG/ClinGen joint consensus guidelines for CNV interpretation53.
Among the recessive disorder-related genes, some loci present unique technical challenges that cannot be reliably detected by conventional variant callers, as described above. To overcome these limitations, specialized approaches were employed: an in-house developed caller was used for detecting common α-globin gene deletions (HBA1/HBA2), and Illumina’s SMNCopyNumberCaller was used for precise quantification of SMN1 and SMN2(ref.55).
STRs
STRs were analyzed at loci defined by the Illumina repeat catalog (https://github.com/Illumina/RepeatCatalogs). STR calls were considered pathogenic if the repeat size was greater than the pathogenic reportable threshold summarized in gnomAD on the basis of the literature.
Defining GCF and cGCF
To characterize carrier frequencies at the gene level, we adopted the concept of GCF, defined as the fraction of participants carrying any P/LP variant(s) in the gene.
To facilitate further analysis across groups of genes, we introduced the concept of cGCF, which is defined as the sum of GCFs for all genes within a specific gene list or tier. These metrics provide a robust framework for quantifying carrier frequencies at multiple levels of granularity, enabling population-specific insights and facilitating tier-based gene classification.
Clinical utility
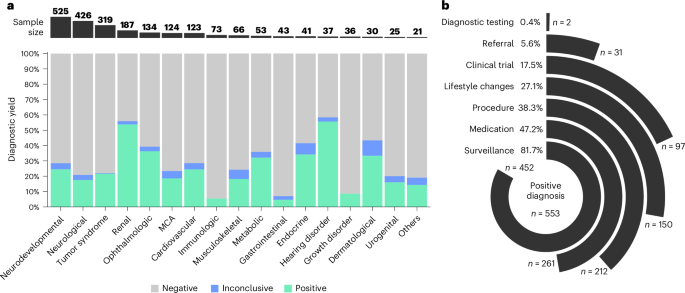
Clinical utility is defined as the percentage of individuals experiencing potential changes to clinical management after a diagnosis, which helps to accelerate decision-making and the consensus formulation process for all relevant stakeholders. The potential change in clinical management was classified into seven categories according to Riggs et al. and the UK 100,000 Genomes Project19,56: (1) referral to specialist(s); (2) indication for further diagnostic tests to evaluate possible complications; (3) initiation or contraindication of interventional or surgical procedures; (4) surveillance for potential future complications; (5) initiation or contraindication of medications; (6) lifestyle changes; and (7) clinical trial eligibility (meet enrollment criteria for phase 2 or higher interventional (related to drugs, medical devices, procedures and vaccines as defined in https://clinicaltrials.gov/) or observational (focused on assessing non-interventional biomedical or health outcomes) trial studies listed in https://clinicaltrials.gov/ or https://www.clinicaltrialsregister.eu/ that were related to the patient’s target gene and disease at the time of diagnosis).
Diagnostic odyssey
The diagnostic odyssey is defined as the time from when the disease’s symptoms are first noted in the proband (odyssey start date) to the time when a genetic diagnosis is reached. We determined the odyssey start date by retrieving the earliest record in the clinical management system that describes the symptoms of the primary indication(s) when referred to the HKGP. The date of genetic diagnosis was determined on the basis of the date at which the HKGP issued the report to the referring clinician. The diagnostic odyssey was calculated as the date of genetic diagnosis minus the odyssey start date, rounded to the nearest year; for odysseys shorter than 1 year, duration was calculated in months.
Founder mutation screening
Novel potential founder mutations were assessed in this study. The following selection criteria were applied for novel founder mutations: (1) repeated occurrence among the participants in this study, (2) absence in the gnomAD non-East Asian genome dataset and (3) absence in ClinVar. For known variants, Chinese-specific founder mutations were directly collected from the literature and compared with our findings. Shared haplotype analysis was conducted for both novel and known potential founder mutation loci among related participants carrying the mutation. This analysis used IBDseq57 for common variants (minor allele frequency >0.5% in this study).
Estimation of ACF
To estimate the ACF, all possible mating combinations among unrelated Chinese participants included in this study were evaluated. Specifically, (1) all pairings, irrespective of sex, were considered for autosomal recessive genes (\({C}_{2}^{n}\) pairings in total; n is the number of unrelated Chinese participants), and (2) only female‒male pairings were assessed for X-linked genes. A virtual couple was classified as ‘at-risk’ if both individuals carried P/LP variants in any of the same autosomal recessive genes or if the female carried P/LP variants in any X-linked genes. The ACF estimated through random mating was then compared to the observed frequency of actual couples carrying P/LP variants in the same gene within this cohort.
Re-tiering CS genes based on ACMG guidelines for the Chinese population
Genes were re-tiered on the basis of ACMG CS guidelines, with carrier frequency thresholds applied to the gene-specific GCF derived from Chinese population data in the HKGP. Tier 1 was unchanged and includes CFTR, SMN1/SMN2, HBA1/HBA2 and HBB. Tier 2 included genes associated with severe or moderate phenotypes and a carrier frequency of at least 1/100 in autosomes in our Chinese population, whereas tier 3 included genes with carrier frequencies of at least 1/200 in sex chromosomes or autosomes. This tiering approach was designed to reflect population-specific genetic characteristics while maintaining consistency with the ACMG’s evidence-based recommendations. cGCFs for different tiers were compared for this Chinese tier and ACMG pan-ethnic tiers for the Chinese population and other populations in the gnomAD 4.0 database13.
Pharmacogenomics
Gene selection and individual selection
To profile the actionable pharmacogenomic variants, we consolidated a gene list of 25 pharmacogenes with PharmGKB Clinical Annotation Level 1A or 1B (Supplementary Table 9). Among the 25 pharmacogenes analyzed, seven pharmacogenes (CACNA1S, CFTR, DPYD, G6PD, MT-RNR1, RYR1and VKORC1) are associated with congenital diseases as classified by ClinGen with definitive, strong or moderate gene−disease validity or as ‘green’ (diagnostic) or ‘amber’ (borderline) in relevant disease panels in Genomics England PanelApp and PanelApp Australia (similar gene selection approach for the diagnostic cohort). To avoid confounding effects from these conditions, individuals from the HKGP Chinese cohort were excluded from the analysis if their own or their offsprings’ primary phenotypes matched the associated congenital diseases. The remaining individuals were included for the pharmacogenomic analysis of known alleles and novel variants.
Known pharmacogenomic variants
Genotyping of known alleles of the 25 selected pharmacogenes was conducted using various tools: (1) Cyrius version 1.1.1 (ref. 58) for CYP2D6 alleles, (2) HLA-HD version 1.7.0 (ref. 59) for HLA-A and HLA-B alleles, (3) Aldy version 4.6 (ref. 60) for other pharmacogenes with star allele nomenclature and (4) VCF-derived for pharmacogenes defined by dbSNP rsIDs. Allele function and phenotype were determined on the basis of information sourced from CPIC and PharmGKB (accessed 12 November 2024). Variants listed in the AMP’s minimum sets for pharmacogenomic testing are also labeled in the same table.
To investigate the discrepancy between the Chinese population and the population with maximum sample size in CPIC, we followed the definitions and methods described by Hernandez et al.17 to compare the differences in the frequencies of altered functional alleles.
To further investigate the significance of the clinical impact of the actionable phenotypes in pharmacogenes, we categorized actionable phenotypes according to the three sections defined by the FDA Tables of Pharmacogenetic Associations (www.fda.gov/medical-devices/precision-medicine/table-pharmacogenetic-associations) (Supplementary Table 11).
Novel variants in LoF pharmacogenes
To further investigate novel deleterious variants in pharmacogenes, SNVs, indels, CNVs and SVs were detected using the same methodology described earlier. This analysis focused on nine pharmacogenes for which no-function alleles have been defined to be associated with actionable phenotype by CPIC or PharmGKB (CYP2B6, CYP2C9, CYP2C19, CYP2D6, DPYD, G6PD, NUDT15, SLCO1B1and TPMT). These genes were selected based on the rationale that LoF is a mechanism associated with their actionable phenotype. Only putative protein-disrupting variants, including frameshift, inframe, splicing and nonsense variants in these genes with PVS1 strength reaching ‘very strong’ from AutoPVS1, were included in this study after manual investigation on IGV for to ensure high-quality variants.
Estimated actionable prescriptions in Hong Kong
To examine the pharmaceutical landscape in Hong Kong, the prescription records of all medications from hospitals under the Hong Kong Hospital Authority between 1 December 2023 and 30 November 2024 were retrieved from the Clinical Data Analysis and Reporting System (CDARS) database. The top 50 drugs were selected on the basis of the total prescription count during this period. We estimated the number of actionable prescriptions by multiplying the frequency of pharmacogenomic actionable phenotypes, as defined in PharmGKB and CPIC and identified in HKGP’s data, for each individual pharmacogenomic gene. To further study the clinical relevance, we analyzed these prescribed drugs using the FDA’s Table of Pharmacogenomic Biomarkers in Drug Labeling (www.fda.gov/drugs/science-and-research-drugs/table-pharmacogenomic-biomarkers-drug-labeling) and identified clinically consequential pharmacogenomic information with three key labeling sections: adverse reactions, warnings and precautions and dosage and administration.
Results reporting
Primary findings
Building upon patient and clinician feedback, we will continue to prioritize returning clinically significant findings directly related to the referral indication and clinical phenotype.
Additional medically actionable findings
Dominant disorders
For participant opt-in for feedback of additional findings of GS, we developed a plan for reporting and returning findings in 13 genes (of which 12 are associated with dominant disorders)—MLH1, MSH2, MSH6, MUTYH, APC, BRCA1, BRCA2, VHL, MEN1, RET, LDLR, APOBand PCSK9—based on clinical actionability and severity. In compliance with ACMG guidelines and reporting guidance, only P/LP variants will be reported (https://search.clinicalgenome.org/kb/genes/acmgsf). This structured approach ensures responsible return of high-impact genetic information while respecting clinical context and participant preferences.
Recessive disorders
For reporting and returning additional findings of MUTYH-associated polyposis, only individuals with two identified disease-causing variants will receive results. Regarding expanded CS, we are at the crossroads. Although we will continue to return carrier status upon patient request, this study reinforces our decision to develop a Chinese-specific CS panel rather than relying solely on resources based on European ancestries, such as ACMG and ‘Mackenzie’s Mission’. We have demonstrated our capability to identify and return these results to patients.
Pharmacogenomics
Given the potential for broad impact, we are now initiating comprehensive review with our scientific and ethics advisory committees to explore strategies for pharmacogenomics implementation.
Statistics and reproducibility
All statistical analyses were performed using R version 4.3.3. Diagnostic yield comparisons for the diagnostic cohort and cGCF comparison in recessive genes were performed by the one-sided χ2 test (Extended Data Table 2 and Supplementary Table 8).
ACF comparisons were performed by two-sided Fisher’s exact test for each gene, and the P value was further corrected by Bonferroni correction for multiple testing on multiple genes (Supplementary Table 7). The significance level was set as P < 0.05 for all analyses in this study.
No statistical method was used to predetermine sample size. The sample size for the diagnostic cohort was determined by including all the HKGP participants who finished genetic diagnosis by November 2024 in HKGI. The sample size for the HKGP Chinese cohort was determined by including all unrelated Chinese participants who finished variant analysis by the same cutoff date.
For both cohorts, individuals with sequencing data who failed the quality control were excluded in this study. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.